Abstract
Background
Early detection of colorectal cancer (CRC) is crucial to reducing tumor-related mortality. Evaluating aberrantly methylated DNA in stool is promising for CRC screening. However, DNA methylation in the colonic epithelium of background mucosa may compromise stool DNA (sDNA) test results. Thus, we compared aberrant methylation of cancer-related genes in preoperative and postoperative sDNA, with the aim of demonstrating that a cancer-specific methylated allele in sDNA originates from CRCs.
Methods
Patients who were to undergo CRC resection in Kyushu University Hospital during 2003-2010 were prospectively enrolled. Preoperative (pre) stool samples from 54 patients, postoperative (post) samples from 52 of the patients and tumor samples were collected. Aberrant promoter methylation of CDH4 and GATA5 was assessed in the primary tumors by methylation-specific polymerase chain reaction (MSP) and in stool samples by real-time MSP.
Results
Aberrant methylation of CDH4 and/or GATA5 was detected in 45 of CRC tissue samples (83.3%) and identified in 23 pre sDNA samples (42.3%) from CRC patients. Aberrant methylation was not found in pre sDNA obtained from CRC patients without aberrant methylation of these genes or in post sDNA in any patient. The detection rate of methylated alleles did not correlate with depth of invasion or tumor stage.
Conclusion
Our findings demonstrate that aberrantly methylated alleles identified in sDNA originate from CRCs. Although tumor-specific aberrant methylation is found in sDNA from patients harboring early and advanced CRC throughout the colon and rectum, the sensitivity of this test needs to be improved for early detection of CRC.
Introduction
Colorectal cancer (CRC) is one of the most common malignancies and one of the leading causes of cancer death in Western countries and Japan. Thus, early detection is key in reducing the rate of death from this disease. At present, a fecal occult blood test (FOBT) has been introduced as a screening test for early detection of CRC. However, because of its wide range of sensitivity, the FOBT is considered unacceptable for CRC screening in average-risk adults (1, 2). In addition, this noninvasive test also shows relatively low specificity due to bleeding or leaking blood from surface vessels of mucosa that arises from unrelated non-neoplastic sources (3).
In 1992, Sidransky et al (4) identified the K-RAS oncogene mutation in stool DNA (sDNA) from patients with CRCs and advanced adenomas. Since then, analyzing sDNA has become one of the most promising screening tests for CRC detection. For example, APC mutations in sDNA were found in 57% of patients with CRC and advanced adenoma in 1992 (5). Further, p53 mutations were also identified in sDNA in 64% of patients with CRCs harboring mutated p53, and some of these patients were negative for FOBT (6). Detection of tumor-specific genetic alterations in sDNA offers an advantage over FOBT/fecal immunochemical testing (FIT) in providing qualitative results, because these alterations somatically occur during initiation and promotion of CRCs. Assessing sDNA for gene mutations could discover some but not all CRCs, as the genetic mutations that contribute to CRC are complex, and no single gene mutation occurs in every CRC. Thus, the sensitivity of screening tests for each genetic mutation is comparatively low.
Recently, epigenetic inactivation of tumor suppressor genes, mostly from aberrant methylation of promoter regions, has been identified in various cancers, including CRCs. These methylated segments of DNA are thought to serve as tumor-specific markers in sDNA (7, 8). This aberrant methylation of promoter regions in specific genes consistently affects the same promoter sites, occurs at an early stage of colorectal carcinogenesis and is more frequently recognized than genetic alterations (9). Thus evaluation of methylated sDNA may have the potential to achieve high sensitivity for early detection of CRCs.
It is possible that the mutated or aberrantly methylated DNA in stool could come from colonic polyps apart from CRCs, non-neoplastic epithelium of colorectal mucosa or supracolonic digestive cancers or lesions. Caldas et al (10) detected K-ras mutations in the stool of patients with pancreatic adenocarcinoma together with benign chronic pancreatitis, and found that DNA markers could survive digestive processes during intestinal transport. Because promoter methylation in several genes was occasionally or constantly found in normal colonic mucosa in an age-related manner (11, 12), this background methylation may affect specificity for the tests. This possibility cannot be ruled out by only comparison of sDNA among CRC patients and healthy counterparts without confirming colonoscopy (CS).
Promoter hypermethylation of the GATA5 gene is a common and specific event in colorectal carcinogenesis, and GATA5 also exhibits tumor-suppressive effects in colorectal cancer cells in vitro (13). The CDH4 gene encodes retinal cadherin (R-cadherin) and is frequently silenced by promoter hypermethylation in human colorectal and gastric cancers (14). In our previous research, aberrant methylation of CDH4 and GATA5 genes was detected in 78% and 60% of over 100 cases of CRCs but not in normal colonic mucosa (15). To clarify that sDNA markers occur exclusively in CRCs, we conducted a prospective study to examine promoter methylation of CDH4 and GATA5 in sDNA collected before and after tumor resection from CRC patients. We also compared the results of sDNA testing and location of tumors, depth of tumors and tumor stages.
Patients and Methods
Study population
Fifty-four patients who underwent CRC resection at Kyushu University Hospital from 2003 to 2010 were prospectively recruited to the study. Written informed consent to analyze sample DNA was obtained from all patients, and the Kyushu University Hospital Human Research Ethics Committee approved this study. Of the 54 total patients, 33 were male and 21 were female, and the patients averaged 66.7 years in age, range; 37 to 88. Primary tumor and the corresponding preoperative (pre, n=54) and postoperative (post, n=52) stool samples were collected from patients. Tumor stage and other clinicopathological data were obtained by medical records. To evaluate anastomotic recurrence and metachronous multicentric CRC, CS was performed within 6 months after surgery in patients who did not undergo CS before surgery, and within 12 months for the remaining patients. Forty-six patients (85.2%) underwent CS in our institution before and after surgery, while CS data were not available for analysis in 8 patients who did not undergo CS because of obstruction by tumors and/or social reasons. In this study, advanced polyps, which have been recognized as precancerous lesions for CRC, were defined as polyps equal to or larger than 10 mm. Recurrence was investigated by regular checkup of patients using computed tomography every 6 months for the first 5 years. The mean follow-up period of recurrence-free patients was 51.2 months.
Collection and extraction of primary tumor tissue DNA and sDNA
Genomic DNA was extracted from fresh-frozen CRC samples and formalin-fixed and paraffin-embedded CRC samples using the QIAamp DNA Mini Kit (Qiagen, Hilden Germany). Pre stool samples were collected 1 to 3 days before surgical resection, and post stool samples were obtained 5 to 10 days after resection. The stool samples were immediately preserved at −80°C until analysis, and DNA extraction from stool samples was performed using the QIAamp DNA Stool Kit (Qiagen). Approximately 100 mg of stool samples was subjected to DNA extraction using specific conditions to increase the amount of human DNA obtained compared with nonhuman DNA (such as bacterial or lumen cells), as described in the manufacturer's protocol. DNA was eluted in 50 μL of elution buffer and quantified.
Bisulfite modification and methylation-specific polymerase chain reaction assay
Genomic DNA (1 µg) from tumor and stool samples was treated with the EpiTect Bisulfite Kit (Qiagen) to convert unmethylated cytosines to uracils, with no effect on methylated cytosines. Treated DNA was eluted in 40 μL of elution buffer. The methylation status of each gene in the tumor samples was verified by methylation-specific polymerase chain reaction (MSP) as described by Herman et al (16). The polymerase chain reaction (PCR) primer sequence for MSP was located in the promoter region of each gene. The primer sequences for methylated CDH4 were 5′-GTTTTCGGTGTCGGGTATC-3′ (forward) and 5′-CGACAACTTACCCGAAACG-3′ (reverse), and for methylated GATA5, 5′-TTCGGGTCGTTGTGAGGTTTC-3′ (forward) and 5′-CAAAATCACGTAACTCTACG-3′ (reverse). MSP annealing temperature for CDH4 was 67°C, and for GATA5, 64°C. MSP was carried out on 1 μL of bisulfate-treated DNA with the following amplification conditions: 5 minute at 95°C, followed by 40 cycles for 30 seconds at 95°C, annealing for 30 seconds, and 72°C for 45 seconds at 72°C and a final extension for 5 minutes at 72°C. Because there are no established optimal positive and negative control DNAs for sDNA analysis, CpGenome Universal Methylated DNA (Chemicon International, Temecula, CA, USA) and serum DNA from a healthy volunteer (17) were bisulfate-modified and used as a positive and a negative control, respectively. All PCR reactions were performed with the control samples as well as template-free distilled water control. PCR products (5 μL) were separated by gel electrophoresis in 3% agarose gel, and the gel was stained with ethidium bromide and visualized by UV illumination.
Real-time MSP
Real-time MSP was performed on a 7500 real time PCR system (Applied Biosystems, Foster City, CA, USA), and 1 μL of bisulfate-modified DNA from stool samples was used in each amplification. Template-free distilled water and negative and positive controls were included for each amplification. CpGenome Universal Methylated DNA was also serially diluted to create a standard curve. The PCR parameters were 5 minutes at 95°C, followed by 55 cycles for 30 seconds at 95°C, 30 seconds at annealing temperature and 45 seconds at 72°C. A melting curve was examined to confirm a single product to eliminate nonspecific amplification.
Statistical analysis
All statistical analyses were performed with JMP version 9.0.2 (SAS Institute, Cary, NC, USA). Correlation between clinical and demographic characteristics, including sex, location of tumor, differentiation of tumor, depth of tumor, tumor stage, presence or absence of remnant advanced polyps and methylation status of CDH4 and GATA5 were analyzed with the chi-square test and Fisher's exact test for categorical variables. Relationship between other characteristics, age, size of tumor and methylation status of those genes were evaluated with Student's t-test. All p values are 2-sided, and a p value ≤0.05 was considered statistically significant.
Results
Aberrant methylation of CDH4 and GATA5 in colorectal cancers
CDH4 and GATA5 promoter aberrant methylation was detected in 70.4% (38/54) and 59.3% (32/54) of CRCs, respectively. Altogether, 46 of 54 CRCs (83.3%) had CDH4 and/or GATA5 aberrant methylation, while the remaining 9 CRCs (16.7%) showed no methylation of either gene. The associations between CDH4 and GATA5 aberrant methylation and clinicopathological features of the tumors are shown in Table I. The incidence of CDH4 and/or GATA5 aberrant methylation did not differ regarding age, sex, tumor location, tumor size, depth of tumor invasion or stage. Differentiated CRCs more frequently had aberrant methylation of the genes, compared with undifferentiated CRCs (87.5% vs. 50.0%, respectively; p=0.041; Tab. I). Among 46 patients who underwent CS before and after surgery, advanced polyps in remnant colon and rectum were found in 7 patients (15.2%) and most were removed endoscopically during CS after surgery.
Associations Between Cdh4 And Gata5 Aberrant Methylation In Tumors And Clinicopathological Features Of Crcs
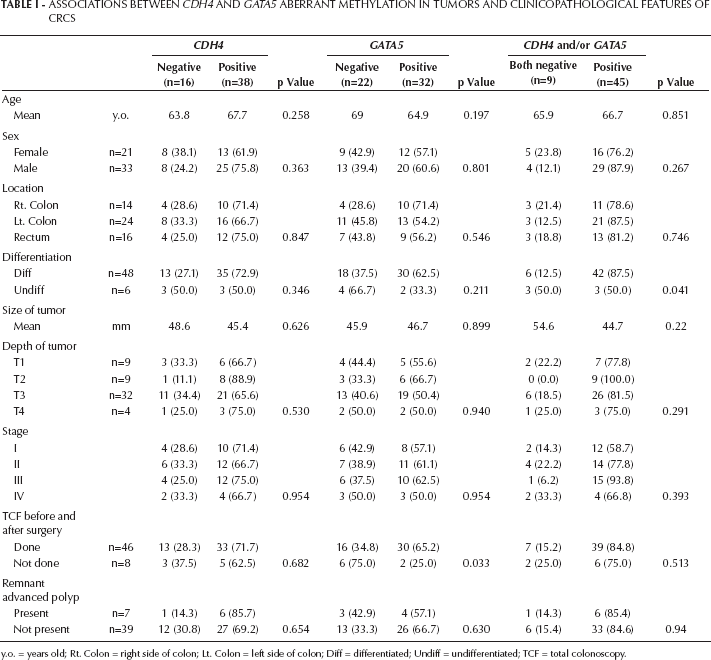
y.o. = years old; Rt. Colon = right side of colon; Lt. Colon = left side of colon; Diff = differentiated; Undiff = undifferentiated; TCF = total colonoscopy.
Methylated allele of CDH4 and GATA5 in preoperative and postoperative sDNA
Real-time MSP revealed the presence of methylated CDH4 and GATA5 allele in 19 (35.2%) and 13 (24.1%) of pre sDNA from CRC patients, respectively. We divided the CRCs into 4 categories according to methylation status in primary tumors, and the results of real-time MSP on pre and post sDNA is summarized in Table II. Methylated CDH4 allele was detected exclusively in pre sDNA from patients who harbored CRCs with aberrant methylation of CDH4. Similarly, methylated GATA5 allele was seen only in pre sDNA from CRCs with aberrant GATA5 methylation. Methylated alleles of CDH4 and/or GATA5 were detected in pre sDNA from 23 of the total 54 CRC patients (42.3%). Among 46 patients having CRCs with CDH4 and/or GATA5 aberrant methylation, 51.1% contained a methylated allele of these genes in pre sDNA, while none of the sDNA from CRC patients without CDH4 or GATA5 aberrant methylation contained a methylated allele (p=0.001). Methylated alleles of these genes were not found in post sDNA regardless of methylation status in primary CRCs (Tab. II; p<0.0001).
Detection Of Methylated Alleles In Preoperative (Pre) And Postoperative (Post) Sdna Samples
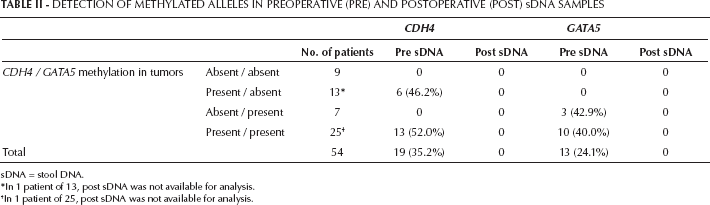
sDNA = stool DNA.
In 1 patient of 13, post sDNA was not available for analysis.
In 1 patient of 25, post sDNA was not available for analysis.
Relation between clinicopathological features of the patients and aberrant methylation in sDNA
To evaluate the characteristics of CRC patients with methylated allele in pre sDNA among 46 patients with CRCs harboring CDH4 and/or GATA5 aberrant methylation, we further analyzed the correlation between the status of methylated alleles in pre sDNA and clinicopathological features of tumors (Tab. III). CRCs having CDH4 and/or GATA5 methylated allele in pre sDNA were relatively larger than those without the methylated allele in pre sDNA, but the difference was not statistically significant (mean: 49.5 mm vs. 39.7 mm, respectively; p=0.11). CDH4 and/or GATA5 methylated allele was almost equally found in pre sDNA from CRCs located in the right colon, left colon and rectum, and the frequency of CDH4 and/or GATA5 methylated allele in pre sDNA did not differ with depth of tumor or tumor stage. Among the 46 total CRC patients, pre and/or post CS was performed in 41 patients and revealed remnant advanced polyp (≥10 mm in size) in 6 patients. The methylated alleles were detected equally in pre sDNA from patients with remnant advanced polyps and those without (50.0% vs. 48.6%, respectively; p=0.693). Within the follow-up period, 13.6% (3 of 19) of stage I-III CRC patients having CDH4 and/or GATA5 methylated allele in pre sDNA developed tumor recurrence, compared with 3.9% (1 of 26) of those without the methylated allele in pre sDNA (p=0.32). None of the patients who underwent curative resection developed other gastrointestinal malignancies during the follow-up period, except for 1 patient with intraepithelial pharyngeal carcinoma at 2 years after surgery.
Correlation Between Gene Methylated Status In Preoperative Sdna And Clinicopathological Features Of Tumors *
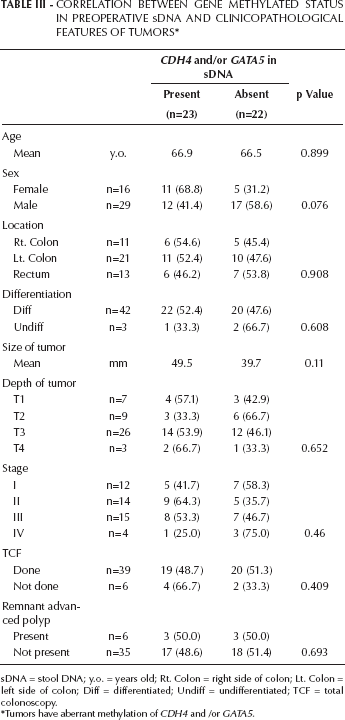
sDNA = stool DNA; y.o. = years old; Rt. Colon = right side of colon; Lt. Colon = left side of colon; Diff = differentiated; Undiff = undifferentiated; TCF = total colonoscopy.
Tumors have aberrant methylation of CDH4 and /or GATA5.
Discussion
In this study, we examined whether aberrant methylated alleles of cancer-related genes in sDNA originated from CRC by comparing the pre and post sDNA from patients who underwent surgical resection. Consistency of the presence of the methylated allele in the primary CRC and pre sDNA and the lack of methylated allele in post sDNA proved that the detected methylated allele in pre sDNA was derived from resected CRCs. Pre and/or post CS demonstrated that 7 CRC patients had advanced polyps. Thus, these polyps might not be the source of the methylated allele in pre sDNA. However, we did not examine the methylation of CDH4 and GATA5 in the polyps. Thus, analysis of post sDNA could mostly eliminate the possibility that these methylated alleles came from additional digestive tract tumors located upstream, polyps in other parts of the colon or morphologically normal mucosa. Although there was little possibility of an inadequate amount or quality of human genomic DNA in post sDNA for real-time MSP compared with pre sDNA in detecting these alterations in residual malignant or premalignant cells, the lack of succeeding malignancy (except for 1 pharyngeal carcinoma) during the follow-up nearly eliminates the coexistence of malignant or premalignant cells prone to becoming detectable cancers in the digestive tract at the time of operation. In contrast, it may be uncertain that the test was truly a false negative, if a test based on detection of molecular markers in sDNA was positive and if the subsequent CS failed to show any evidence of CRC or advanced polyp. This is particularly important in broadly applying the use of methylated genes as CRC markers, because the genes methylated in the majority of CRCs become methylated to some degree in the background and in morphologically normal colorectal epithelium in an age-related manner (12).
Aberrantly methylated genes in sDNA for CRC detection have been assessed by MSP (2, 18) and more recently, methylation levels of methylated genes in sDNA in CRC patients and non-neoplastic patients were evaluated by real-time quantitative MSP (13, 19). The quantitative MSP estimating the area under the receiver operating characteristic (ROC) curve could distinguish the patients with CRC or advanced polyp from noncancer individuals, and was advocated as a novel approach for early CRC detection with applicable sensitivity and specificity (9). In this study, we also employed real-time MSP in the anticipation of high sensitivity to detect aberrantly methylated alleles that comprise a tiny fraction in heterogeneous DNA samples, but we did not assess ROC curves, because the amplified allele was not found in post sDNA. Although previous studies demonstrated promoter methylation of CDH4 (14) and GATA5 (13) in some populations of normal colonic tissues, methylation status of these genes was not identified in post sDNA or pre sDNA from the patients with CRC without methylation of those genes. It could be possible that some of the colonocytes from normal mucosa were shed into the stool to some degree. Thus, differences between our results and those of previous studies might be due to the differences of the selected sequences for MSP primers and the annealing temperatures, because the degree of methylation in each gene could differ across the 5′ promoter regions, and the variation of specificity of analyses is mainly determined by primer sequences and annealing temperature of MSP. We applied relatively higher annealing temperatures to eliminate mis-annealing of the primers and subsequent amplification of bisulfite-treated DNA templates that contained nonmethylated cytosines to some degree instead of complete methylated cytosines throughout the sequences of the primer sites.
Compared with FOBT/FIT, which provides up to a 10% positive predictive value (PPV) and lower specificity for symptom-free patients (20), sDNA tests using designated tumor-specific markers can give a higher PPV to detect cancer or precancerous lesions (advanced polyps). Ahlquist et al demonstrated that an abundance of colonocytes retaining expression of tumor-associated antigens exfoliated into mucus over CRC. This luminal shedding of colonocytes appeared to be much greater from CRC than normal colonic epithelium (21), and the above exfoliation of colonocytes was a continuous process. Thus, detecting tumor-specific DNA markers in these colonocytes could have an advantage over FOBT/FIT, which depends on bleeding or leaking, which usually happens in an intermittent manner (22). Moreover, this shedding of the colonocytes containing specific DNA marker into stool could occur at earlier stages than that into blood or urine (21).
Several drawbacks need to be addressed to apply this method in routine clinical practice. These include the difficulty in the reproducible retrieval of human DNA from huge amounts of nonhuman DNA in the stool and a sufficient quality of DNA to allow bisulfite modification for detection of aberrant methylation, which is only present in a small fraction of total DNA samples. In addition, the established use of comprehensive markers of several individually heterogeneous CRCs and the improvement of sensitivity without decreasing specificity are required. The specificity of our sDNA tests was 100%, but the sensitivity was relatively low, while the employment of SFRP2 methylation for sDNA tests allowed a higher sensitivity of 77% but showed a relatively lower specificity of 77% for CRC detection (8). Finally, compared with FOBT/FIT, sDNA tests are more elaborate and more costly (23).
The detection rate of methylated alleles in pre sDNA did not differ among the patients with CRCs of various depths of invasion and location of the tumor, and there was no significant difference in the size of tumors between CRCs harboring methylated allele in pre sDNA and those without methylated alleles. While FOBT/FIT has shown lower sensitivity for Dukes’ stage A tumors than for Dukes’ stage B/C/D, and less efficiency to detect proximal colon cancers compared with distal colon cancers (3), sDNA tests could be equally sensitive to early CRC and to CRC in the right colon. These advantages are important when the opportunity to perform CS is limited, and it is substituted for by sigmoidoscopy or another less sensitive modality to detect proximal colon cancers or relatively smaller CRCs.
This study had several limitations. First, the number of patients was too small to draw firm conclusions. Second, only the CDH4 and GATA5 marker genes were examined in this study, thus CRCs having no methylation of these promoters, such as undifferentiated CRCs, could not be detected by this sDNA test. Identification of additional markers and/or the combination of markers among many genetic and epigenetic alterations in CRCs is required. Ultimately, our results needs to be verified in larger prospective studies, and the efficacy and total cost of sDNA tests compared with FOBT/FIT, computed tomography colonography and CS should be evaluated.
Footnotes
list of Abbreviations
Acknowledgements
We appreciate the technical support from the Research Support Center, Graduate School of Medical Sciences, Kyushu University.
