Abstract
Background
Some studies have reported that deletions at chromosome arm 9p occur frequently and represent a critical step in carcinogenesis of some neoplasms. Our aim was to evaluate the deletion of locus 9p21 and chromosomes 3, 7 and 17 in localized prostate cancer (PC) and correlate these alterations with prognostic factors and biochemical recurrence after surgery.
Methods
We retrospectively evaluated surgical specimens from 111 patients with localized PC who underwent radical prostatectomy. Biochemical recurrence was defined as a prostate-specific antigen (PSA) >0.2 ng/mL and the mean postoperative follow-up was 123 months. The deletions were evaluated using fluorescence in situ hybridization with centromeric and locus-specific probes in a tissue microarray containing 2 samples from each patient. We correlated the occurrence of any deletion with pathological stage, Gleason score, ISUP grade group, PSA and biochemical recurrence.
Results
We observed a loss of any probe in only 8 patients (7.2%). The most common deletion was the loss of locus 9p21, which occurred in 6.4% of cases. Deletions of chromosomes 3, 7 and 17 were observed in 2.3%, 1.2% and 1.8% patients, respectively. There was no correlation between chromosome loss and Gleason score, ISUP, PSA or stage. Biochemical recurrence occurred in 83% cases involving 9p21 deletions. Loss of 9p21 locus was significantly associated with time to recurrence (p = 0.038).
Conclusions
We found low rates of deletion in chromosomes 3, 7 and 17 and 9p21 locus. We observed that 9p21 locus deletion was associated with worse prognosis in localized PC treated by radical prostatectomy.
Introduction
Prostate cancer (PC) is the most common noncutaneous cancer and the second leading cause of death in males in Western countries (1). Its biological behavior is heterogeneous; PC is clinically insignificant in a considerable proportion of patients but is aggressive in others, and the current prognostic factors are not able to precisely discriminate between them (2).
Screening PC using prostate-specific antigen (PSA), although controversial, has made early diagnosis possible. In the United States, where screening was embraced, more than 75% of PCs are stage T1c (1). However, overdiagnosis is a well-known side effect of PSA-based screening and may lead to the diagnosis of clinically insignificant tumors. Confirming this hypothesis, the largest screening trial found that 781 men should be screened to avoid 1 death due to PC at a 13-year follow-up (3). This finding emphasizes that PSA is not able to differentiate aggressive from indolent tumors and that a new biomarker is required.
Radical prostatectomy (RP) is a treatment option for organ-confined tumors and demonstrates good results in large surgical series; however, up to a quarter of patients experience recurrence after surgery (4). Prior identification of patients with poor prognosis and higher chances of recurrence is important to determine cases requiring adjuvant treatments. The well-established classic prognostic parameters, such as Gleason score, stage and PSA, can estimate aggressiveness and recurrence in PC. However, these parameters are often deficient, even when used in combination, which may be due to the biological variability of PC (5, 6).
These reasons underscore the need for new biomarkers that will not only enable the identification of PC patients that may benefit most from surgery, but will also prevent the over-treatment of patients with indolent tumors, thereby improving PC management and treatment results. From a molecular point of view, cancer is the consequence of a combination of single-nucleotide polymorphisms, insertions, deletions, gene fusions and epigenetic events, leading to changes in the expression of oncogenes and suppressor genes that play important roles in carcinogenesis. Therefore, the search for new biomarkers has focused on genetic changes associated with tumor development and disease prognosis (7, 8).
Loss of heterozygosity in a tumor suppressor locus is a common genetic alteration associated with malignancy, and the loss of various loci on chromosome 9 has been reported in various cancers (9–11). Global analyses of copy number profiles of primary tumors and metastases have identified recurrent aberrations associated with PC progression, such as broad deletions on the 1p, 6q, 8p and 9p chromosome arms and gains on the 1q, 3q, 7q and 8q genes (12). In vitro studies showed that the 9p21 and 7q22 loci are frequently deleted regions in PC cell lines (13). In agreement with this, loss of heterozygosity at the 9p21 locus was observed in 72% of 40 PC surgical specimens (14).
Although previous studies have examined genic deletions in PC, most of them have focused on describing the occurrence of deletions rather than examining their correlations with postsurgical tumor outcomes (15, 16). The aim of this study was to assess the occurrence of losses at chromosomes 3, 7, and 17, and locus 9p21 in surgical specimens of clinically localized PC assembled in a tissue microarray using fluorescence in situ hybridization (FISH), and to evaluate the association between these losses and prognostic factors and biochemical recurrence following RP.
Materials and Methods
Case selection
We retrospectively evaluated patients with clinically localized PC who underwent curative RP between January 1994 and April 2000, all performed by the same surgeon. For this case-control study, we selected a case group of 51 patients with low-, intermediate- or high-risk PC who experienced biochemical recurrence after surgery. Then, 60 patients who did not experience biochemical recurrence within 10 years after surgery were selected as controls; they were matched to the first group according to the PC stratification of risk (17). We stratified the patients into 3 risk groups according to a combined evaluation of pre-operative PSA, pathological stage and Gleason score. Low-risk PC patients were characterized by a PSA <10 ng/mL, a Gleason score ≤6 and a pT2a stage. The PC risk was considered to be intermediate when the PSA level was 10-20 ng/mL, Gleason score of 7, and a pT2b stage. A high risk was defined as a PSA level ≥20 ng/mL, a Gleason score ≥8, and a pathological stage higher than pT2c. This risk stratification is similar to that proposed by D'Amico et al; the difference is the inclusion of the pathological stage and Gleason score in our study, which are more reliable in terms of prognostic strength when compared with the clinical parameters employed in criteria of D'Amico et al (18).
We also considered the tumor grade according to the recently modified International Society of Urological Pathology (ISUP) grading criteria that stratify patients into 5 groups. With this classification, the Gleason scores 3 + 3, 3 + 4, 4 + 3, 4 + 4 and 9-10 became grade I, II, III, IV and V groups, respectively (19, 20).
The median follow-up time was 123 months. The participants included 97.3% white and 2.7% Asian patients. The demographic and clinical data according to recurrence status are shown in Table I. Tumor recurrence was defined as a PSA level exceeding 0.2 ng/mL at follow-up. The tumor-node-metastasis (TNM) staging designations were assigned according to the TNM 2010 classifications. All patients provided informed consent to participate in the study and allow their biological samples to be analyzed. Approval for the study was granted by the Institutional Board of Ethics of UNINOVE (n°206.920).
Demographic and clinical data: patients with recurrence vs. no recurrence
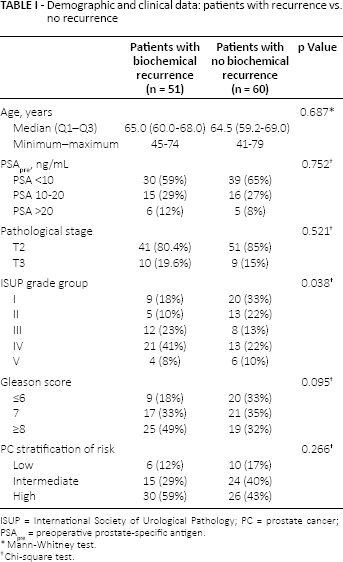
ISUP = International Society of Urological Pathology; PC = prostate cancer; PSApre = pre-operative prostate-specific antigen.
Mann-Whitney test.
Chi-square test.
Tissue microarray
Histological examination of the surgical specimens was performed in formalin-fixed and paraffin-embedded sections stained with hematoxylin and eosin. The slides containing the primary tumor for each patient were selected by sampling the area representative of the final pathological Gleason score. Two areas from each tumor were marked with permanent ink. Marked areas were included in the tissue microarray, which was constructed using a Beecher manual arrayer (Beecher Instruments, Sun Prairie, WI, USA) (21). Two cylinders with a diameter of 0.6 mm were removed from each patient's paraffin block, which contained the previously marked areas of the PC, and were transferred in 3-mm intervals to a recipient paraffin block. Next, the tissue microarray recipient block was cut into consecutive 3-μm histological sections, and the slides were used for FISH analysis.
Fluorescence in situ hybridization
The detection of chromosomes and locus deletions were performed using FISH. We employed commercially available centromeric probes specific to chromosomes 3, 7 and 17, and a locus-specific probe for 9p21 from Abbott (Des Plaines, IL, USA). The reactions followed the manufacturer's recommendations (22). The slides were serially dehydrated in 70%, 85% and 100% ethanol solutions for 2 minutes each and then placed in 2X saline sodium citrate (SSC) solution (pH 7.0) at 75°C for 10 minutes. Subsequently, each slide was immersed in a proteinase K/2X SSC solution (0.25 mg/mL) at 37°C for 10 minutes and in a 2X SSC solution at room temperature for 5 minutes. The tissue samples were again placed in 70%, 85% and 100% ethanol solutions for 2 minutes each, air dried and exposed to the Televysion probes. The slide was then covered with a 24×32-mm cover slip and sealed with rubber cement. Hybridization was accomplished by heating the slide plate at 80°C for 10 minutes and incubating it overnight in a humidified chamber at 37°C. Following the hybridization, the glue and cover slip were gently removed, and the slide was immediately immersed in a urea/0.1X SSC buffer at 45°C for 30 minutes. Subsequently, the slides were washed with 2X SSC at room temperature for 2 minutes, again serially dehydrated with ethanol, and air dried. Finally, the slide was counterstained with 10 μL of 4′,6-diamino-2-phenyl-indole/antifade and covered with a cover slip.
The analysis was performed with a fluorescence microscope using the CytoVision system for documentation. All cells in each cylindrical sample were analyzed. Samples were considered to have a deletion when there was no detectable signal for the specific probe in the fragment, so that only samples with homozygous deletions were considered positive, as previously described (23). We correlated the occurrence of each deletion with the Gleason score, ISUP grade group, PSA serum levels, pathologic stage and biochemical recurrence. All cytogenetic analyses were performed and reviewed by the same uropathologist without knowledge of the evolution of each particular case.
Statistical analysis
Statistical analyses were performed using SPSS 17.0 for Windows (IBM Corp., Armonk, NY, USA). All of the reported p values were 2-sided. Mann-Whitney, Pearson, chi-square and Fisher's exact tests were used to explore the bivariate associations between deletions and the continuous and categorical variables. Biochemical recurrence-free survival curves were generated using the Kaplan-Meier method. We performed the log-rank test to show differences between the curves.
Results
The cases and controls exhibited similar characteristics regarding their risk stratification, which demonstrated the homogeneity of this case-control study (Tab. I). The mean and median follow-up times for the entire sample were 116 and 123 months, respectively. The ISUP grading system was significantly higher in cases with biochemical recurrence (p = 0.038). During the follow-up time, 8 patients had a diagnosis of bone metastasis, 5 patients died by PC and 6 by other causes; thus the 10-year specific and overall survival rates were 95.4% and 94.5%, respectively.
There was a loss of 18 specimens during the tissue microarray assembly; as such, deletions were evaluated for 93 cases. We observed a loss of signal of any probe in 8 patients (7.2%), of whom 75% experienced biochemical recurrence. The most common deletion was the loss of locus 9p21, which occurred in 6 patients (6.4%). One patient exhibited simultaneous deletions of chromosomes 3 and 7, and another had deletions of chromosomes 3 and 17. The clinical and pathological characteristics of patients with genetic changes are described in Table II.
Clinical and pathological characteristics of patients according to type of deletion
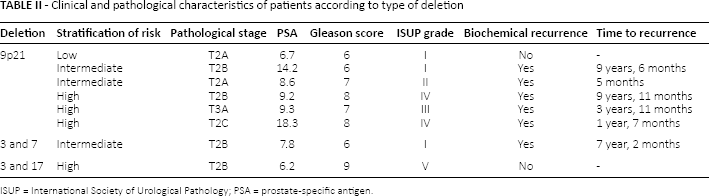
ISUP = International Society of Urological Pathology; PSA = prostate-specific antigen.
There were no significant correlations between the presence of each deletion and Gleason score, ISUP grade group, PSA serum levels or pathological stage. We examined the occurrence of any deletion in relation to prognostic factors, but again, no significant correlation was observed (Tab. III). When we examined the occurrence of any deletion with biochemical recurrence, we observed a marginally higher chance of relapse in patients with a deletion at the 9p21 locus (p = 0.084). We noted that 5 of the 6 cases (83%) with the 9p21 deletion had tumor recurrence, whereas tumor recurrence was observed in 39 out of 87 controls (45%). There were no correlations between tumor recurrence and loss of chromosomes 3, 7 or 17 (Tab. IV).
Correlation between deletions and classic prognostic factors in prostate cancer
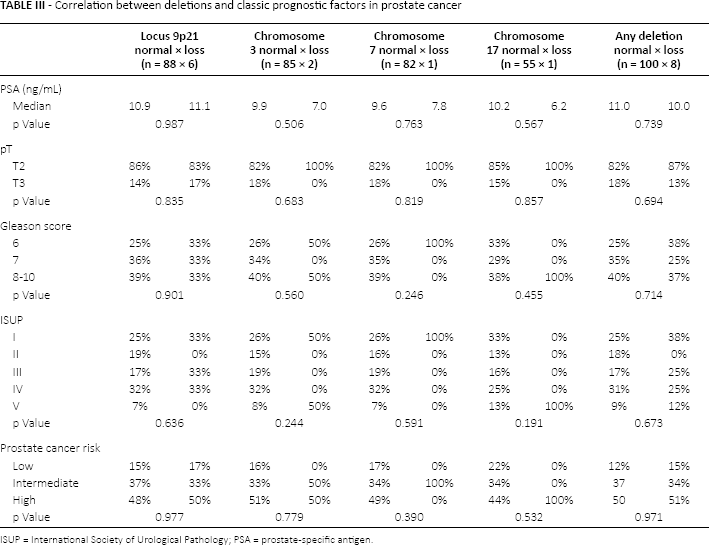
ISUP = International Society of Urological Pathology; PSA = prostate-specific antigen.
Correlation between biochemical recurrence and deletions
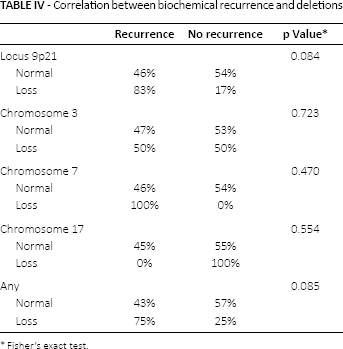
Fisher's exact test.
The Kaplan-Meier curve of biochemical recurrence-free survival according to 9p21 deletion is shown in Figure 1. The curve indicates that loss of the 9p21 locus was significantly associated with a shorter time to biochemical recurrence in localized PC treated with RP. The mean biochemical recurrence-free survival time was 80.5 and 124.3 months for deletion and control cases, respectively (log-rank test, p = 0.038). No significant correlation was observed between time to biochemical recurrence and loss of chromosome 3, 7 or 17. Regarding patients who developed bone metastases, only 1 had any chromosomal loss (9p21 deletion locus), which prevented us from making correlations. We did not find any chromosomal deletion in the 9 patients who died by PC or other causes.
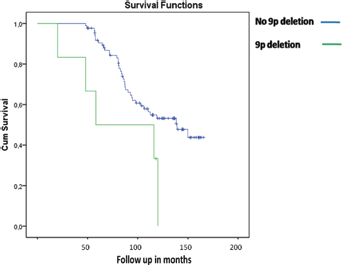
Kaplan-Meier analysis of biochemical recurrence-free survival stratified by locus 9p21 status.
Discussion
This is the first study in the literature to demonstrate that the deletion of the 9p21 locus was independently associated with a shorter time to biochemical recurrence for localized PC following RP. We also demonstrated that deletions of chromosomes 3, 7 and 17 are extremely rare in localized tumors, and therefore have no role as potential biomarkers in PC. The strong points associated with our results were the extended postoperative follow-up, the fact that all patients underwent surgery by the same surgeon, and having cytogenetic alterations evaluated by the same uropathologist. The employment of a tissue microarray for expression analysis also facilitated homogeneity in our findings.
PC is a clinically heterogeneous disease, encompassing aggressive tumors that threaten patients' lives, as well as a considerable number of indolent tumors where treatment may be more harmful than the tumor itself. Problematically, the available tools are not adequate for distinguishing between these 2 phenotypes. The clinical heterogeneity of PC, coupled with its high prevalence, raises challenges in the management of this disease. Therefore, the most necessary task in PC-related clinical oncology is the discovery of new prognostic biomarkers (24).
The identification of loss of heterozygosity on specific genetic loci and other genetic changes has been helpful for understanding carcinogenesis, and has led to the description of new biomarkers and tools for PC. Recently the Oncotype Prostate DXR was indicated as a test for counseling male patients with regard to the best course of therapy: active surveillance versus treatment (25). This tool is a quantitative RT-PCR assay performed on tissue obtained via needle biopsies, and incorporates 12 cancer-related genes that are algorithmically combined to calculate a Genomic Prostate Score, which correlates with the probability of harboring adverse pathologies at surgery (26).
Analyses of copy number alterations in PC have identified mutations associated with tumor development such as deletions of 1p, 6q, 8p and 9p and losses of large regions of chromosomes 13, 15 and 18 (21, 27–29). The locus 9p21 seems to be a critical region where losses have been correlated to worse prognoses in several cancers. Our group previously demonstrated that the loss of the 9p21 locus occurred in 15% of a sample of 94 clear cell carcinoma patients who underwent partial or radical nephrectomy. It was an independent predictor of specific mortality in multivariate analysis, and was associated with a 28-fold greater risk of death (23). Frequent allelic loss of 9p21 has also been reported in lung, ovarian, bladder, oral and head-and-neck cancers (10, 11).
The aim of the current study was to identify chromosome deletions that may contain important genes for the progression and recurrence of PC, to establish new markers of prognosis. Although we observed deletions of locus 9p21 in only 6 cases, the majority of those cases showed biochemical recurrence at follow-up. Additionally, we found that cases with locus 9p21 deletion had a mean time of 80 months to biochemical recurrence, which was significantly shorter than the mean time to recurrence for controls, 120 months. Based on these observations, we hypothesize that the deletion of the 9p21 locus may play a role in PC carcinogenesis and probably contains a tumor suppressor gene.
Contradictory to our findings, Visakorpi et al (16) evaluated 31 primary and 9 recurrent PC cases. Through comparative genomic hybridization analysis, they found that 74% of primary tumors showed DNA sequence copy number changes and that losses were 5 times more common than gains. Loss of 9p was observed in 16% of cases, and the other most common losses were 13q (32%), 6q (22%) and 16q (19%). They found that recurrence was significantly associated only with gains on chromosome arm 8q (89% of recurrence cases vs. 6% of controls) and chromosomes X (56% vs. 0%) and 7 (56% vs. 10%). They concluded that losses of several chromosomal regions are common in primary tumors and suggested that deletional inactivation of putative tumor suppressor genes in these 3 chromosomal sites is likely to underlie PC development (16).
Corroborating our findings, Perinchery et al mapped the 9p21 region in 40 primary tumors using 15 microsatellite markers and found that 72.5% of cases showed losses of heterozygosity at chromosome arm 9p (14). Only 6 of 15 microsatellite markers exhibited loss of heterozygosity in PC specimens, and the most common deletions were found on 9p21 at loci D9S1748 (50%), D9S171 (51.4%) and D9S270 (21.8%). There were also deletions on 9p22 at locus D9S162 (8.3%), on 9p13 at locus D9S273 (13.8%), and on 9p11 at locus D9S153 (7.7%). Deletions on 9p21 were associated with higher pathological stage; loss of heterozygosity at 3 or more loci occurred in 46% of stage T3 cases versus 15% of stage T2 cases, suggesting a higher incidence of loss of heterozygosity in the advanced stage of PC. They also demonstrated that the expression of p16 protein was either absent or very low, suggesting that the candidate target tumor suppressor gene at this locus was the gene that encodes protein p16 (14).
The tumor suppressor gene CDKN2A/ARF is located at the 9p21 region, and it encodes 2 major cell cycle regulatory proteins, p16 and p14, which are involved in the normal retinoblastoma (Rb/E2F) and p53 pathways (27). Loss of the CDKN2A tumor suppressor gene caused by 9p21 deletions would lead to stimulation of the cell cycle and simultaneous Rb and p53 inactivation, as has been demonstrated in bladder cancer (30). In renal cell carcinoma, the p16 gene was also postulated to be the candidate target present at locus 9p21. Grady et al (9) evaluated deletions at different loci of chromosome 9 in 60 renal cell carcinoma cases, and found that 9p21 deletion occurred in 57% cases and was associated with higher pathological staging. In their study, the expression of p16 was absent or low, underscoring the hypothesis that p16 is probably encoded by a gene at the 9p21 locus (9). Another candidate gene at 9p21 locus is the CDKN2B tumor suppressor gene, which encodes protein p15, a cyclin D-dependent kinase inhibitor that regulates the cell cycle via the Rb pathway, and the interferon-alpha and interferon-beta genes (31).
The biological variability of PC may explain the difficulty of establishing accurate prognostic markers. In this context, there is a clear need for more sensitive tests to identify patients with poor prognosis in order to improve clinical practices. Despite the existence of previous descriptions of 9p21 deletions and other copy number alterations in PC, most prior studies have focused on describing the occurrence of these deletions, rather than on examining correlations with tumor outcomes (15). Moreover, no study to date has explored the deletion of 9p21 locus in a group of patients who were homogenously treated, evaluated and followed up for more than 10 years.
We acknowledge that our study is limited by its retrospective nature and that our conclusions require additional testing. However, based on our findings that 9p21 deletion was associated with worse outcome, we postulate that this deletion is a potential biomarker for prognosis in PC. Further prospective studies with larger series should be conducted to confirm our results and identify the gene at this locus. If these results are confirmed, they could be applied in clinical practice to improve the prognostic evaluation of patients with PC. In combination with other molecular and biochemical assays, tests for 9p21 deletions may help to identify patients with greater likelihood of PC recurrence so that they can be offered adjuvant treatment, and at the same time, this may spare many patients from unnecessary adjuvant therapies and their associated complications.
Conclusions
We found that deletions on chromosomes 3, 7 and 17 were rare events in our sample of localized PC cases. We also observed that locus 9p21 deletions were significantly correlated with shorter times to biochemical recurrence after surgery.
Footnotes
Abbreviations
Financial support: No grants or funding have been received for this study.
Conflict of interest: The authors have no conflicts of interest to declare.
