Abstract
Introduction
The excision repair cross-complementation group 2 (ERCC2) ATP-dependent helicase is an essential member of the DNA repair pathway. It has been observed to be differentially expressed in different cancers, which shows its involvement in carcinogenesis.
Aim
In the present study we have tried to determine the association of expression patterns of this gene with head and neck carcinogenesis.
Method
We first carried out a systematic review of the available studies on the role of ERCC2 in head and neck cancer (HNC). In order to test the hypothesis that the expression patterns of XPD/ERCC2 play a critical role in HNC pathogenesis, we then conducted a population based case-control study on 81 head and neck tumor samples and adjacent normal-tissue control samples. Reverse transcriptase polymerase chain reaction (RT-PCR) and quantitative polymerase chain reaction (qPCR) were used to assess ERCC2 deregulation at the mRNA level.
Result
Expression analysis showed that the ERCC2 expression level was significantly upregulated (p<0.05) in HNC tissues compared with adjacent normal tissues. Furthermore, the expression pattern of ERCC2 was correlated with the expression pattern of Ki-67 and a significant correlation (r = 0.230, p<0.03) was observed between ERCC2 and Ki-67. Spearman's correlation also showed a significant correlation between ERCC2 expression and tumor stage (r = 0.271, p<0.02) and grade (r = 0.228, p<0.02) of HNC.
Conclusions
Our data suggest that deregulation of ERCC2 in HNC has the potential to predict a more aggressive cancer phenotype and may be considered a possible biomarker for improved diagnosis and prognosis of HNC.
Introduction
Head and neck cancer (HNC) is the sixth most common type of cancer across the world. Approximately more than 600,000 new patients are diagnosed and 350,000 individuals die of this disease worldwide each year (1). It has been identified as a common malignancy in the Pakistani population, accounting for 21% of cancers in men and 11% in women (2, 3). The main risk factors for HNC include tobacco use, alcohol consumption, ionizing radiation, reactive oxygen species, occupational exposure, HPV infection, and genetic susceptibility (4-5-6-7-8-9-10-11-12-13). Since DNA is frequently damaged by these different endogenous and exogenous carcinogens, this genomic instability plays a pivotal role in the development of HNC (14). The damage is routinely removed by different repair pathways, among which the nucleotide excision repair pathway (NER) is an important DNA repair pathway that removes DNA adducts induced by mutagens and carcinogens in a number of sequential steps (15, 16).
The excision repair cross-complementation group 2 (ERCC2) gene, also called xeroderma pigmentosum group D (XPD), is located on chromosome 19q13.3 and comprises 23 exons; it is one of the main components of the NER pathway (17). It is an ATP-dependent helicase and an essential part of the transcription repair factor complex TFIIH (18). This gene encodes a protein which contributes to 3 critical cellular mechanisms: (i) repair of damaged DNA in the NER pathway, (ii) general transcription, and (iii) cell cycle regulation through its interaction with cyclin-activating kinase (CAK), which is a key activator of cyclin-dependent kinases (19). In the NER pathway, ERCC2 removes some DNA cross-links, UV lesions, bulky chemical adducts, and thymidine dimers (20). The helicase activity of ERCC2 unwinds DNA around the damaged site in the 5’-3’ direction so as to allow the damage-specific nucleases to cleave the DNA on both sides of the damage (21).
Genetic variations in DNA repair genes such as ERCC2 affect the DNA repair capacity. Polymorphisms of the ERCC2 gene are thought to be associated with a high risk of various cancer types such as colorectal cancer, gastric adenocarcinoma, lung carcinoma, urinary bladder carcinoma, skin melanoma, breast cancer and HNC (22). ERCC2 polymorphisms may lead to reduced helicase activity and ultimately DNA repair capacity, which can be important in the carcinogenesis and development of HNC (23). Three polymorphisms of ERCC2 have been extensively studied worldwide. These include C→A, leading to Arg156Arg in exon 6, G→A, leading to Asp312Asn in exon 10, and A→C, leading to Lys751Gln in exon 23 (24).
With regard to expression deregulation of the ERCC2 gene, a limited number of studies on the role of its expression in different cancers are available. We have therefore performed quantitative reverse-transcriptase PCR to determine the relative expression levels of ERCC2. Furthermore, we have correlated the expression of ERCC2 with the expression of the proliferation marker Ki-67.
Materials and methods
Publication Search
To identify the relevant literature, a PubMed search was performed for ERCC2 studies that used human blood and tissue samples obtained from HNC patients along with corresponding control and noncancerous tissues. The following criteria were adopted for the inclusion of studies in this systematic review:
studies on mutation screening and expression analysis of ERCC2 in patients with HNC
studies using blood samples, HNC tumor samples, control blood samples and adjacent noncancerous tissues, respectively, for comparison
studies performed with mutation detection and expression analysis techniques
studies published as full articles in English
Studies using cancer cell lines, serum or saliva samples were not included. Review articles and meta-analyses were also excluded from the present study.
Tumor Sample Collection
Surgical sections of head and neck squamous cell carcinoma (HNSCC) along with adjacent normal tissues (from a microscopically confirmed uninvolved area more than 2 cm away from the tumor section) were collected from 81 HNSCC patients immediately after surgery at the Pakistan Institute of Medical Sciences, Islamabad. Inclusion criteria for patients were: (i) histologically confirmed HNSCC; (ii) no previous chemotherapy, radiotherapy or surgery for the tumor; (iii) no previous history of cancer; and (iv) planned surgical treatment. Tumor and control tissues were stored in RNA at -80°C. Frozen sections of collected tumor and control tissues were analyzed for the presence of tumor cells by a consultant pathologist using hematoxylin and eosin staining. A specifically designed consent form was completed for each patient and signed by the patient. Information about addiction, ethnic group, gender and age was recorded. The study was approved by the ethics committees of the COMSATS Institute of Information Technology and the Pakistan Institute of Medical Sciences. Clinical or pathological staging and identification of the lesions were based on the International Union Against Cancer TNM Classification of Malignant Tumors of 2003.
RNA Extraction and cDNA Synthesis
RNA was isolated from HNC tumor samples and control samples using the standard Trizol reagent method (25). Extracted RNA was analyzed on 1% TAE gel to confirm the isolation of RNA. After isolation, the RNA yield was quantified with a UV spectrophotometer. Absorbance was measured at wavelengths of 260 and 280 nm. Extracted RNA was stored at -80°C. cDNA was synthesized with the SuperScript III First-Strand Synthesis System (Invitrogen). Single-stranded cDNA was synthesized from purified total RNA and stored at 4°C.
Reverse Transcriptase Polymerase Chain Reaction (RT-PCR)
RT-PCR was used to analyze RNA expression. RT-PCR was carried out to assess the expression variations of the ERCC2 gene in HNC tumor samples. The reaction conditions optimized for this reaction were as follows: initial denaturation for 2 minutes at 95°C and 35 cycles of 94°C for 15 seconds, annealing at 56°C for 15 seconds and 72°C for 30 seconds, and final extension at 72°C for 10 minutes. β-actin was amplified as internal control to check the specificity of the reaction. Amplified products were either stored at 4°C or electrophoresed on a 2% agarose gel after completion of the PCR reaction. Their densities were analyzed using the Quantity One software (Bio-Rad Laboratories).
Quantitative Polymerase Chain Reaction (qPCR)
For qPCR, primers specific for ERCC2, Ki-67 and the reference gene β-actin were obtained from IDT DNA Technology. Each qPCR was performed in a 10-µL reaction mixture containing 1 µL cDNA, 1 µL of each forward and reverse primer, 5 µL SYBR Green Master Mix, and 2 µL RNase-free water. qPCR was performed using the StepOnePlus PCR system (Applied Biosystems) at 56°C under standard conditions. The relative mRNA expression of ERCC2 and Ki-67 was computed using the 2-delta delta Ct analysis method with β-actin as the reference gene.
Statistical Analysis
One-way ANOVA, Tukey's post hoc test, Bartlett's test and Student's t-test were used to assess the association of ERCC2 and Ki-67 expression with clinical and histopathological parameters (e.g., TNM stage and grade). Spearman's correlation coefficient was used to assess correlations among gene-gene expression and gene-clinical and histopathological parameters. Statistical analysis was performed using the GraphPad Prism 5 software and SPSS software package.
Results
Study Characteristics
A total of 197 potentially relevant studies were retrieved after a comprehensive search of the PubMed database (Fig. 1), and 124 of these studies were excluded for being not relevant to HNC or ERCC2. A further 45 studies including 18 meta-analyses, 5 reviews, 10 studies carried out on cell lines, 4 studies with no control data, 6 studies with missing data, and 2 studies in languages other than English were excluded. Consequently, 26 studies on the association of ERCC2 polymorphisms with the risk of HNC and only 2 studies on expression analysis of the gene in HNC were included in the systematic review, as shown in Figure 1.
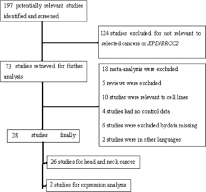
Flowchart of the process used for selection of eligible studies.
The characteristics of the 28 eligible studies, including year of publication, ethnicity of studied population, type of cancer, method of genotyping, and sample size for ERCC2 are listed in Table I. All studies were published between 2000 and 2014.
Literature search in PubMed database for eligible studies reporting on the ERCC2 gene in head and neck carcinoma
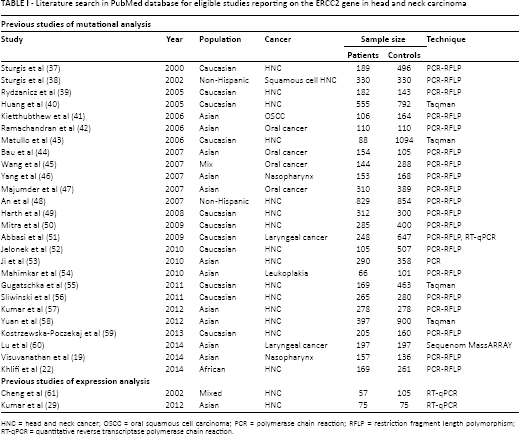
HNC = head and neck cancer; OSCC = oral squamous cell carcinoma; PCR = polymerase chain reaction; RFLP = restriction fragment length polymorphism; RT-qPCR = quantitative reverse transcriptase polymerase chain reaction.
Expression of ERCC2 measured by RT-PCR
In the present study cohort, the expression profiles of ERCC2 were analyzed using RT-PCR in 81 HNC samples and compared with adjacent noncancerous tissues. The demographic characteristics of the study cohort are summarized in Table II. In most tumor tissue samples, ERCC2 expression was upregulated (p<0.04) compared with noncancerous tissues (Fig. 2). β-actin expression was used as internal control in this experiment to show that an equal amount of cDNA was used in all samples (Fig. 2). Uniform expression of β-actin indicates an equal concentration of RNA and valid RT-PCR.
Demographic and clinical parameters of the study cohort
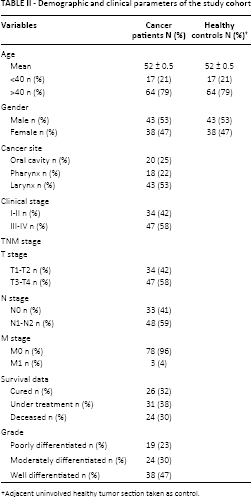

Adjacent uninvolved healthy tumor section taken as control.
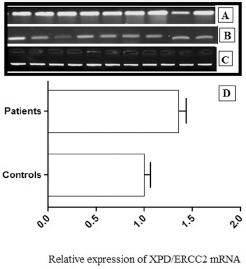
Expression pattern of ERCC2 gene in tumor tissue samples. ERCC2 expression in HNC tumor tissues (
Expression of XPD/ERCC2 measured by qPCR
In order to further confirm the expression patterns of the ERCC2 gene in the same study cohort, qPCR was used. Significant upregulation of ERCC2 was observed in HNC tumor tissues compared with adjacent noncancerous tissue samples (Fig. 3). A statistically significant increase in ERCC2 mRNA levels was observed in larger tumors (T3-T4) compared with smaller tumors (T1-T2). With regard to other parameters, increased expression of ERCC2 was observed in tumors with more advanced clinical stage, lymph node involvement and metastases compared with early-stage tumors, but this increase was not statistically significant (Fig. 3). Significantly increased ERCC2 expression was observed in poorly to moderately differentiated tumors compared with well differentiated tumors. Moreover, significant upregulation was observed in HNC patients still receiving cancer treatment (chemotherapy and radiotherapy) compared to cured and deceased HNC patients. With regard to tumor site, significantly increased ERCC2 mRNA levels were observed in patients with oral carcinoma compared to patients with laryngeal carcinoma and pharyngeal carcinoma (Fig. 4).
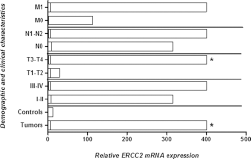
mRNA expression of ERCC2 in HNC tumor samples. Column plot comparing the ERCC2 mRNA levels of HNC tumor and normal control samples, HNC tumor samples at clinical stage I-II and clinical stage III-IV, HNC tumor samples at different pT stages, HNC tumor samples with lymph node involvement (N1-N2) and without lymph node involvement (N0), and HNC tumor samples with metastasis (M1) and without metastasis (M0) *p<0.05.
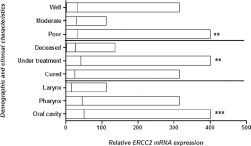
mRNA expression of ERCC2 in HNC tumor samples. Column plot comparing the ERCC2 mRNA levels of HNC tumor samples in different areas, in different grades of HNC tumors, and with survival data. The p values were computed using one-way ANOVA, Tukey's post hoc test and chi-square test
Expression of Ki-67 measured by qPCR
Expression analysis of the proliferation marker Ki-67 was also carried out by qPCR in 81 HNC tissues and compared with adjacent uninvolved normal tissue used as control. Highly significant (p<0.008) upregulation of the Ki-67 gene was observed in tumor tissues compared with adjacent normal tissues (Fig. 5). Upregulated expression of Ki-67 was observed in advanced clinical stage (III-IV), tumor stage (T3-T4), lymph node status (N1-N2) and metastatic stage (M1-M2) and in poorly differentiated tumors, but this upregulation was statistically nonsignificant (Fig. 5). Highly significant (p<0.006) upregulation of Ki-67 was observed in deceased patients compared with cured patients and those still under treatment for HNC (Fig. 6). Relative expression of the Ki-67 gene was compared in different areas of the head and neck using ANOVA. Significant upregulation (p<0.001) was observed tumors of the oral cavity and pharynx compared with tumors of the larynx (Fig. 6).
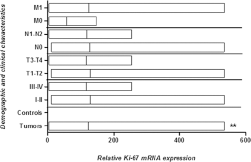
mRNA expression of Ki-67 in HNC tumor samples. Column plot comparing the Ki-67 mRNA levels of HNC tumor and normal control samples, HNC tumor samples at clinical stage I-II and clinical stage III-IV, at different pT stages, with lymph node involvement (N1-N2) and without lymph node involvement (N0), and with metastasis (M1) and without metastasis (M0) *p<0.05; **p<0.01.
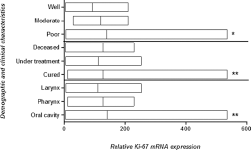
mRNA expression of Ki-67 in HNC tumor samples. Column plot comparing the Ki-67 mRNA levels of HNC tumor samples in different areas, with different grades of HNC tumor samples, and with survival data. The p values were computed using one-way ANOVA, Tukey's post hoc test and chi-square test *p<0.05; **p<0.01.
Correlations between different Clinicopathological Characteristics
With regard to the relationship between different clinicopathological characteristics, a positive correlation was observed between gender versus area (r = 0.312**, p<0.005), pT versus pC (r = 0.930**, p<0.001), pT versus pN (r = 0.990**, p<0.001), pT versus grade (r = 0.231*, p<0.03), pC versus pN (r = 0.920**, p<0.001), and pC versus grade (r = 0.227*, p<0.04). A negative correlation was observed between age versus pT (r = -0.342**, p<0.002), age versus pC (r = -0.323**, p<0.003), age versus pN (r = -0.317**, p<0.004), and age versus area (r = -0.307**, p<0.005) (Tab. III).
Correlations between NER pathway gene (ERCC2), proliferation marker (Ki-67) expression and clinicopathological characteristics of primary HNC †
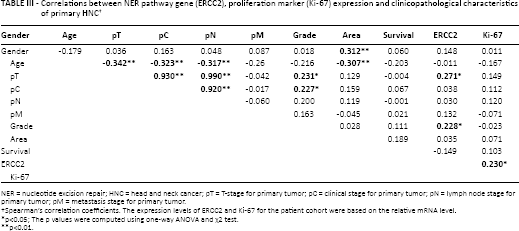
NER = nucleotide excision repair; HNC = head and neck cancer; pT = T-stage for primary tumor; pC = clinical stage for primary tumor; pN = lymph node stage for primary tumor; pM = metastasis stage for primary tumor.
Spearman's correlation coefficients. The expression levels of ERCC2 and Ki-67 for the patient cohort were based on the relative mRNA level.
p<0.05; The p values were computed using one-way ANOVA and χ2 test.
p<0.01.
With regard to correlations between histopathological parameters and gene expression, a positive correlation was observed between pT versus ERCC2 (r = 0.271*, p<0.02) and grade versus ERCC2 (r = 0.228*, p<0.02). In case of gene-gene interaction, a positive correlation was observed between ERCC2 and Ki-67 (r = 0.230*, p<0.03) (Tab. III).
From this correlation study it is important to note that in HNC patient groups positive correlations were consistently observed between ERCC2 expression, T stage and tumor grade. As illustrated in Table IV, statistical analysis revealed that the ERCC2 expression level in HNC patients was associated with T stage and tumor grade (p<0.0007 and p<0.0003, respectively).
Association of increased ERCC2 expression in HNC with tumor size and grade
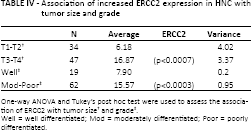
One-way ANOVA and Tukey's post hoc test were used to assess the association of ERCC2 with tumor size
Well = well differentiated; Mod = moderately differentiated; Poor = poorly differentiated.
Discussion
Genetic variations in different genes involved in repair mechanisms may contribute to an increased incidence of different cancers including HNC. Among these mechanisms, the NER pathway is an important DNA repair pathway which removes DNA adducts induced by mutagens and carcinogens in a number of sequential steps (15, 16). DNA adducts are mainly formed by exposure to UV light, ionizing radiation, drugs and reactive oxygen species (26). The ERCC2 gene is an important member of the NER pathway and helps to remove DNA adducts. In many earlier studies, differential expression of ERCC2 has been reported in various cancer types. However, the results have been inconclusive and further studies are warranted to elucidate the role of ERCC2 expression in carcinogenesis. In this context we conducted a case-control study to analyze the expression patterns of ERCC2 in HNC patients of Pakistani origin.
We first carried out a systematic review, which showed that to date most of the studies (n = 26) on the role of ERCC2 in HNC have focused on mutation profiling of XPD/ERCC2 in HNC and little data is available regarding variations in the expression of ERCC2 in HNC: only 2 studies analyzed ERCC2 expression levels in HNC. We therefore designed a case-ontrol study to (i) evaluate the role of the selected DNA repair pathway gene ERCC2 in HNC tumor tissues, and (ii) further correlate the expression levels of the gene with different histopathological parameters of the study cohort.
In the first step, we analyzed the ERCC2 expression levels in tumor samples of 81 HNC patients along with adjacent normal control tissue samples using RT-PCR. An upregulated level of ERCC2 was observed in HNC tumor tissues compared with adjacent controls tissues. In the second step, we confirmed the results by assessing the ERCC2 expression profiles using qPCR in the same study cohort. Statistically significant upregulation was observed in HNC samples compared with control tissue samples. A similar trend in expression had earlier been observed in 21 HNSCC patients by qPCR analysis (27). However, qPCR-based expression analysis of ERCC2 has also found significant downregulation of the gene in HNSCC (75 cases) and esophageal squamous cell carcinoma (100 cases) compared with the respective controls (28, 29). The difference between the expression patterns of patients and controls suggests genetic heterogeneity in gene expression, which might be the result of different genetic variations, epigenetic characteristics, or environmental factors. Another factor that could generate variations in the expression levels of biomarkers in HNC is that HNC is a complex disease involving different anatomical sites (30). In this study, the ERCC2 gene was found to be upregulated further in more advanced tumor stages and higher tumor grades of HNC. A similar pattern of positive correlation between gene expression and T staging has earlier been observed in other DNA repair genes (APEX1 and OGG1) in HNC patients (31, 32). This study confirmed that deregulation of the DNA repair gene ERCC2 in HNC resulted in higher tumor aggressiveness.
In the third step, we assessed the expression levels of the proliferation marker Ki-67 in our study cohort and observed significant upregulation in HNC tumor tissues compared with adjacent control tissues. A similar pattern of Ki-67 overexpression has been reported in different cancers (33, 34). Jiang et al (35) reported that Ki-67 overexpression is related to increased tumor proliferation, recurrence and metastasis. As regards the association between Ki-67 expression and histopathological parameters in the study cohort, upregulation of Ki-67 was observed in more advanced stages and grades of HNC. Increased Ki-67 expression is indicative of more aggressive tumors (31).
The expression levels of ERCC2 and Ki-67 were correlated using Spearman's correlation coefficient. A significant positive correlation was observed between ERCC2 and Ki-67. In the present study, Ki-67 upregulation combined with deregulation of the DNA repair gene ERCC2 appeared to be linked to excessive proliferation in HNC tumor samples, resulting in more aggressive tumors and consequently an increased level of carcinogenesis. Similar results have been reported in studies where Ki-67 expression was correlated with other genes such as p53 (36), XRCC1 and OGG1 (31), and APEX1 (32).
Based on these results, we can conclude that ERCC2 is overexpressed in HNC tumor tissue compared with adjacent control tissue and that the upregulation is associated with an aggressive disease phenotype. Our data also suggest that deregulation of the DNA repair pathway gene combined with overexpression of the proliferation marker Ki-67 may illuminate the association of the DNA repair pathway with the proliferation pathway.
Although this is not first report in relation to the global context, Pakistani populations have not been analyzed for such genetic variations and their association with different types of cancers. Nevertheless, our sample size (81 tumor tissues and adjacent control tissues) is comparatively small. Further, validation studies with larger sample size and additional stratification to control for potential confounding factors such as smoking status and DNA adduct level are required to validate our findings.
In conclusion, our study has demonstrated a significant association between the ERCC2 expression pattern and HNC at the mRNA level in the Pakistani population. Our study also confirms that ERCC2 expression levels are significantly increased in HNC compared with noncancerous healthy tissues. The ERCC2 expression level is further enhanced in more aggressive HNC tumors. In line with previous findings, inhibition of DNA repair activities of ERCC2 due to expression deregulation or any inhibitors could potentially sensitize the tumor cells to therapeutic agents, making ERCC2 an attractive molecular target in the treatment of cancer.
Footnotes
Acknowledgment
All authors wish to acknowledge the patients who contributed to this research work. We are also thankful to the hospital staff of the Nuclear Oncology and Radiotherapy Institute Islamabad (NORI) and Military Hospital Rawalpindi (MH), Pakistan, for their help and cooperation.
Financial support: None.
Conflict of interest: All authors declare that they have no conflict of interests.
