Abstract
Investigations into the underlying genetic contributions to human disease are transitioning from small family-based traditional linkage analyses to large population-based studies designed to identify genetic factors in more complex and common diseases that have the greatest impact on human health. These types of studies have driven the need for larger numbers of samples for analysis and more efficient and effective methods for DNA purification, especially for large samples that provide sufficient quantities of DNA for extensive analysis. The Autopure LS™ Nucleic Acid Purification Instrument, by Gentra Systems, Inc., a platform capable of high-throughput sample purification from large samples, was developed to meet the demands of these large studies. This article presents data demonstrating the equivalency of DNA purified using the Autopure LS automated instrument and the manual method based on the same purification process. In addition, we present data demonstrating the in-lab time savings realized by automating the purification process.
INTRODUCTION
Increasingly investigations into the underlying genetic contribution to human disease rely upon large population-based association studies capable of detecting genetic polymorphisms that influence disease susceptibility, initiation, progression, severity, prognosis and response to therapy. These types of investigations have been initiated for a wide range of diseases including diabetes, a variety of cancers, cardiovascular disease, obesity, schizophrenia, Alzheimer's, hypertension and asthma, among many others. 1––––––7 In addition, similar studies are now being incorporated into clinical trials as part of the drug discovery process to better understand the genetic contributions to Adverse Drug Responses (ADR) which account for thousands of deaths each year.
With the numbers of samples predicted to increase into the millions, there is a need to automate the process of purifying DNA from a variety of biological samples to support these studies. 8,9 The longitudinal nature of these studies and the extensive analyses of these samples require that large quantities of DNA (sometimes milligram quantities) suitable for long term storage be effectively and efficiently purified from a variety of sample types. The most common source of DNA for these studies is whole blood although additional sources such as biopsy samples, cultured cells and buccal cells are also important and increasingly common.
Liquid phase DNA purification methods are often employed for these samples which are normally collected in larger volumes (e.g. 10 ml whole blood). Liquid phase chemistries, such as the Puregene® DNA Purification Kit (Gentra Systems, Minneapolis, MN), offer the distinct advantages of scalability and quality of DNA in comparison to solid phase chemistries. The scalability of the purification chemistries allows for efficient purification of DNA in high yields when working with such large samples. In addition, the high purity achieved with the Puregene method allows the DNA to be stored for long periods of time (stability studies have demonstrated that the Raregene purified DNA is stable for at least 10 years upon storage at 4°C). This is an important consideration for association studies and clinical trials where the DNA is often banked for future analyses.
This article presents the results of performance evaluations of the Autopure ls robotic system which automates the Puregene DNA purification process for large blood samples in the 1 to 10 mL volume range and other sample types.
MATERIALS AND METHODS
INSTRUMENT DESCRIPTION
The Autopure ls instrument automates Gentra's Puregene chemistry and features on-board sample tracking, precision reagent dispensing, controlled mixing and pouring, gravimetric weight verification and indexed centrifugation. Batches of eight or 16 samples can be processed in two size ranges, 1–5 mL or 5–10 mL volumes. Samples are poured into specially designed 50 mL input tubes with bar coded red caps and placed into an input rack supplied with the system. An equal number of bar coded blue capped tubes are placed into the input rack to receive the purified DNA. After the operator places the input rack into the instrument and starts the purification process, the instrument begins by recording the input and output tube weights. Because sample volumes vary, the blood weights are adjusted with the first reagent, RBC Lysis Solution to ensure that the centrifuge is balanced. The system runs Microsoft® Windows operating system software and outputs the data into Microsoft® Excel file format.
SAMPLE SET UP
Blood samples were collected in 10 mL Vacutainer® Brand blood collection tubes (EDTA K3 No. 16852, Becton Dickinson, Franklin Lakes, NJ) and stored at 4°C until use. White cell counts were determined using a Coulter Counter® CBC-5 (Coulter Electronics, Inc., Hialeah, FL) calibrated using CBC-7 Hematology Controls (R&D Systems, Minneapolis, MN). DNA was purified from the blood samples within 96 hours of being drawn. DNA purified manually using the standard Puregene DNA Purification Kit (Cat. No. D-50K, Gentra Systems, Inc., Minneapolis, MN) was compared to DNA purified using the Autopure ls instrument. DNA was purified from 16 whole blood samples using the Autopure ls instrument according to the manufacturer's directions using the Puregene reagents supplied (Gentra Systems, Inc., Minneapolis, MN). Upon completion of the purification process, approximately 80 minutes for 16 samples, the purified DNA was rehydrated overnight on a rotator (Clay Adams Nutator®, Fisher Scientific Catalog No. 14–062).
GENOMIC DNA ANALYSIS
To assess DNA integrity, a volume of two μL was removed from each purified DNA sample and separated by 0.7% agarose gel electrophoresis. The DNA was electrophoresed for one hour at 75 volts using 0.125 μg/mL ethidium bromide in the gel and running buffer. The gel was photographed on a UV transilluminator using a Kodak Digital Imaging System EDAS 120 LE (Rochester, NY). Lambda DNA digested with Hind III was used as a size reference.
UV ABSORBANCE DETERMINATION
DNA yields were determined using a Beckman DU-64 UV spectrophotometer (Fullerton, CA). A 10 μL volume of each DNA sample was diluted in 190 μL ultrapure deionized water and vortexed at high speed for five seconds. To obtain the DNA concentration, the 320 nm reading (background) was subtracted from 260 nm, and the resulting value was multiplied by the DNA extinction coefficient of 50. Then, DNA yield was calculated by multiplying each concentration by the respective DNA volume as estimated by weight in grams. The A260/A280 ratio was determined by subtracting the background (A320) reading from each of the two readings before they were divided.
DNA RESTRICTION ENZYME DIGESTION
DNA quality was evaluated further by analyzing digestion with a panel of six restriction endonucleases. In a digest volume of 25 μL, two units of EcoR I, Hae III, Msp I, Pst I, Hind III, Xba I (New England Biolabs, Beverly, MA and Sigma, St. Louis, MO) were added to one μg DNA and allowed to digest for 30 minutes at 37°C. To examine the samples for digestion, a volume of two μL was analyzed by 0.7% agarose gel electrophoresis as described above.
DNA AMPLIFICATION
Purified DNA was evaluated for amplification performance using primers specific to a 1.5 kb target in the CYP2D6 locus, one of the cytochrome P450 genes (10). For each reaction, a quantity of 100 ng DNA was amplified in a 25 μL volume containing: 1X Taq Polymerase Buffer, 0.05 U/μL Taq Polymerase, 1.5 mM MgCl2, and 0.2 mM each dNTP (Promega, Madison, WI) as well as forward and reverse primers at 1 μM (Research Genetics, Huntsville, AL). The amplification conditions for the CYP2D6 target were: 30 cycles: 94°C 30 seconds, 63°C 30 seconds, and 72°C 30 seconds; 72°C hold six minutes, 4°C hold. A volume of 10 μL from each reaction was analyzed by 2% agarose gel electrophoresis as described above.
RESULTS
The Autopure ls instrument automates the Puregene purification process consisting of six primary steps as described in Figure 1. The method is based on the preferential precipitation of proteins from a crude lysate and subsequent precipitation of the DNA with isopropanol.
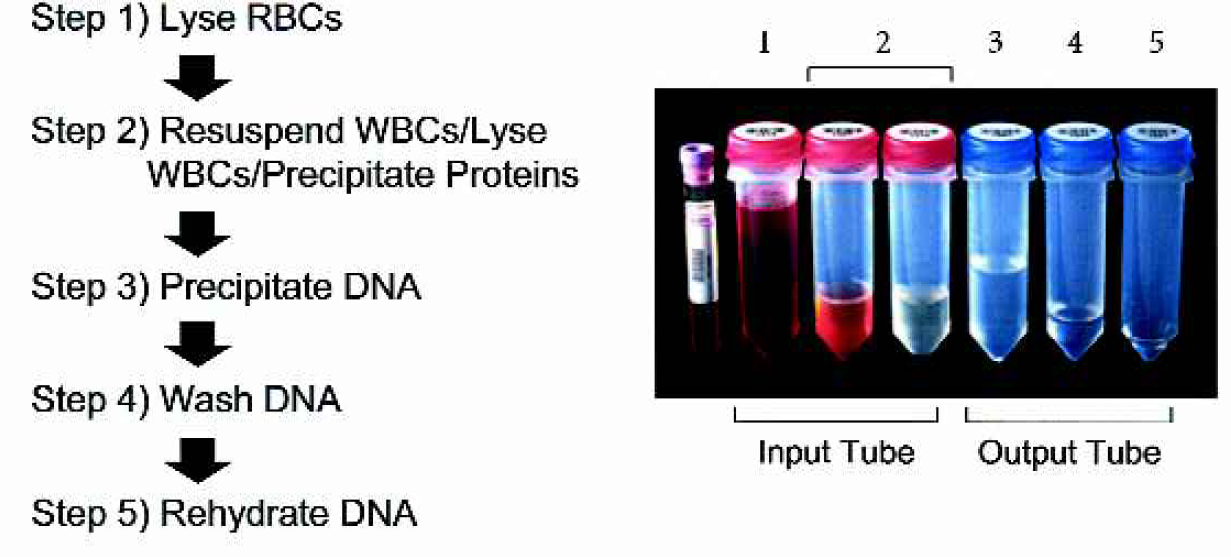
The Autopure ls automated DNA purification method. Purification of DNA from whole blood on the Autopure ls instrument is performed in five steps, 1) lysis of the red blood cells followed by centrifugation to collect the white blood cells, 2) lysis of the white blood cells and precipitation of protein, followed by centrifugation to pellet the protein, 3) precipitation of the DNA with isopropanol followed by centrifugation to pellet the DNA, 4) washing of the DNA with 70% ethanol and 5) addition of rehydration solution to rehydrate DNA.
In order to demonstrate the effective purification of DNA on the Autopure ls, samples were purified in parallel using the standard manual Puregene method and the Autopure ls instrument using replicate human whole blood samples collected from a single donor.
Electrophoretic analysis of the purified samples demonstrate that the DNA purified on the instrument were equivalent in molecular weight in comparison to the DNA samples purified manually (Figure 2). All samples appeared intact and undegraded as evidenced by the lack of low molecular weight DNA.
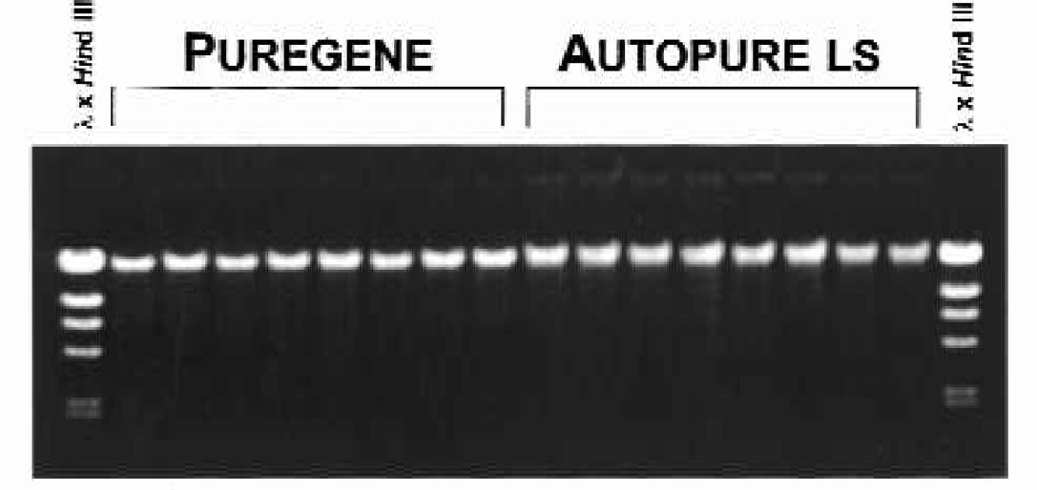
DNA purified with the manual Puregene method and the Autopure ls instrument results in equivalent quality and yields of DNA. DNA was purified from replicate 10 ml human whole blood samples with the manual Puregene method and the Autopure ls as indicated. Purified DNA (100 ng) was analyzed by gel electrophoresis on a 0.7% agarose gel (see Materials and Methods).
The quantity and quality of the purified DNA samples were also analyzed spectrophotometrically to determine DNA yield (Figure 3) and A260/A280 ratio (Figure 4). DNA yields were 348.3 (+/−4.3) μg DNA (n = 8) for the manually purified samples and 350.2 (+/–5.1) μg DNA (n = 8) for Autopure ls purified samples. The A260/A280 ratios were essentially identical for DNA samples purified with either the manual or the automated method: 1.839 +/–0.007 (n = 8) for manual samples and 1.840 +/–0.016 (n = 8) for samples run on the Autopure ls instrument.
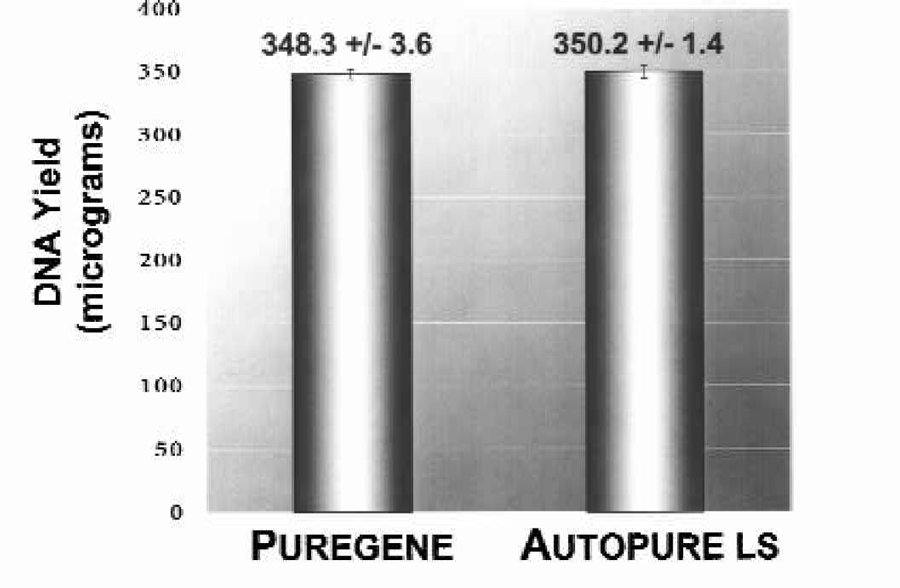
DNA purified with the manual Puregene method and the Autopure ls instrument results in equivalent yields of DNA.
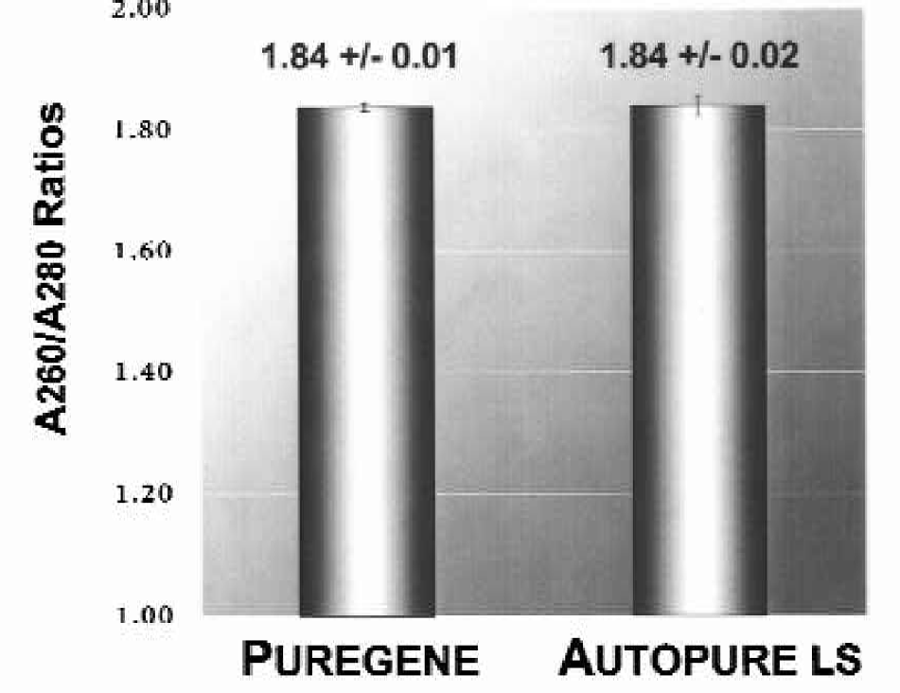
DNA purified with the manual Puregene method and the Autopure ls instrument results in equivalent quality of DNA. DNA was purified from replicate 10 ml human whole blood samples with the manual Puregene method and the Autopure ls instrument as indicated. Purified DNA was analyzed by UV spectrophotometry to determine A260/A280 ratios.
To further assess the quality of the DNA, the samples were digested with six restriction enzymes and showed no difference in restriction enzyme digestion ability (Figure 5). Another measure of DNA quality was PCR amplification performance. We observed no detectable difference in amplification ability of the DNA samples purified manually and those purified on the Autopure ls instrument for the 1.5 kb CYP2D6 target (Figure 6). DNA purified with the Autopure ls instrument also amplified very efficiently with primers specific for a variety of other loci including HLA-H, MTHFR, HLA-A, Factor V, D1S80, and Factor II (data not shown).
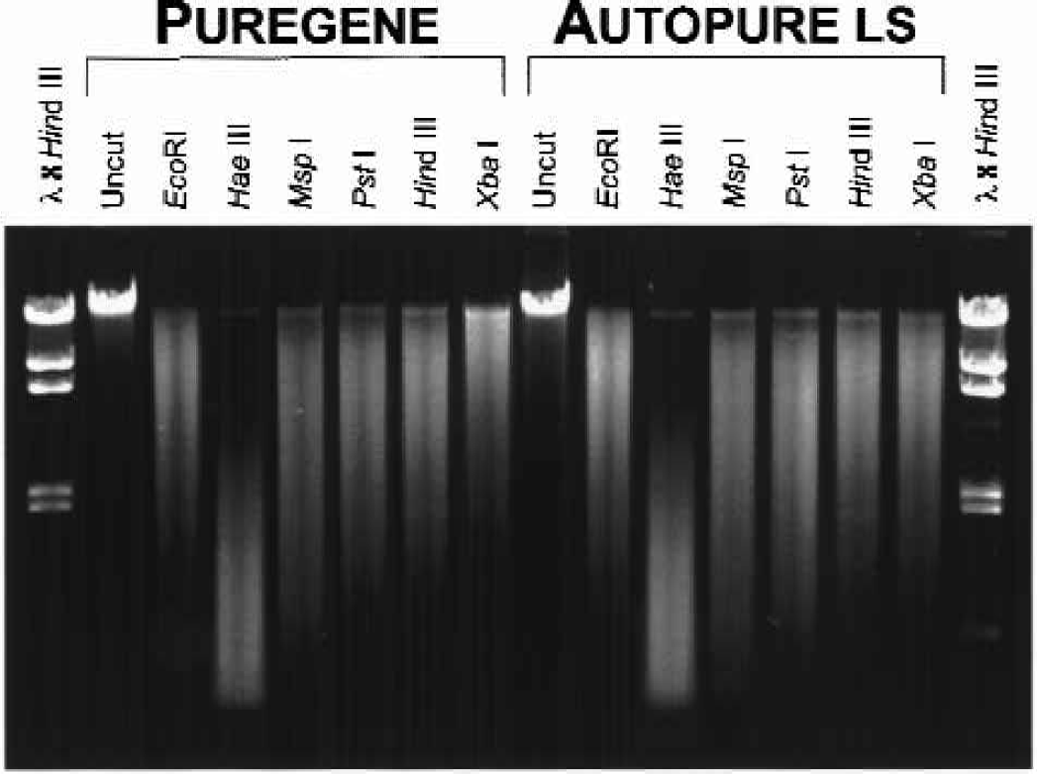
DNA purified with the manual Puregene method and the Autopure ls instrument perform equivalently in restriction endonucleases digestions. DNA was purified from replicate 10 ml human whole blood samples with the manual Puregene method and the Autopure ls instrument as indicated. DNA (2 μg) was digested with a 2-fold excess of enzyme for 30 minutes at 37 °C. Digested DNA was then analyzed by electrophoresis on a 0.7% gel
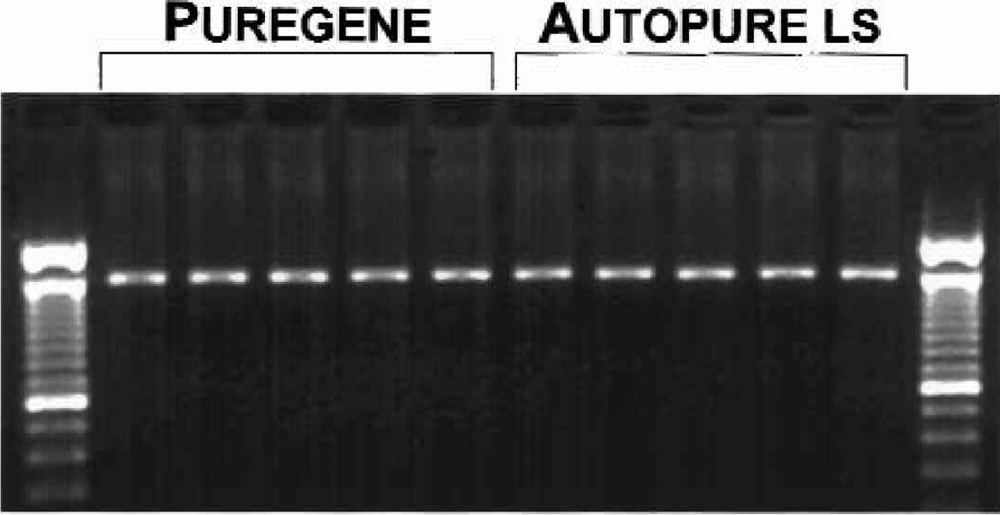
DNA purified with the manual Puregene method and the Autopure ls instrument perform equivalently in PCR amplifications. DNA was purified from replicate 10 ml human whole blood samples with the manual Puregene method and the Autopure ls instrument as indicated. DNA was amplified with primers specific for the CYP2D6 locus, resulting in a 1.5kb amplicon (see Materials and Methods). PCR products were analyzed by gel electrophoresis on a 1.0% agarose gel.
Lastly, the quality of the DNA was assessed by Southern blot analysis. The blot was probed with a T cell receptor (TCR) specific probe commonly used for detection of TCR gene rearrangements. As seen in Figure 7, all samples were efficiently digested with the restriction endonuclease and a single germline band was easily detected. Again, the DNA purified on the Autopure ls instrument performed equivalently to the DNA purified manually.
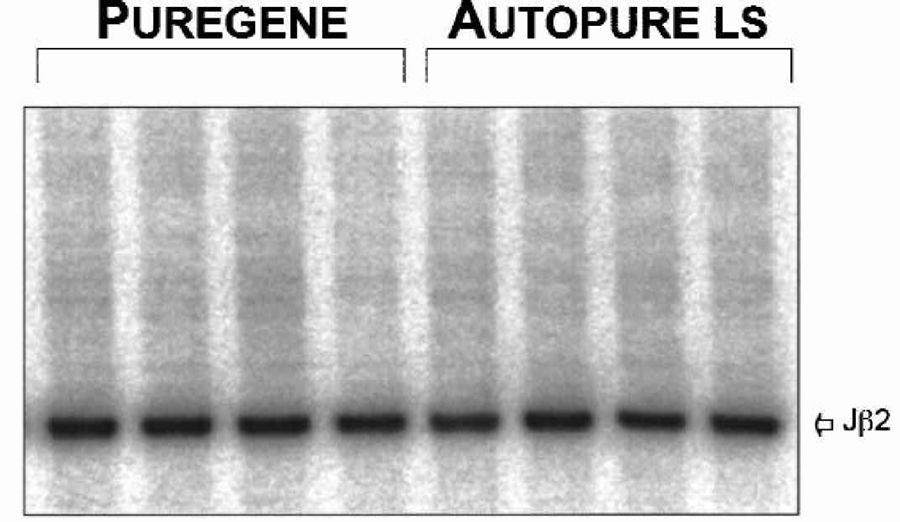
DNA purified with the manual Puregene method and the Autopure ls instrument perform equivalently in Southern blot analyses. DNA was purified from independent human whole blood samples with the manual Puregene method and the Autopure ls instrument as indicated. DNA was analyzed by Southern blot using probes specific for the Jβ2 locus to analyze T-cell receptor gene rearrangements.
While the Autopure ls instrument was designed to purify DNA from large samples like the 10 ml whole blood samples described above, there is a growing need to purify DNA from buccal samples that typically yield far less DNA. We performed preliminary feasibility experiments with buccal mouthwash samples to assess the ability of the Autopure ls instrument to purify DNA from these samples. As seen in Figure 8, proteinase digestion was found to dramatically increase the yield and consistency of DNA recovery from these samples (3.85μg vs 11.85μg). The yield of DNA from the buccal mouthwash samples was found to be highly variable but the quality, as evidenced by high molecular weight DNA, was comparable to DNA isolated from whole blood. We also found that the DNA performed well in PCR amplifications and restriction endonucleases digestions (data not shown). These results suggest that the Autopure ls instrument is capable of purifying limited quantities of DNA from difficult samples.
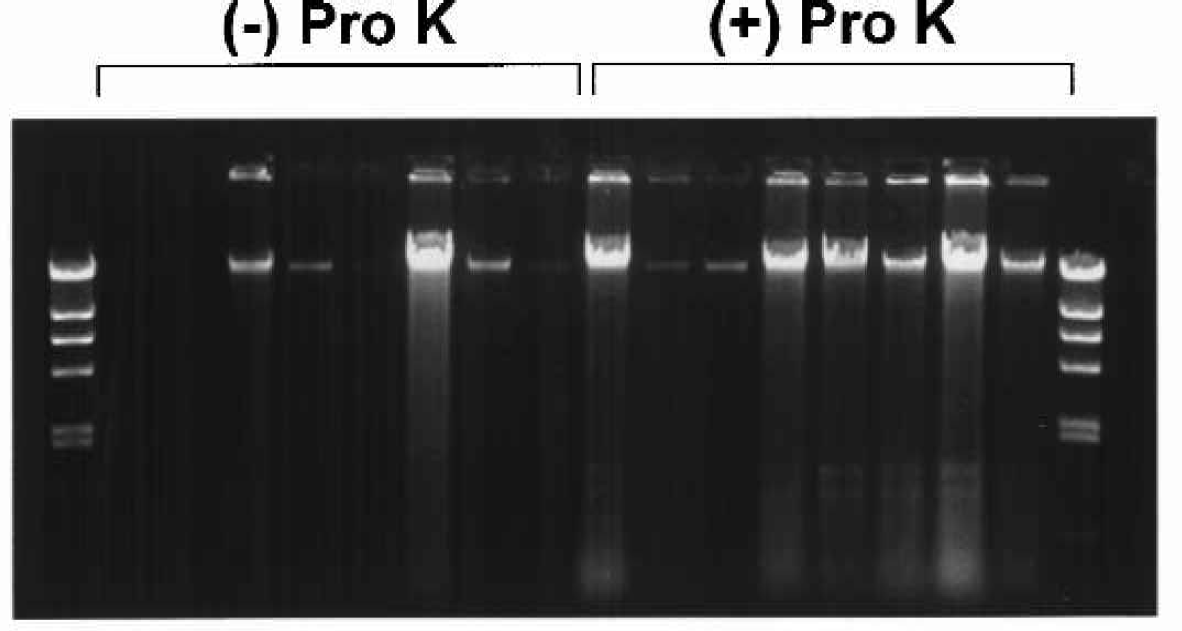
Effective automated purification of DNA from buccal mouth-wash samples. Buccal mouthwash samples were collected from 16 different donors. Samples were then treated with Proteinase K as indicated. DNA was then purified from the samples on Autopure ls instrument subassemblies. The purified DNA was analyzed by gel electrophoresis on a 0.7% agarose gel. As seen above, pre-treatment with Proteinase K resulted in a significantly greater yield of DNA.
The purification process on the Autopure ls instrument is completely automated. Operation of the Autopure ls instrument requires less than 10 minutes to set up a rack of 16 samples. The samples are scanned into the instrument, transferred to a 50 ml processing tube and then loaded into a sample rack. In order to quantitate the anticipated impact of automation on productivity, we studied the amount of time the lab spent on DNA purification and genotyping before and after integration of the Autopure ls instrument. As shown in Figure 9, the instrument dramatically increased the amount of time available for genotyping. Prior to automation of the DNA purification process, the lab had only 36% of its time available for genotyping. Since implementing the Autopure ls instrument for purification of DNA, the lab now has over 70% of its time dedicated to genotyping. We estimate that this has saved over 2,400 hours of labor that was previously spent on manual purification of samples.
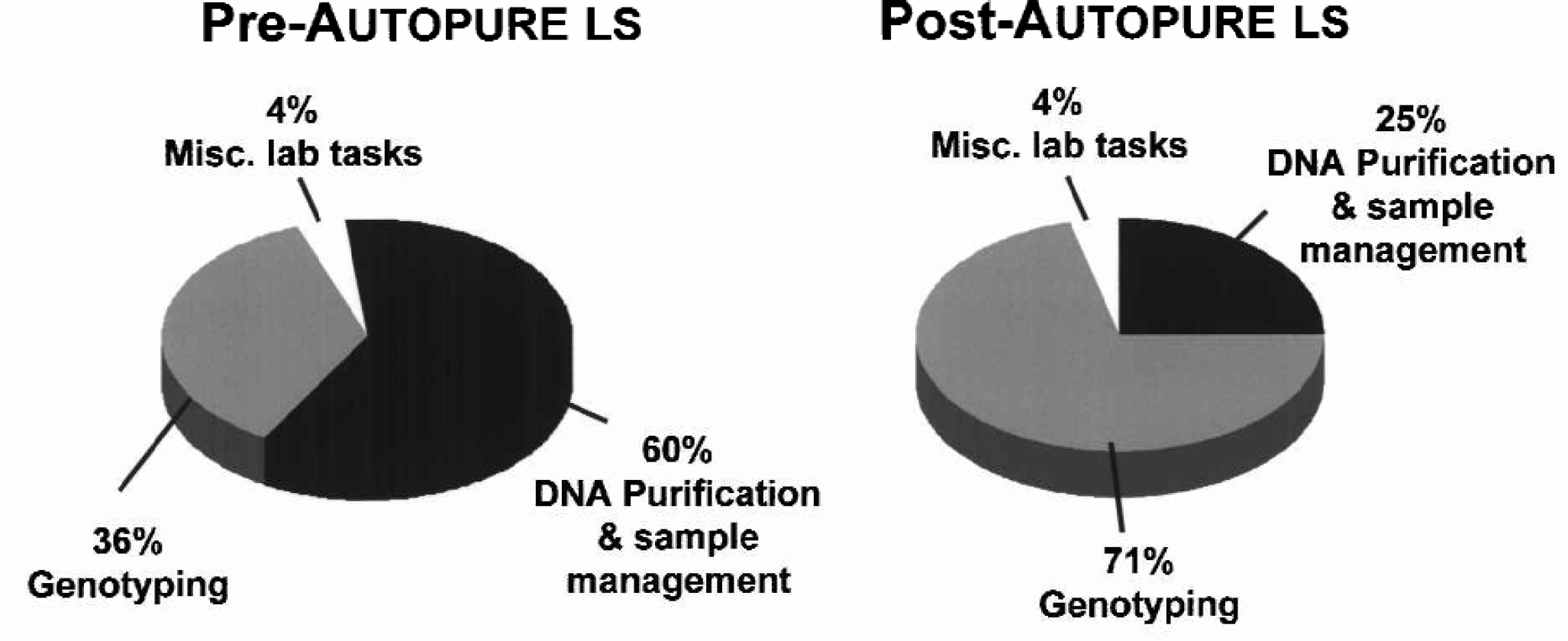
The effect of automated DNA purification on labor allocation in the laboratory. The amount of time spent on three major activities in the genotyping laboratory was measured before and after the Autopure ls instrument installation as indicated. As seen in the figure, automation of the DNA purification process resulted in a significant increase in the time available and dedicated to genotyping.
In conclusion, the performance of the Autopure ls instrument has proven to be equivalent to the manual Puregene DNA purification process. The quantity, quality and performance of the purified DNA in downstream assays were found to be consistent and reliable. The DNA performed without failure in a variety of downstream assays including PCR, Southern blot, restriction endonucleases digestion, Invader® assays (Third Wave Technologies, Madison, WI, data not shown) and MASSArray™ (Sequenom, data not shown). The Autopure ls instrument has also proven to dramatically decrease the hands-on time required for DNA purification and radically transform the laboratory workflow. Given the demonstrated efficiency and effectiveness of DNA purification on the Autopure ls, the instrument promises to be a useful tool for laboratories performing large sample nucleic acid purifications where yield, quality and stability of the purified samples is critical.
