Abstract
This study aimed to explore the mechanisms of bridging integrator-3 dysregulation, its prognostic value, and related signaling pathways in colorectal cancer . Bioinformatic analysis was performed based on the data from The Cancer Genome Atlas–colorectal cancer and Human Protein Atlas. Colorectal cancer cell lines, LoVo and HT29 cells, were used as in vitro cell model to assess the effect of demethylation on bridging integrator-3 expression. Results showed that bridging integrator-3 was downregulated in colorectal cancer tissues compared to normal colon and rectum tissues. Heatmap of bridging integrator-3 messenger RNA expression, exon expression, and DNA methylation indicated a negative correlation between bridging integrator-3 expression and methylation of some CpG sites within the coding sequence. Demethylation treatment significantly increased bridging integrator-3 expression in LoVo and HT29 cells. Low bridging integrator-3 messenger RNA and exon expression were associated with significantly worse overall survival (P = .015 and .013, respectively). Multivariate analysis confirmed that low bridging integrator-3 messenger RNA expression was an independent prognostic factor of unfavorable overall survival (Hazard Ratio (HR) = 1.596, 95% confidence interval: 1.024-2.486; P = .039). High bridging integrator-3 DNA methylation was also associated with significantly worse overall survival (P = .013). Kyoto Encyclopedia of Genes and Genomes analysis indicated that the genes correlated with bridging integrator-3 (absolute Pearson r ≥ 0.3, n = 121) were enriched in sphingolipid signaling pathway, natural killer cell-mediated cytotoxicity, p53 signaling pathway, and apoptosis. Based on these findings, we infer that DNA hypermethylation might be an important mechanism of suppressed bridging integrator-3 expression in colorectal cancer . Its low expression is an independent predictor of unfavorable survival in patients with primary colorectal cancer . Bridging integrator-3 might act as a tumor suppressor via modulating natural killer cell-mediated cytotoxicity, p53 signaling pathway, and apoptosis.
Introduction
Colorectal cancer (CRC) is one of the most common cancers in the world. 1 During the past decades, survival of patients with CRC have improved significantly due to early diagnosis and advances in therapeutic strategies. 1 However, a large proportion (about 40%-50%) of patients with CRC still develop local or distant recurrence after surgery. 2 Currently, the American Joint Committee on Cancer (AJCC) tumor, node, metastases classification is the most valuable tool in staging CRC and for making corresponding therapeutic strategies. 3 However, the therapeutic outcomes of patients at similar clinical stages varied significantly. 4 Therefore, it is necessary to explore new predictive markers for individualized therapies.
BIN3 is a gene-encoding bridging integrator-3 in human, which is a member of the Bin, Amphiphysin, Rvs (BAR) domain protein family. 5 This protein has a conserved role in activating cell division control protein 42 (CDC42) in human myocytes and muscle cells, thereby promoting correct cell division. 5,6 This gene has an interesting location at human chromosome 8p21.3, a region that has been implicated widely in cancer suppression. 7 Recent studies suggest that BIN3 is a tumor-suppressive gene in some cancers. Mice with BIN3 knockdown had increased susceptibility to lymphomas and lung cancers. 8 Primary mouse embryonic fibroblasts from BIN3-null mice had stronger potentials in proliferation, invasion, and resistance to anoikis, which are common characteristics of cancerous cells. 8 Another recent study also observed the tumor suppressive effect of BIN3 in breast cancer. 9 Its inactivation can protect the cancer cells from anoikis. 9 These findings suggest that BIN3 might play a tumor suppressor role in multiple cancers.
In this study, we explored BIN3 expression profile and its prognostic value in CRC and studied the possible mechanisms of its dysregulation. In addition, we also investigated the potential signaling pathways it might be involved in.
Materials and Methods
Data Mining Using FireBrowse
Bridging integrator-3 messenger RNA (mRNA) expression in digestive system neoplasms and in the adjacent normal tissues was examined using data from The Cancer Genome Atlas (TCGA). Data analysis was performed by using FireBrowse (http://firebrowse.org/) that provides access to analyze data generated by TCGA.
Data Mining in the Human Protein Atlas
Bridging integrator-3 protein expression in cancer tissues and in the corresponding normal tissues was reviewed by data mining in the Human Protein Atlas (http://www.proteinatlas.org/). 10 The images of BIN3 immunohistochemistry staining in CRC tissues and in normal colon and rectum tissues were also obtained from this database.
Data Mining Using UCSC Xena browser
Heatmap of BIN3 mRNA expression, BIN3 exon expression, and BIN3 DNA methylation in patients with primary CRC (TCGA-CRC) was reviewed using University of California, Santa Cruz (UCSC) Xena browser. Kaplan-Meier plots of overall survival (OS) of the patients were also generated by using UCSC Xena browser. The patients were divided into high and low BIN3 expression groups according to median BIN3 mRNA or median BIN3 exon expression and were divided into high and low DNA methylation groups according to median DNA methylation.
KEGG Analysis
Bridging integrator-3 correlated genes (absolute Pearson r ≥ 0.3) in TCGA-CRC were identified by using cBioPortal for Cancer Genomics. Then, these genes were loaded into ClueGo in Cytoscape 11 for analysis of Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways.
Cell Culture
Human CRC cell lines, LoVo and HT29 cells, were obtained from Shanghai Cell Biology, Institute of the Chinese Academy of Sciences (Shanghai, China). The cells were cultured in Roswell Park Memorial Institute (RPMI)-1640 medium supplemented with 100 units/mL penicillin, 100 μg/mL streptomycin, and 10% fetal bovine serum (Invitrogen, Carlsbad, California).
Demethylation Treatment
Demethylation treatment was performed by using 5-Aza-2’-deoxycytidine (5-AZA-dC; Sigma-Aldrich, St Louis, Missouri). LoVo and HT29 cells were subjected to 1 or 2.5 μM 5-AZA-dC treatment for 24 hours.
Quantitative Real-Time PCR
Total RNA from the cell samples was isolated using the Trizol Reagent (Invitrogen ) and then was reversely transcribed into complementary DNA (cDNA) using the iScript cDNA Synthesis Kit (Bio-Rad, Hercules, California). Then, quantitative real-time polymerase chain reaction (qPCR) was performed to detect gene expression using gene-specific primers and TaqMan Master Mix (Applied Biosystems, Foster City, California), with an ABI 7900HT Fast Real-Time PCR System (Applied Biosystems). Primers for BIN3 were forward, 5′-CTTACTCTCCAATCCCCTCT-3′ and reverse, 5′-TCGATCACAGTCTTCTGGA-3′. The relative mRNA expression was normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH) RNA levels. Relative mRNA expression levels were calculated using the 2−ΔΔCT method.
Statistical Analysis
Statistical analysis was performed by using SPSS 19.0 (SPSS Inc, Chicago, Illinois). Continuous variables were reported as means (standard deviation [SD]). The association between BIN3 RNA expression and the clinicopathological features was evaluated using χ2 tests. Log-rank test was performed to determine the significance of the difference between OS curves. Prognostic values were analyzed by univariate and multivariate Cox regression models. The group difference was compared by 2-tailed Student t test or analysis of variance with Student Newman-Keuls test as a post hoc test. P < .05 was considered statistically significant.
Results
BIN3 is Downregulated in CRC
By retrieving data in TCGA-CRC, we characterized BIN3 mRNA expression in some digestive system neoplasms (Figure 1A). BIN3 expression was slightly lower in CRC tissues than that in normal tissues (fold change: 0.88; Figure 1A). To further characterize its expression at the protein level, data mining was performed in the Human Protein Atlas. Results showed that a majority of breast, prostate, thyroid, and endometrial cancers had moderate granular cytoplasmic staining, whereas the other cancer tissues were negatively or weakly stained (Figure 1B). The normal colon and rectum usually had medium BIN3 staining (Figure 1B and D). In comparison, among 12 cases of CRC tissues, only 3 cases had medium BIN3 staining (Figure 1C and E), whereas 2 cases had weak staining (Figure 1C and F) and 7 cases had no detectable BIN3 expression (Figure 1C and G).
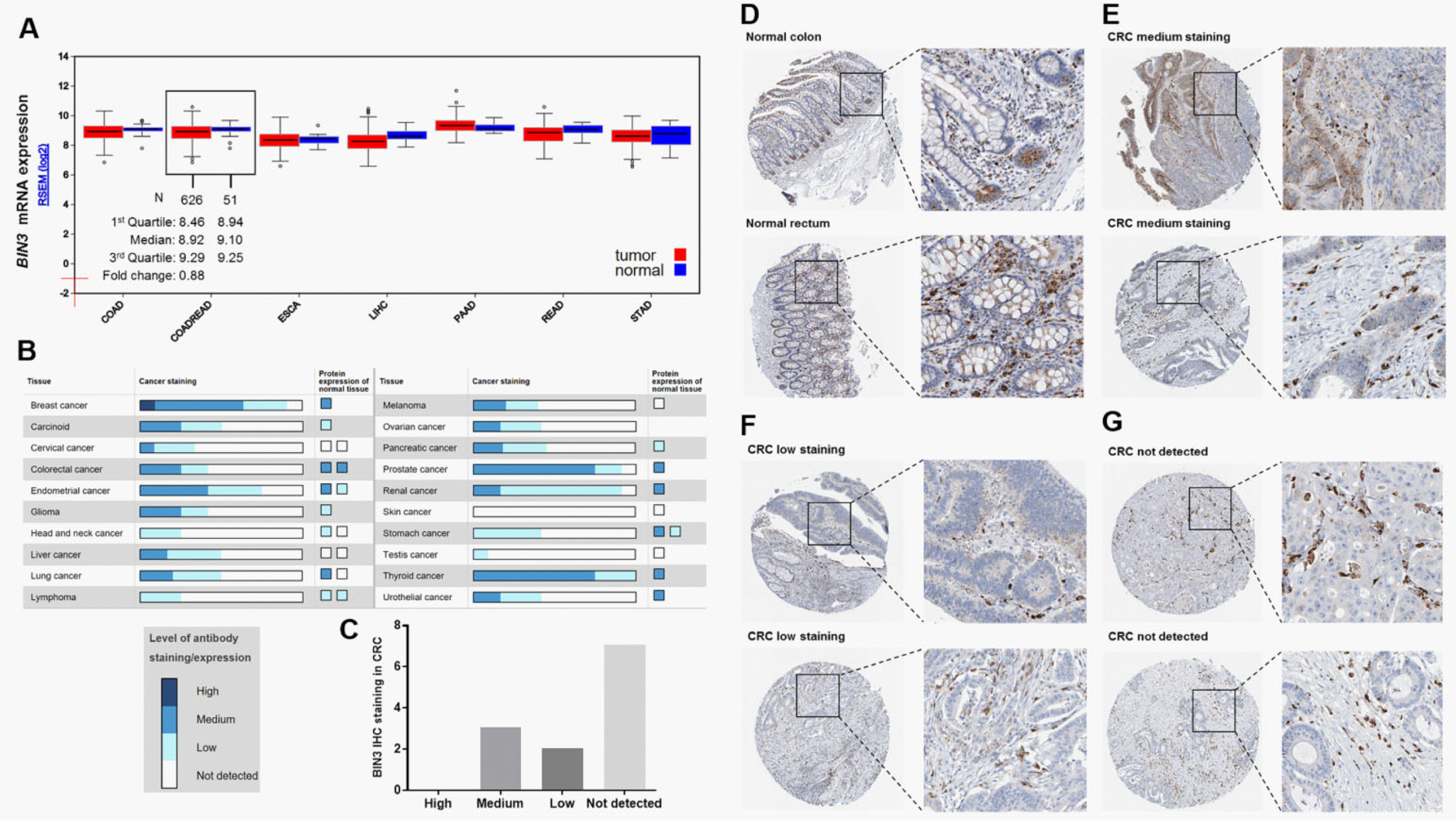
Bridging integrator-3 mRNA and protein expression in CRC. A, BIN3 mRNA expression in some digestive system neoplasms. Box plots were obtained by data mining in FireBrowse. B, BIN3 IHC staining summary in some cancers and in the corresponding normal tissues. Data were obtained from the Human Protein Atlas. C, BIN3 IHC staining summary in 12 cases of CRC. D-G, Representative images of BIN3 IHC staining in normal colon and rectum (D). Medium (E), low (F), and not detectable (G) BIN3 IHC staining in CRC tissues. Images were obtained from the Human Protein Atlas. BIN3 indicates bridging integrator-3; CRC, colorectal cancer; IHC, immunohistochemistry; mRNA, messenger RNA.
BIN3 Downregulation is a Result of DNA Hypermethylation
Then, we further investigated the possible underlying mechanisms of suppressed BIN3 expression in CRC. By data mining in TCGA-CRC (N = 370), we generated a comparable heatmap showing BIN3 mRNA expression, exon expression, and DNA methylation (Figure 2A). In the heatmap, BIN3 mRNA expression was positively correlated with exon expression (Figure 2A). Interestingly, the methylation status of some CpG sites (black frame, Figure 2A) was negatively correlated with the expression of BIN3 mRNA and BIN3 exon. Therefore, we hypothesized that the expression of BIN3 might be suppressed by DNA hypermethylation. To verify this hypothesis, LoVo and HT29 cells were treated with 1 or 2.5 μM 5-AZA-dC. After the treatment, the cells were subjected to qPCR and Western blotting assay of BIN3 expression. Following studies showed that demethylation significantly restored BIN3 expression at both mRNA and protein level in the cells (Figure 2B–D).
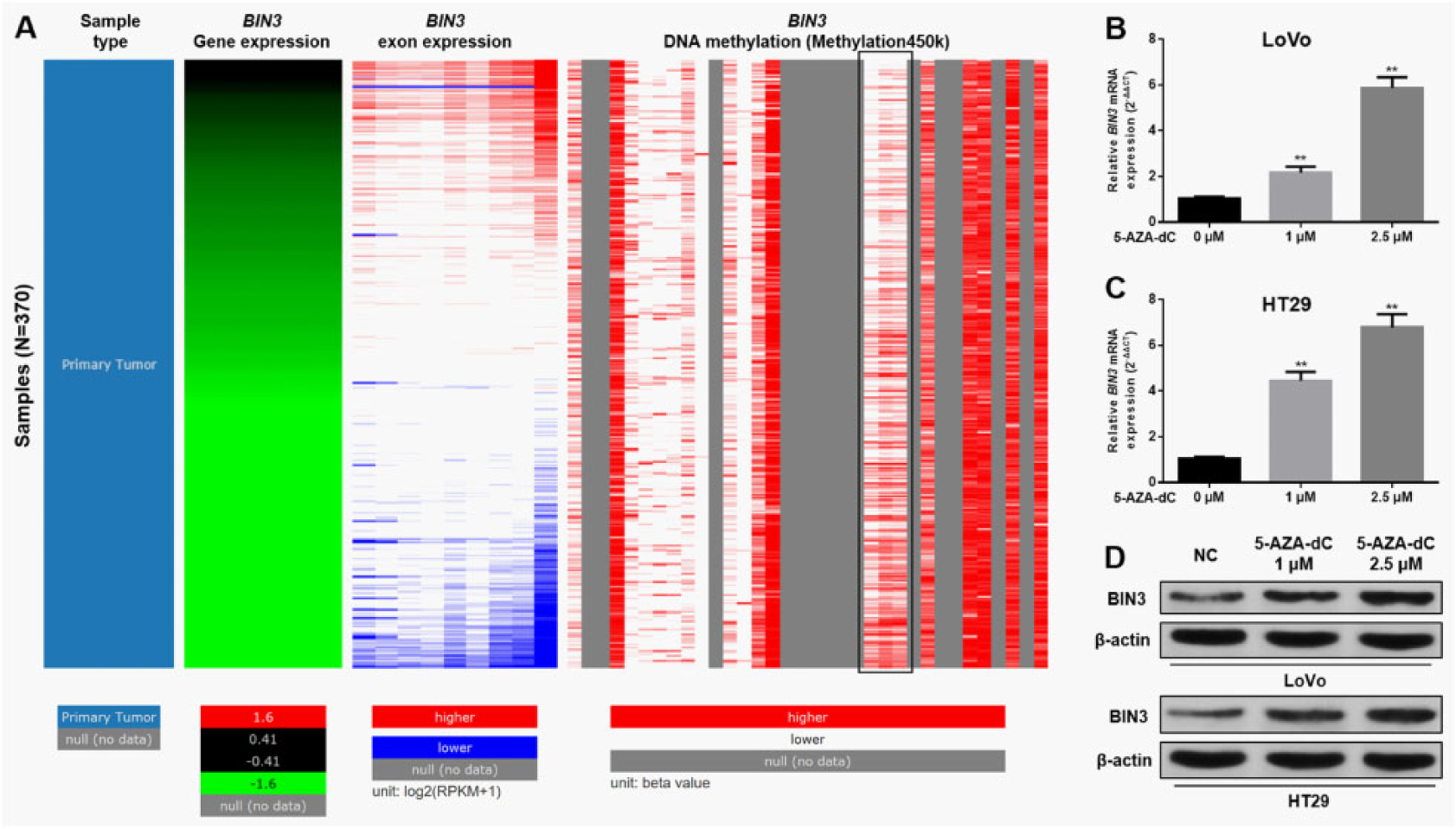
Bridging integrator-3 downregulation is a result of DNA hypermethylation. A, Heatmap of BIN3 mRNA expression, exon expression, and DNA methylation. Only data from patients with primary CRC were used. Analysis was performed using UCSC Xena Browser. B-D, qPCR (B-C) and Western blotting (D) of BIN3 expression in LoVo and HT29 cells 24 hours after 5-AZA-dC treatment. **P < .01. BIN3 indicates bridging integrator-3; CRC, colorectal cancer; mRNA, messenger RNA; qPCR, quantitative real-time polymerase chain reaction; 5-AZA-dC, 5-Aza-2’-deoxycytidine.
Low BIN3 Expression is an Independent Indicator of Unfavorable OS of CRC
The association between demographic/clinicopathological parameters and BIN3 RNA expression was summarized in Table 1. Low BIN3 expression group had a significantly lower ratio of early stages (I/II) patients (81/175, 46.3%) than high BIN3 expression group (107/175, 61.1%; P = .0053; Table 1). Besides, the low BIN3 expression group also had a significantly higher proportion of lymphatic invasion (58/161, 36.0% vs 41/161, 25.5%, P = .04) and death (52/184, 28.3% vs 33/183, 18.0%, P = .02) compared with the high BIN3 expression group (Table 1). To further assess the prognostic value of BIN3, we examined the association between BIN3 expression and OS in patients with CRC. In TCGA-CRC, low BIN3 mRNA expression was associated with significantly worse OS (P = .015, Figure 3A). Since BIN3 is a protein-coding gene, we also assessed the association between BIN3 exon expression and OS in the patients. Results of the log-rank test showed that low BIN3 exon expression was also associated with unfavorable OS (P = .013, Figure 3B). In univariate analysis, older age (>64), late stage (III/IV), venous invasion, lymphatic invasion, and low BIN3 expression were associated with unfavorable OS (Table 2). Multivariate analysis confirmed that low BIN3 mRNA expression was an independent prognostic factor of unfavorable OS (HR = 1.596, 95% confidence interval [CI]: 1.024-2.486, P = .039; Table 2). Besides, age and clinical stage were also independent prognostic factors of OS (Table 2).
The Association Between Demographic/Clinicopathological Parameters and BIN3 Expression in Patients With Primary CRC in TCGA-CRC.
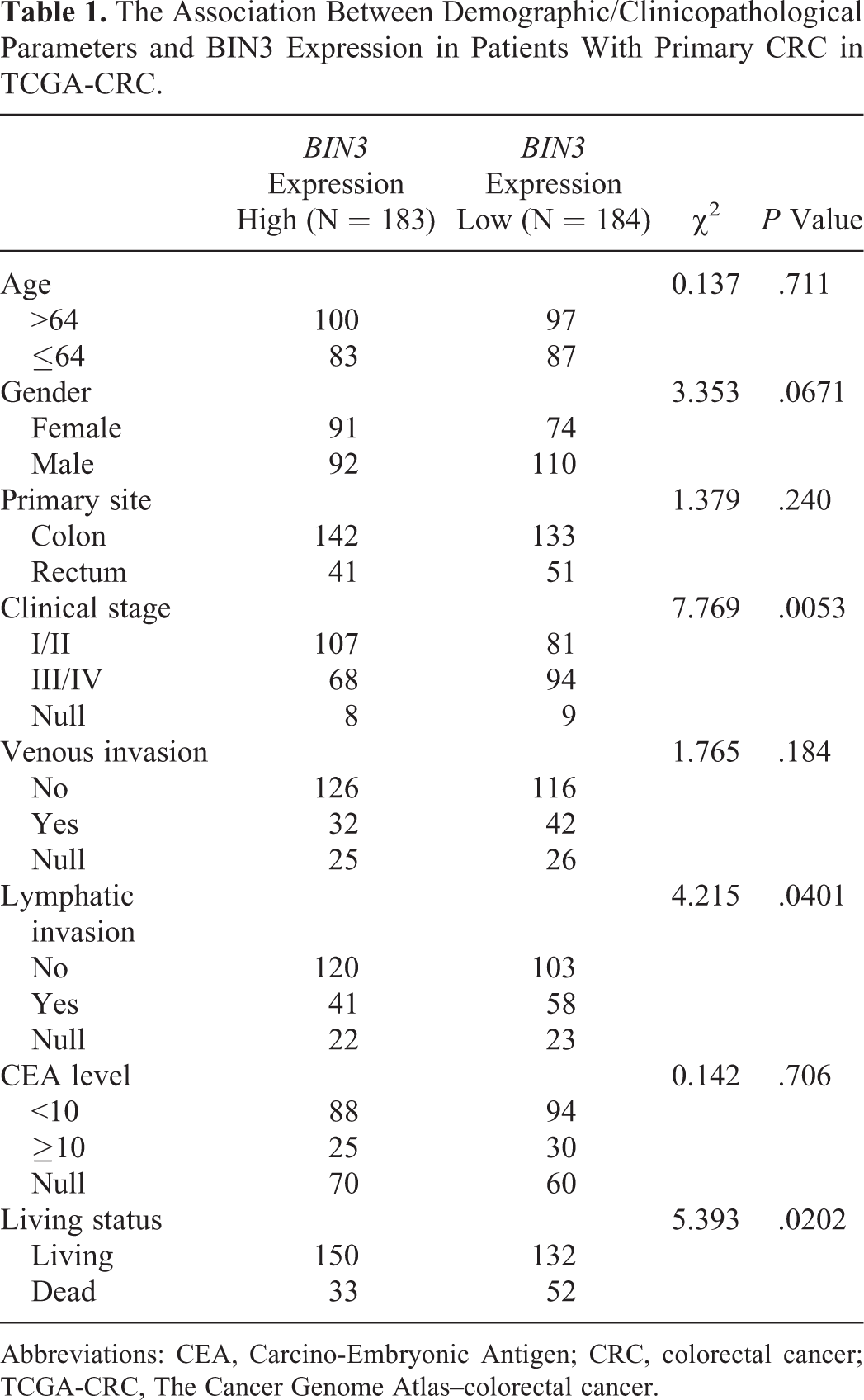
Abbreviations: CEA, Carcino-Embryonic Antigen; CRC, colorectal cancer; TCGA-CRC, The Cancer Genome Atlas–colorectal cancer.

Low BIN3 expression is associated with unfavorable OS of patients with CRC. A-C, Kaplan-Meier plots of OS of patients in TCGA-CRC. Patients were divided into high and low BIN3 expression groups according to median BIN3 mRNA expression (A) or median BIN3 exon expression (B) or were divided into high and low BIN3 DNA methylation groups according to median DNA methylation (C). BIN3 indicates bridging integrator-3; CRC, colorectal cancer; mRNA, messenger RNA; OS, overall survival; TCGA-CRC, The Cancer Genome Atlas–colorectal cancer.
Univariate and Multivariate Analyses of OS in Patients With Primary CRC in TCGA-CRC.

Abbreviations: BIN3 indicates bridging integrator-3; CI, confidence interval; CRC, colorectal cancer; OS, overall survival; TCGA-CRC, The Cancer Genome Atlas–colorectal cancer.
High DNA Methylation Might be an Indicator of Unfavorable OS of CRC
Since we confirmed that BIN3 expression is regulated by DNA methylation, we then examined whether its DNA methylation is associated with OS among the patients. Log-rank test showed that high BIN3 DNA methylation was associated with significantly worse OS (P = .013, Figure 3C).
KEGG Pathway Analysis of BIN3 Correlated Genes in TCGA-CRC
To further investigate the possible functional role of BIN3 in CRC, we identified its correlated genes (absolute Pearson r ≥ 0.3) in TCGA-CRC (n = 121, Supplemental Table 1). Then, the genes were loaded into the online software ClueGo to identify their enrichment in KEGG pathways. Results showed that the genes were enriched in sphingolipid signaling pathway, natural killer cell–mediated cytotoxicity, p53 signaling pathway, and apoptosis (Figure 4 and Supplemental Table 2).

Kyoto Encyclopedia of Genes and Genomes pathway analysis of BIN3 correlated genes in the TCGA-CRC. BIN3 indicates bridging integrator-3; TCGA-CRC, The Cancer Genome Atlas–colorectal cancer.
Discussion
In this study, we firstly characterized BIN3 expression in CRC. Bioinformatic results indicated that BIN3 was slightly downregulated at the mRNA level and was significantly downregulated at the protein level in some CRC cases. Bridging integrator-3 maps to human chromosome 8p21.3. 7 In fact, chromosome 8 (8p) is a host region of many cancer suppressor genes. 12 Its deletion is common in some types of cancers and is one of the most frequent events in cancer progression. 13,14 Besides the chromosomal loss, epigenetic modification, such as DNA hypermethylation, is also an important mechanism of repressed gene expression in this region. For example, secreted frizzled-related protein 1, which maps to 8p11, is a negative regulator of the Wnt pathway. 15 Secreted frizzled-related protein 1 downregulation caused by promoter methylation is common in bladder, colon, breast, cervical, and ovarian cancers. 15 –18 Deleted in liver cancer 1, located at 8p22, encodes a Rho-GTPase activating protein. 19 Deleted in liver cancer 1 can negatively regulate the Rho/ROCK/MLC2 pathway and suppress metastasis and migration of cancer cells. 19 Its promoter methylation was observed in lung, breast, and colon cancers. 19 Cub and Sushi multiple domains 1 (CSMD1) is a tumor suppressor gene in CRC, which maps to 8p23. 20 One recent study found the majority of CRC tumors were methylated at one or more CpG loci within the CSMD1 coding sequence, 20 thereby had downregulated CSMD1 expression. Although we observed suppressed BIN3 expression in some CRC cases, the underlying mechanisms have not been revealed. In this study, by comparing BIN3 mRNA expression, exon expression, and DNA methylation, we observed a negative correlation between BIN3 expression and methylation of some CpG sites within the coding sequence. In addition, by using 5-AZA-dC, a demethylation reagent, we further demonstrated that demethylation increased BIN3 expression in CRC cells. Based on these findings, we infer that DNA hypermethylation might be an important mechanism of suppressed BIN3 expression in CRC.
Loss of BIN3 was correlated with poor prognosis characteristics in patients with breast cancer, such as higher tumor grade, lymph node infiltration, and decreased survival. 9 In this study, we also assessed the prognostic value of BIN3 in patients with CRC. Survival data in TCGA-CRC showed that low BIN3 expression and high BIN3 DNA methylation were associated with unfavorable OS. Moreover, by performing univariate and multivariate analysis, we confirmed that low BIN3 mRNA expression was an independent prognostic factor of unfavorable OS (HR = 1.596, 95% CI: 1.024-2.486, P = .039).
Bin, Amphiphysin, Rvs adapter proteins have been implicated in many cellular processes, such as actin organization, cell polarity, endocytosis, vesicle fusion, and trafficking, specialized membrane organization, stress signaling, transcription, and tumor suppression. 21,22 As an archetypal member of the BAR adapter gene family, the signaling pathways in which BIN3 is involved in cancers have not been fully revealed. In epithelial cancer cells, BIN3 owns the ability to relocate to the cell membrane after cell detachment and to induce a proapoptotic cascade. 9 Mechanistically, via its BAR domain, BIN3 can form a complex with TUBA (the human homolog of gef1p) and localize TUBA to the curved membrane. 9 Once located at the membrane, TUBA activates CDC42 and subsequently leads to the activation of P38α and cell death. 9 Colorectal cancer is also a disease originating from the epithelial cells lining the colon or rectum, but the possible regulative effect of BIN3 in CRC has not yet been reported.
In this study, we identified the genes correlated with BIN3 in CRC and then performed KEGG pathway analysis. Among the identified pathways, natural killer cell–mediated cytotoxicity, p53 signaling pathway, and apoptosis are closely related to tumorigenesis and malignant progression. These findings provide important insights and clues about the functional role of BIN3 and its upstream and/or downstream genes in the pathological development of CRC. Therefore, it is meaningful to further investigate its regulative networks in these pathways in the future.
Conclusion
DNA hypermethylation might be an important mechanism of suppressed BIN3 expression in CRC. Its low expression is an independent predictor of unfavorable survival in patients with primary CRC. Bridging integrator-3 might act as a tumor suppressor via modulating natural killer cell–mediated cytotoxicity, p53 signaling pathway, and apoptosis.
Footnotes
Abbreviations
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplementary material for this article is available online
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
