Abstract
Purpose
This study aimed to determine the prognostic value of mutations in the tumor suppressor gene FBXW7 for clinical outcomes in colorectal cancer (CRC).
Methods
Between January 2000 and December 2009, FBXW7 mutations in tumor tissues from 1,519 CRC patients at Taipei Veterans General Hospital were assessed using a MassArray system. We compared the clinicopathological variables and prognosis between the wild-type and mutant tumor tissue groups.
Results
FBXW7 mutations were present in 114/1,519 CRC patients (7.5%). In stage I/II CRC patients, mutant FBXW7 was more common than wild-type FBXW7 (62.3% vs. 50.8%). CRC patients with FBXW7 mutations did not differ significantly in their 5-year overall survival (OS). Stage I/II CRC patients with FBXW7 mutations had lower OS, but this difference was not significant (71.6% vs. 78.2%). Among FBXW7 tumors, S582L was the most frequent mutation type (19.3%), followed by R465H (16.6%), R505C (14.9%) and R479Q (14.9%). Subgroup analysis of FBXW7 mutants showed that R465H/R465C/R479Q had better 5-year OS than other mutant types (76.9% vs. 56.0%; p=0.012).
Conclusions
There was no strong association between patient prognosis and FBXW7 mutations in our large-scale study.
Introduction
Colorectal cancer (CRC) is the most common cancer and the third leading cause of death in Taiwan. Currently, >10,000 new CRC cases are diagnosed annually (1). In recent years, advances in molecular biology have improved our understanding of the genetic background to CRC development.
FBXW7 (F-box and WD repeat domain-containing 7, also known as hCDC4) is one of the most important human tumor suppressor genes, which undergoes deletions and/or mutations in cancers from a wide range of human tissues, such as breast, colon, endometrium, stomach, lung, ovary, pancreas and prostate (2–3–4). The overall frequency of point mutations in FBXW7 in human cancers is approximately 5%-6% (5, 6), and approximately 10% of human colon cancers involve FBXW7 mutations, causing deregulation of these protein substrates (2, 7). FBXW7 is the substrate recognition component of an evolutionarily conserved SCF (complex of SKP1, CUL1, and F-box protein)–type ubiquitin ligase complex (8). SCF-FBXW7 acts as a tumor suppressor in several tissues and targets multiple transcriptional activators and protooncogenes for ubiquitin-mediated degradation (2). It is part of an ubiquitin ligase complex that negatively regulates proteins involved in cell growth, metabolism and differentiation, including c-Jun (9, 10), cyclin E (11, 12), c-Myc (2, 13), different members of the Notch family (14, 15), Aurora-A (16–17–18), Kruppel-like factor 5 (KLF5) (19, 20), myeloid cell leukemia-1 (MCL-1) (21, 22), sterol regulatory element-binding proteins (SREBP) (23–24–25) and mammalian target of rapamycin (mTOR) (26, 27).
FBXW7 has not been previously analyzed as a prognostic marker. In a VICTOR clinical trial, FBXW7 was not predictive of prognosis on either univariate or multivariate analysis with microsatellite instability (MSI) (5, 6). This study evaluated the prognostic role of the MSI status, specific FBXW7 mutations and several clinicopathological features in a hospital-based CRC population to facilitate better stratification of CRC patients.
Materials and Methods
Patients and clinical data
Between March 2000 and July 2009, 1,519 CRC patients who underwent resection were enrolled in this study. Written informed consent for tissue collection was obtained from all patients. Details of the patients’ follow-ups were obtained from the Department of Oncology Medicine, Taipei Veterans General Hospital.
During sample collection, tumors were meticulously dissected, and samples were collected from different tumor quadrants. The tissue fragments were immediately frozen in liquid nitrogen and stored in the Biobank of Taipei Veterans General Hospital for further analysis. Patients who met the following criteria were excluded: familial adenomatous polyposis, personal history of inflammatory bowel disease, pathological diagnosis other than adenocarcinoma, death within 30 postoperative days or emergent operation for obstruction or perforation. The primary end point was overall survival (OS). The pathological staging of cancer was assessed after surgery according to the American Joint Committee on Cancer tumor-node-metastasis grading system (AJCC TNM) (28, 29). All clinical findings were prospectively recorded in detail and stored in digital formats. The surgeries were performed by specialized colorectal surgeons, and pathological reports were provided by a specialized gastrointestinal pathologist, according to the consensus statement of the College of American Pathologists for prognostic factors in CRC – i.e., differentiation grade, mucin component and presence of lymphovascular invasion (29).
Several surveillance procedures were performed before surgery, including colonoscopy, computed tomography (CT) of the area from the neck to the pelvis, and whole body bone scans if patients had symptomatic bone pain or high carcinoembryonic antigen (CEA) levels. Serum CEA levels were measured by one laboratory using an immunoassay, with a recommended upper normal limit of 5 ng/mL. After surgery, patients were monitored every 3 months for the first 2 years and every 6 months thereafter. At each visit, imaging studies, including chest radiography and either abdominal ultrasonography or abdominopelvic CT, were performed. Colonoscopy was performed 6 months to 1 year after surgery and every 3 years thereafter. Unscheduled CT, whole body bone scans or positron emission tomography were performed for patients with increased serum CEA levels and for symptomatic patients.
Proximal colon tumors were tumors located in the cecum, ascending colon, hepatic flexure and transverse colon. Distal colon tumors were tumors of the splenic flexure, descending colon, and from the sigmoid colon to recto-sigmoid junction. The last date of follow-up in this study was December 2013 or until patient death. OS was measured from the definite surgery date until death or last follow-up date.
Tissue samples and mutation analysis
The study was approved by the Institutional Review Board of Taipei Veterans General Hospital (2014-03-008CC), and the tissues used in this study were obtained from the Biobank of the same hospital. DNA extraction was performed using a QIAamp DNA Tissue Kit and a Minelute Virus Kit (Qiagen, Valencia, CA, USA), according to the manufacturer's instructions. The DNA quality and quantity were confirmed using a Nanodrop 1000 Spectrophotometer (Thermo Scientific).
MSI analysis
MSI detection was performed as previously described (30, 31). Five reference microsatellite markers were used according to the international criteria for MSI determination (32): D5S345, D2S123, BAT25, BAT26 and D17S250. Samples with instability in >2/5 marker loci were defined as MSI-H, and those with 1 focus with instability or without evidence of instability were MSI-L/MSS. The analysis was performed twice if the results were equivocal.
PCR conditions for MassArray-based mutation characterization
The present study identified 13 mutations in FBXW7 (Supplementary Table I - Target mutations for the MassDetect CRC panel (v1.0). Available online at www.biological-markers.com), where the design was based on a previous study (33). PCR and the extension primers for the mutations were designed using MassArray Assay Design 3.1 (Sequenom, San Diego, CA, USA). DNA from tumors and plasmas was amplified using the MassDetect CRC panel PCR primer pools. Unincorporated nucleotides were inactivated by shrimp alkaline phosphatase, and a single base extension reaction was performed using a mixture of dideoxynucleotides and extension primers that immediately hybridized adjacent to the mutations. PCR was performed in 384-well plates in 5-μL volumes, i.e., 1 pmol of the corresponding primers, 10 ng tumor DNA, and HotStar Reaction Mix (Qiagen). The PCR conditions were 94°C (15 minutes), followed by 40 cycles at 94°C (20 seconds), 56°C (30 seconds) and 72°C (60 seconds), and a final extension at 72°C (3 minutes). During primer extension, each sample was denatured at 94°C, followed by 40 cycles at 94°C (5 seconds), 52°C (5 seconds) and 72°C (5 seconds). The PCR products from multiplexed reactions were spotted onto SpectroCHIP II arrays, and DNA fragments were resolved using a MassArray Analyzer 4 System (Sequenom). Each spectrum was analyzed using Typer 4.0 (Sequenom) to detect mutations. Putative mutations were filtered manually. The coverage rate of MassArray design was 39.43% (321/814) and most of the detected mutants were missense mutations (Supplementary Table I - Target mutations for the MassDetect CRC panel (v1.0). Available online at www.biological-markers.com).
Statistical analysis
The statistical end point of the analyses was OS from the surgery date. The group distributions for each clinicopathological trait were compared using the 2-tailed Fisher's exact and chi-square tests. Numerical values were compared using Student's t-test. Data were expressed as means ± standard deviation. Kaplan-Meier survival curves were constructed, and statistically significant differences in survival were identified by the log-rank test. Statistical analyses were performed using SPSS (version 16.0 for Windows; SPSS, Chicago, IL, USA).
Results
In the study cohort, the mean age was 70.0±11.5 years (range 28.7-107.4). The median follow-up duration of survival was 67.0 months (range 1-163). Among the 1,519 patients, 999 (65.8%) were men and 520 (34.2%) were women. In the cohort, the tumor locations comprised proximal colon (408, 26.9%), distal colon (646, 42.5%) and rectum (465, 30.7%). Staging was as follows: 214 (14.1%) stage I cancers, 571 (37.6%) stage II, 476 (31.3%) stage III and 258 (17.0%) stage IV. Patient demographic data are shown in Table I. FBXW7 mutations were identified in 114 (7.5%) patients, and 159 (10.5%) patients had MSI-H tumors.
Demographic Data For Patients With Colorectal Cancer (N=1,519)
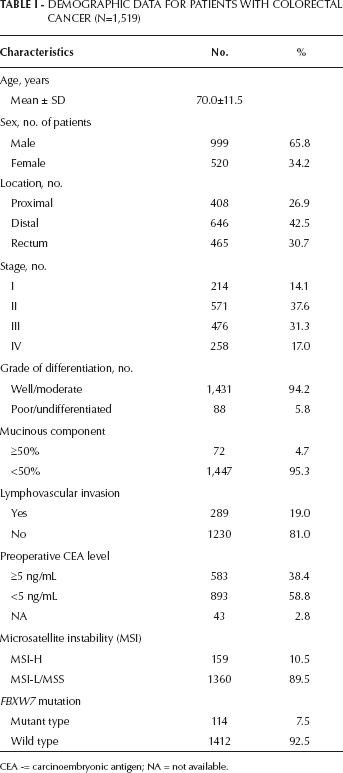
CEA -= carcinoembryonic antigen; NA = not available.
S582L was the most common type of FBXW7 mutation (19.3%), followed by R465H (16.6%), R505C (14.9%) and R479Q (14.9%) (Tab. II). The vast majority of the mutations were single nucleotide changes, primarily missense substitutions. We also found that 4.5% of tumors had double FBXW7 mutations.
Distributions of FBXW7 Mutations (N=114)
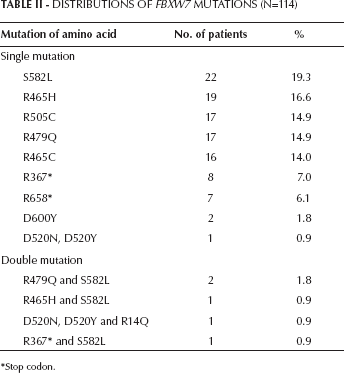
Stop codon.
To elucidate the clinical effect of FBXW7 mutations, we analyzed the relationships between FBXW7 mutations and CRC clinicopathological features (Tab. III). We found that FBXW7 mutant CRCs had significant associations with earlier stages (62.3% vs. 50.8%, p=0.045) and MSI-H tumors (16.7% vs. 10.0%, p=0.025).
Relationships Among FBXW7 Mutation Status and Clinicopathological Features
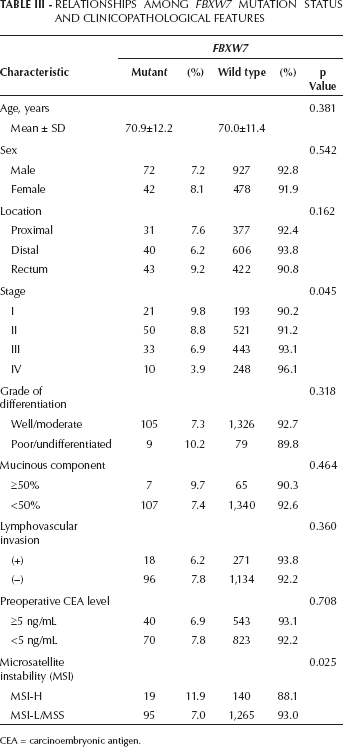
CEA = carcinoembryonic antigen.
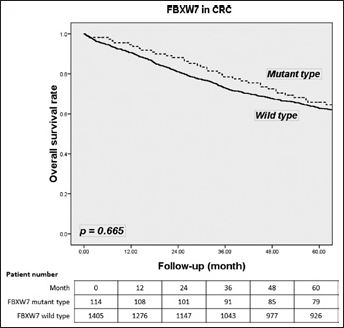
FBXW7 mutations had no effect on survival with CRC tumors (65.8% vs. 62.9%; p=0.665) (Fig. 1). In stage I/II CRC, FBXW7 mutations tended to be related to a lower survival rate than wild-type FBXW7 (71.6% vs. 78.2%; p=0.161) (Fig. 2). In stage III/IV CRC, FBXW7 mutations had no significant association with OS compared with wild-type FBXW7 (56.4% vs. 47.2%; p=0.208) (Fig. 3). Subgroup analysis of mutant FBXW7 revealed that R465H/R465C/R479Q had better 5-year OS than other mutant types (76.9% vs. 56.0%; p=0.012) (Fig. 4). Among the 71 stage I/II CRC patients with FBXW7 mutations, R465H/R465C/R479Q had significantly better OS than FBXW7 wild types and other mutant cases (85.6% vs. 78.2% vs. 60.9%; p=0.017) (Fig. 5). In contrast, there were no significant differences among the stage III/IV CRC patients (65.1% vs. 50.2% vs. 47.1%; p=0.272).
Five-year overall survival (OS) rate for specific FBXW7 mutations.
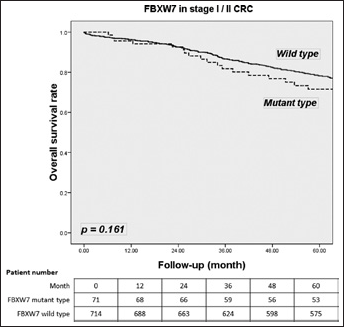
Five-year overall survival (OS) rate according to the FBXW7 mutations in early stages (stage I/II) for colorectal cancer (CRC) patients.
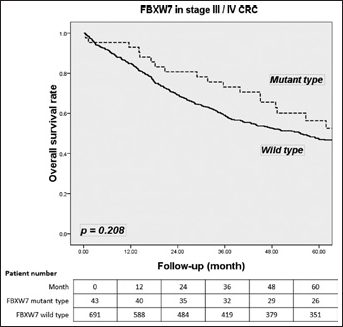
Five-year overall survival (OS) rate according to the FBXW7 mutations in late stages (stage III/IV) for colorectal cancer (CRC) patients.
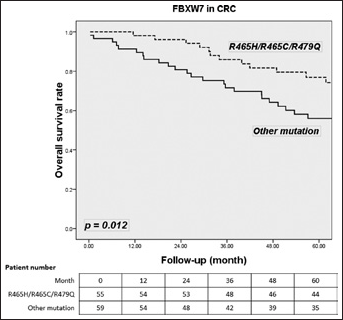
Five-year overall survival (OS) rate for the major hotspots (R465H/R465C/R479Q) and other mutations in 114 colorectal cancer (CRC) patients with FBXW7 mutations.
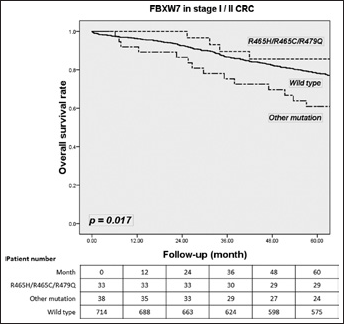
Five-year overall survival (OS) rate for the major hotspots (R465H/R465C/R479Q), other mutations and wild-type FBXW7 in stage I/II colorectal cancer (CRC) patients.
Discussion
To our knowledge, this is the largest study to evaluate the role of FBXW7 mutations in CRC patients. We identified 114 FBXW7 mutations (7.5%) in 1,519 CRC patients. No specific demographic or pathological features characterized the FBXW7-mutated population in our study group. We found that FBXW7 mutations were more common in the earlier tumor stages (stage I/II). No previous studies have reported associations between FBXW7 mutations and CRC stage. Previous studies have detected FBXW7 mutations in approximately 6% of cancer patients (5, 6). A substantial variation in the mutation frequency among tumor types was also described, with the highest mutation rate in cholangiocarcinoma (30%), followed by gastrointestinal tumors (including colorectal cancers) and endometrial and prostate cancers (range 4%-15%) (5, 34–35–36–37).
FBXW7 mutations indicated a poor prognosis in stage I/II CRC patients, but the difference was not significant (p=0.161). However, FBXW7 mutations were not prognostic for stage III/IV CRC patients. These results may indicate that FBXW7 mutations play roles in the prognosis of earlier stage CRC. Previous studies lack sufficient information to allow an adequate assessment of FBXW7 function. UICC/AJCC tumor-node-metastasis (TNM) stage remains the gold standard of prognostic factors in CRC (28). The 5-year survival rate for CRC patients depends primarily on the TNM stage. The TNM staging system has aided in prognosis, treatment choice and patient selection for clinical trials (38). However, even the most up-to-date staging (AJCC, 7th edition staging) (39) cannot address all of the survival discrepancies. Therefore, other prognostic factors may be critical for treatment planning (39, 40). We also found that lymphovascular invasion and elevated preoperative CEA levels were significant prognostic factors for poor OS, corresponding with previous reports (41, 42).
In this study, we found that tumors with FBXW7 mutations were significantly more likely to exhibit MSI than tumors with wild-type FBXW7 (16.7% vs. 10.0%, p=0.025). Previous studies found that FBXW7 mutations were not associated with molecular variables including KRAS, NRAS, BRAF, PIK3CA and TP53 (6). In the VICTOR trial cohort, no specific FBXW7 mutation was predictive of prognosis in either a univariate or a multivariate analysis with MSI and chromosomal instability (6). Analysis of germline mutations of the mismatch repair protein was not included in the present population-based study; therefore, we could not identify a significant relationship between FBXW7 mutations and MSI status. In this study, we did not do MMR sequencing to differentiate familial or sporadic MSI. Further MSI tumors caused either by germline mutation of MMR or MLH1 methylation would have higher mutation rates in tandem repeat sequences. Because there are several repeat sequences in the coding region of FBXW7, it is reasonable that MSI tumors have higher frequencies of FBXW7 mutation (33).
The vast majority of mutations we detected were single nucleotide changes, primarily missense substitutions. The coverage rate of MassArray design was 39.43% (321/814). The difficulty in understanding the impact of FBXW7 mutations involves the complex regulation associated with this protein, with alternate splicing, dimerization and subcellular localization. Unlike previous studies (5), we found that in the 114 patients with FBXW7 mutations, the most frequent specific mutations were of S582L (19.3%), followed by R465H (16.6%), R505C (14.9%) and R479Q (14.9%). The prognostic effects of different FBXW7 mutation genotypes were conflicting. In subgroup analysis of FBXW7 mutants, R465H/R465C/R479Q represented 48.2% of all mutations. These major hotspots had better OS than other mutant types (76.9% vs. 56.0%; p=0.012). Among the 71 stage I/II CRC patients with FBXW7 mutations, R465H/R465C/R479Q had significantly better OS than FBXW7 wild types and other mutant cases (85.6% vs. 78.2% vs. 60.9%; p=0.017). In contrast, there were no significant differences among the 43 stage III/IV CRC patients (p=0.272). Akhoondi et al (5) described 2 major hotspots in nucleotides 1393-1394 (Arg465) and 1435-1436 (Arg479), accounting for approximately 43% of all mutations. FBXW7 mutants that correspond to the major mutational hotspots in primary tumors (Arg465 and Arg479) may act in a negative dominant manner to abrogate the wild-type FBXW7 function. Coexpression of wild-type FBXW7 and FBXW7R465C increased the steady-state cyclin E level and its half-life compared with the expression of wild-type α-FBXW7 alone. The amount of cyclin E protein is tightly controlled and destroyed by FBXW7 (12). FBXW7 can specifically associate with phosphorylated cyclin E and catalyze its ubiquitination (12). Cyclin E is a key cell cycle component that is frequently deregulated in cancer (43). FBXW7 mutations that lead to high cyclin E expression play critical roles in human CRC development.
FBXW7 is a homodimer in cells and the presence of an arginine mutant may render the dimer nonfunctional or unstable, effectively lowering the concentration of active FBXW7 complexes (7). Furthermore, genetic alterations in the FBXW7 locus were found in human CRCs, whereas differential expression of FBXW7 isoforms was inversely correlated with advanced stages and OS in CRC patients. This may explain why CRC patients with FBXW7 mutations did not exhibit significantly lower OS.
FBXW7 is a p53-dependent tumor suppressor gene that undergoes mutations and/or deletions in various human tumors (16). FBXW7 loss alters the spectrum of tumors that develop in p53-deficient mice to include epithelial tumors of the lung, liver and ovary (16). FBXW7's functional significance is related to its role in the binding and targeting of mTOR for ubiquitination and proteosomal degradation (26). FBXW7 depletion leads to increases in mTOR and phosphorylated mTOR, and tumor cell lines with FBXW7 deletions or mutations are particularly sensitive to treatment with rapamycin (26). FBXW7 is found within chromosome band 4q32, a chromosomal region (44, 45) that is mutated in approximately 30% of all human cancers. FBXW7 mutations are localized throughout the gene coding region, including the isoform-specific 5′-exons, nuclear localization signal, F-box motif and WD40 repeats (5). In this hospital-based population study, we found that FBXW7 mutations were significantly associated with MHI-H tumors (p=0.025).
Despite these negative findings, our study had some limitations. First, although we used a large population size, the low incidence of FBXW7 (7.5%) and MSI-H (10.5%) resulted in small prognostic subgroups. Therefore, further validation is needed. Second, the lack of information regarding adjuvant therapy obscured the prognostic analysis; therefore, the chemotherapeutic effect may be negligible. Third, the coverage rate of our MassArray design was only 39.43% (321/814) and most of the detected mutants were missense mutations. Fourth, we did not perform an FBXW7 expression assessment by IHC or qRT-PCR.
Conclusions
In summary, FBXW7 may have no role as a prognostic factor in CRC. Stage I/II CRC patients with FBXW7 mutations had poorer survival rates than those with tumors with FBXW7 wild type, although the difference was not significant. Further studies are needed to verify the results. Limited analyses showed that FBXW7 mutations are inversely correlated with prognosis in solid tumors, including CRC. This is the first large-scale study of the relationship between the outcomes for CRC patients undergoing curative surgery and FBXW7 mutations. There was no strong association between patient prognosis and FBXW7 mutations in our large-scale study. Approximately 10% of CRCs harbor FBXW7 mutations; these large datasets will be required to understand how specific mutations influence the prognosis or therapeutic response.
