Abstract
Background
Cancer screening and early detection greatly increase the chances of successful treatment. However, most cancer types lack effective early screening biomarkers. In recent years, natural language processing (NLP)-based text-mining methods have proven effective in searching the scientific literature and identifying promising associations between potential biomarkers and disease, but unfortunately few are widely used.
Methods
In this study, we used an NLP-enabled text-mining system, MarkerGenie, to identify potential stool bacterial markers for early detection and screening of colorectal cancer. After filtering markers based on text-mining results, we validated bacterial markers using multiplex digital droplet polymerase chain reaction (ddPCR). Classifiers were built based on ddPCR results, and sensitivity, specificity, and area under the curve (AUC) were used to evaluate the performance.
Results
A total of 7 of the 14 bacterial markers showed significantly increased abundance in the stools of colorectal cancer patients. A five-bacteria classifier for colorectal cancer diagnosis was built, and achieved an AUC of 0.852, with a sensitivity of 0.692 and specificity of 0.935. When combined with the fecal immunochemical test (FIT), our classifier achieved an AUC of 0.959 and increased the sensitivity of FIT (0.929 vs. 0.872) at a specificity of 0.900.
Conclusions
Our study provides a valuable case example of the use of NLP-based marker mining for biomarker identification.
Introduction
Screening and early detection of cancer aim to detect malignancy or precursor lesions at an early stage when treatment of cancer is most effective. However, for most cancer types, the effective early screening biomarkers have not been discovered or fully validated.1,2 Moreover, even for the cancer types with many research and commercial products, the existing biomarkers still cannot fully meet the clinical needs.1,2 As a representative example, colorectal cancer (CRC) is a cancer type with numerous studies on screening and early detection, and several biomarker sets have been developed for clinical use.3,4 The most commonly used biomarkers in CRC screening are the fecal occult blood test (FOBT) and fecal immunochemical test (FIT), which detect blood in the stool.3–5 Because of the low sensitivity of guaiac-based FOBT (gFOBT) and antibody-based FIT for early-stage CRC, several stool DNA tests have been developed,5–7 and combining stool cancer DNA detection with FIT achieves a sensitivity of ∼90%.5,8 Meanwhile, several blood-based cancer screening tests are also used for CRC screening.9–14 However, such tests have higher costs or lower performance than stool-based tests for CRC screening. Although existing stool-based CRC screening tests have a positive impact in improving patient outcomes, 3 it is hard to achieve positive predictive values higher than 50%, thus, effective markers are still needed. 15
Generally, there are two ways for effective biomarker screening using publicly available information. The first way is data mining on high throughput omics datasets. However, many markers identified by high-throughput omics technologies are not repeatable in different public datasets, thus such data mining needs to use multiple datasets, which needs to be performed by experienced data scientists. In addition, for many disease types and clinical needs, there is a shortage of public omics datasets and generating new data can be quite costly. Another option is to find supportive evidence for potential biomarkers by text-mining of biomedical literature. However, the manual searching of literature is time intensive and often returns a lot of results supported by only one or two small studies, which makes it difficult to identify useful information. Therefore, although not fully validated, the use of natural language processing (NLP)-based automatic text-mining methods with semantic search and information retrieval systems is a promising tool to identify well-supported biomarkers from the scientific literature.16–19
To explore the usage of such a text-mining method on marker identification, we select a new category of markers, gut microbiota, which has been linked to CRC by several studies in recent years.20–22 Here, we used an NLP-enabled text-mining system, MarkerGenie, 23 to identify potential stool bacterial markers for early detection and screening of CRC. After experimental validation using digital droplet polymerase chain reaction (ddPCR), we achieved a CRC diagnostic performance of 0.959 area under the curve (AUC) by building a model with the abundance of five bacteria and the FIT results. Our study provides a valuable case applying NLP-based marker mining for cancer diagnosis.
Materials and methods
Patients and sample collection
Stool samples for 117 CRC patients and 77 healthy donors were retrospectively collected at Huazhong University of Science and Technology Union Shenzhen Hospital, China. All samples were collected before colonoscopy surgery and without the intake of laxatives or antibiotics. Only stool samples from individuals with confirmed CRC (based on histopathology) and healthy donors (with no polyp, tumor, or inflammatory bowel disease detected during colonoscopy) were included in the study. A FIT was performed for each sample using the Luminex platform by Tellgen Corporation (Shanghai, China). This study was approved by the Ethics Committee of Huazhong University of Science and Technology Union Shenzhen Hospital (Approval No. KY-2020-045-01). Informed consent was obtained from all enrolled participants. Detailed sample information is presented in Table S1.
Biomedical literature mining
Biomedical literature mining was performed using MarkerGenie V1.0 23 by keyword “colorectal cancer” or “CRC” without any filtering, and only bacterial markers were retained. Subsequently, manual checks were performed based on the biomedical literature evidence count and related sentences in the literature of each marker listed in the results of MarkerGenie.
Qualification of bacteria abundance in stool samples
Fecal samples were thawed from −80°C, and approximately 200 mg stool was used for DNA extraction using QIAamp PowerFecal Pro DNA Kit (Qiagen) according to the manufacturer's instructions. DNA was quantified using Qubit 1× dsDNA HS Assay Kit (Thermo Scientific).
As previously described, multiplex ddPCR of three or four targets was achieved by using a ratio-based probe-mixing strategy. 24 For each target, the probes contained different ratios of two different fluorophores (FAM, HEX), and the probes were mixed at different concentrations to place targets in a defined position on a 2-D plot. Signal readout and preliminary analysis were performed using QuantaSoft software in the QX200 Droplet Reader (Bio-Rad). In addition, QuantaSoft Analysis Pro software provided further analysis of the bacteria abundance by distinguishing the clusters of each target, and the bacteria abundance was normalized to “copy number per 100 μL.”
Primer and probe design for multiplex ddPCR
To design primers and probes for multiplex ddPCR, we developed a K-mer-based primer designing pipeline named “AmpGenie” (manuscript under preparation). Briefly, species-specific K-mers were selected as candidate primers, and primers with suitable GC (guanine and cytosine) content in the entire sequence (40∼60%) and the last five bases (20∼60%) were retained. Subsequently, primers with more than four di-nucleotide sequences or with homopolymer sizes > 4 were removed. After pairing primers based on suitable amplicon size, a probe sequence was designed, and hairpins, homodimers, and heterodimers were filtered using Primer 3.25,26 For multiplex PCR, the non-specific alignment of each two primers was screened using Bowtie 2. 27 Primer combinations with a Tm value difference less than 1°C and without non-specific alignment were selected for experimental validation.
Classifier construction and validation
Classifiers were constructed using Python 3.9. The scikit-learn package was used to perform 5-fold cross-validation. In each fold, the abundance of each marker was Z-score scaled in training cohort, and then a regularized logistic regression model was trained in the training cohort and validated in the test cohort. Finally, the performance in the test cohort of each fold was concatenated to evaluate the performance of the markers.
Statistical analysis
The statistical analyses were performed using R (v4.1.1). Rank-sum test was used to compare variables between groups, using the ggsignif packages (0.6.3) in R. For the receiver operating characteristic (ROC) curve analyses, the area under the curve (AUC), sensitivity, and specificity were calculated using the pROC package (v1.18.0) in R. The Pearson correlation was used to calculate the correlation between the two datasets.
Results
NLP-based marker selection and filtering
To identify bacterial diagnostic markers for CRC, an NLP-enabled text-mining system, MarkerGenie, was used to extract the relationships between marker and disease. 23 We first evaluated the performance of MarkerGenie across various disease areas by comparing them with the Disbiome database. 28 As shown in Figure S1, for the majority of diseases, over half of the bacterial biomarkers with at least three supporting evidences in the Disbiome database were also successfully identified by MarkerGenie, with at least three supporting articles, highlighting the effectiveness of MarkerGenie. Subsequently, we used the keywords “colorectal cancer” and “CRC”, and only selected bacterial markers for CRC biomarker screening. Subsequently, several criteria were used for marker filtering:(a) only bacterial species were retained; (b) more than three articles must support the marker; (c) there was no conflict among different studies (e.g., some bacteria were reported to have increased in abundance in some studies, but decreased in others), and only the markers with direct evidence were retained. (If some bacteria were reported to have increased abundance in CRC, or were included in predictive models for CRC, the article was a “direct evidence” type. Also, the evidence in some articles could not directly support the marker; for example, the bacteria were reported to secrete a toxin, which could induce CRC. This kind of article should not be regarded as a “direct evidence” type); (d) probiotics were excluded, because the effectiveness of probiotics in cancer development has not been fully demonstrated, and might not be suitable for cancer screening (Figure 1(a)); and (e) 16 bacteria met the criteria, and 12 bacteria were selected for further validation, because for some markers of the same genus, we only selected one for validation (Figure 1(b)). Moreover, we noticed that the two species with the most published studies, Escherichia coli and Bacteroides fragilis, had several studies with conflicting conclusions. Thus, we only included genotoxic Escherichia coli with the pks gene29,30 and enterotoxigenic Bacteroides fragilis (ETBF) with the bft gene,31,32 which have been proven to involve in CRC tumorigenesis. Therefore, the candidate marker list for CRC screening consisted of a total of 14 bacteria (Figure 1(b)).

NLP-based bacteria marker selection for CRC. (a) Criteria used for marker filtering. (b) The markers selected for experimental validation; the markers of the same group were amplified in the same PCR reaction. The markers labeled by * are strains.
Stool bacterial marker abundance validation using ddPCR
We established a multiplex ddPCR system to experimentally quantify the copy numbers of the bacterial markers in stool. The primers were validated using plasmid-based standards (Figure S2), and three or four markers were amplified in each tube (Figure S3). We then compared copy numbers of bacterial markers between CRC patients and healthy donors. Seven markers (Peptostreptococcus stomatis, Streptococcus gallolyticus, Parvimonas micra, Clostridium symbiosum, Fusobacterium nucleatum, Solobacterium moorei, and Gemella morbillorum) showed a significant difference using an alpha of 0.05 (P = 1.0 × 10−13, 1.8 × 10−11, 0.034, 2.7 × 10−13, 1.0 × 10−15, 3.7 × 10−8, and 0.046, respectively, Wilcoxon rank-sum test, Figure 2). However, the other seven markers did not have a significant difference in our cohort (Figure S4). To explore further we collected seven publicly available large-scale CRC metagenomic datasets, and analyzed the relative abundance, prevalence, fold-change, and P-value of the bacterial markers.21,22,33–36 Most markers with no significant difference in our study also had low prevalence or low fold-change in the public datasets with the exception of Porphyromonas asaccharolytica (Figure 3(a–d)). Most of the public datasets were based on the Caucasian population; for the only public datasets with primarily an Asian population the abundance of Porphyromonas asaccharolytica was much lower, which might explain the low prevalence of Porphyromonas asaccharolytica in our cohort (Figure 3(a)).
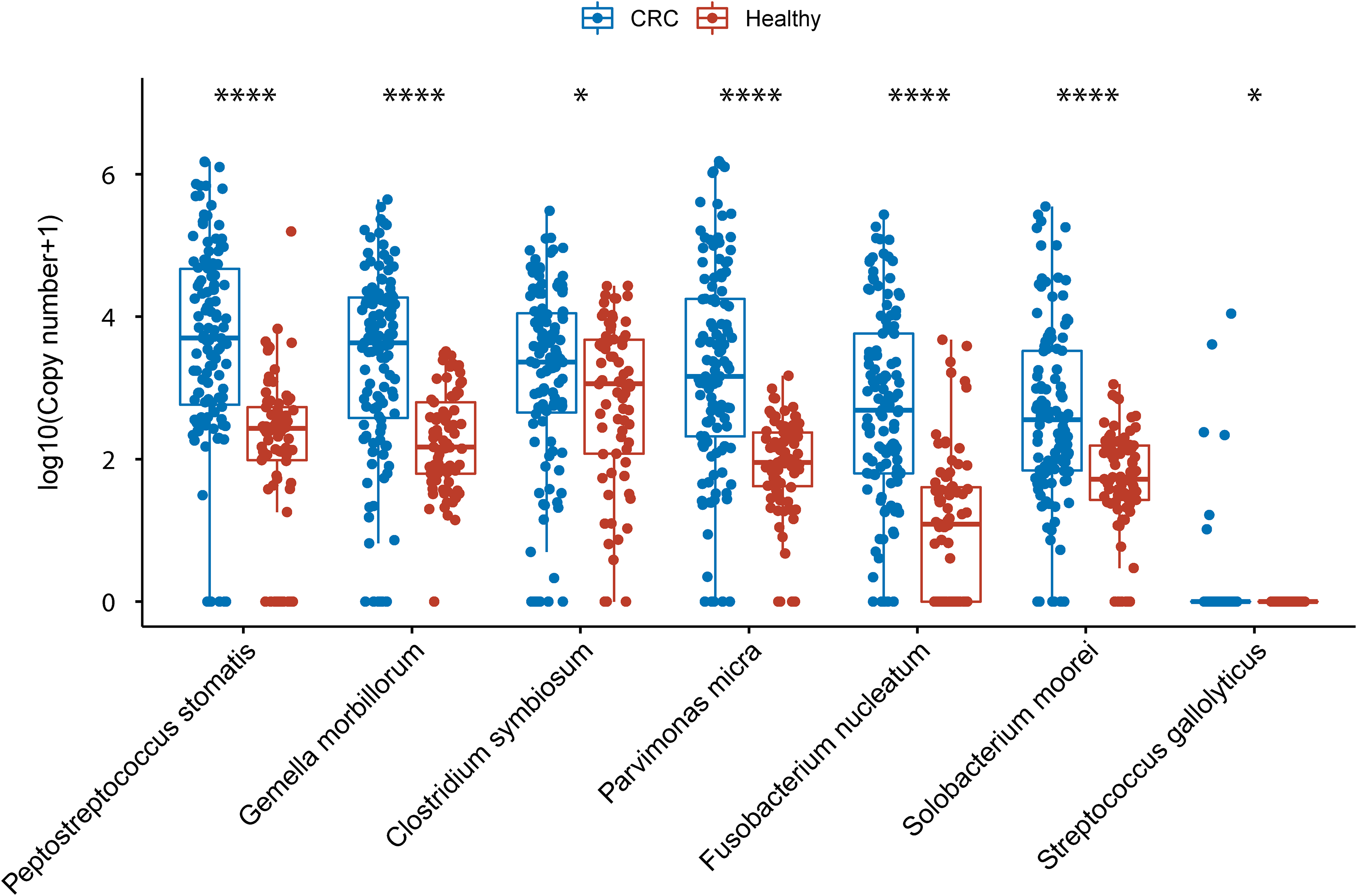
Relative abundance of bacteria markers with significant difference in stool between CRC patients and healthy donors. (* P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001, Wilcoxon rank-sum test).
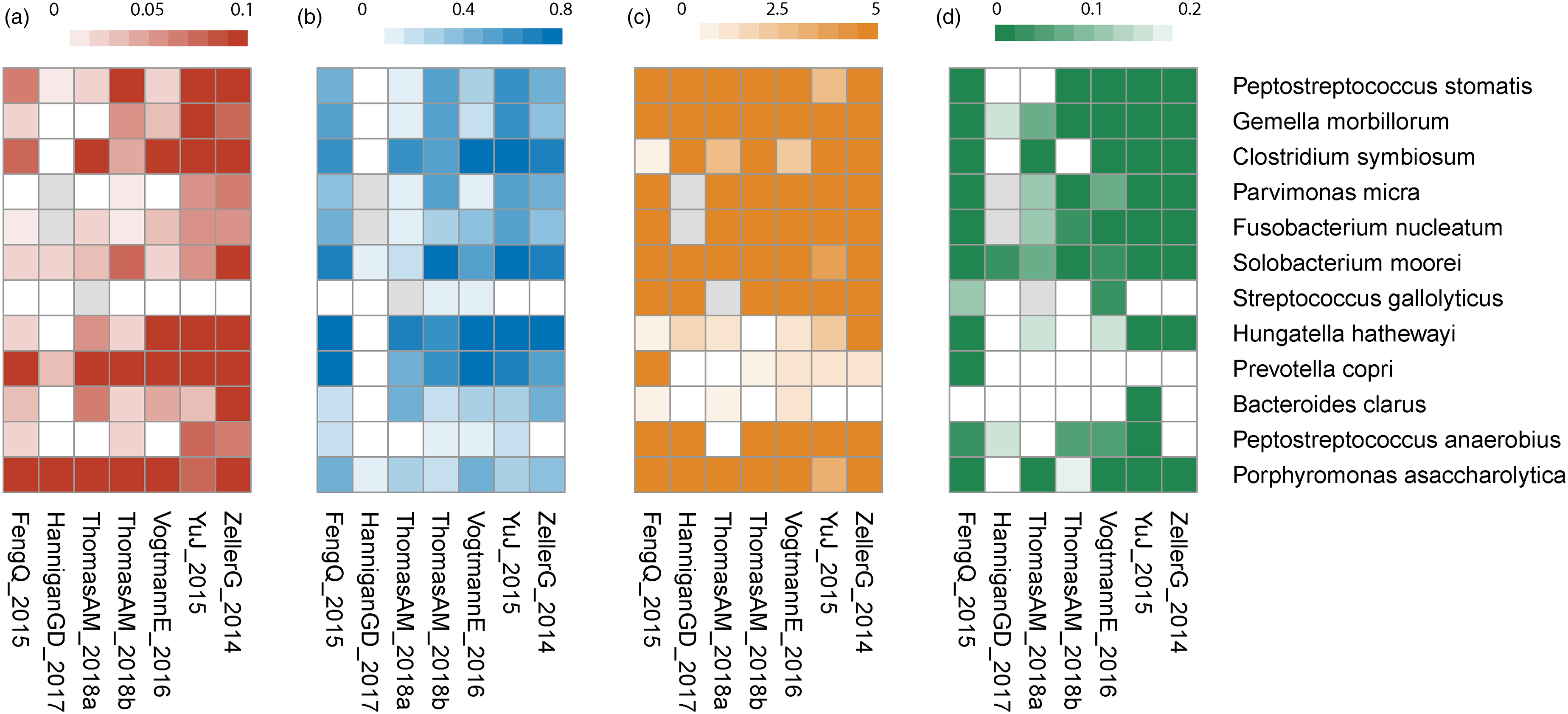
Heatmaps of bacteria markers in seven CRC metagenomic datasets. (a) Relative abundance in CRC patients. (b) Prevalence in CRC patients. (c) Fold-change between CRC patients and healthy donors. (d) P-value between CRC patients and healthy donors, Wilcoxon rank-sum test.
Stool bacterial markers could discriminate between CRC patients and healthy donors
Next, we tested whether these stool bacterial markers were useful for CRC diagnosis. Five bacterial markers (Peptostreptococcus stomatis, Parvimonas micra, Fusobacterium nucleatum, Solobacterium moorei, and Gemella morbillorum) achieved individual AUCs > 0.70 in distinguishing between CRC patients and healthy donors (Figure S5), and principal component analysis based on these five bacterial markers showed samples were grouped according to whether they were or were not CRC patients (Figure S6). Moreover, these markers are highly correlated with each other (Figure S7). Thus, we used the combination of these markers to construct a classifier for CRC patients and healthy donors. Using 5-fold cross-validation, a mean AUC of 0.852 was achieved (sensitivity = 0.692, and specificity = 0.935, Figure 4(a)), suggesting that stool bacterial markers alone are highly predictive for CRC patients.
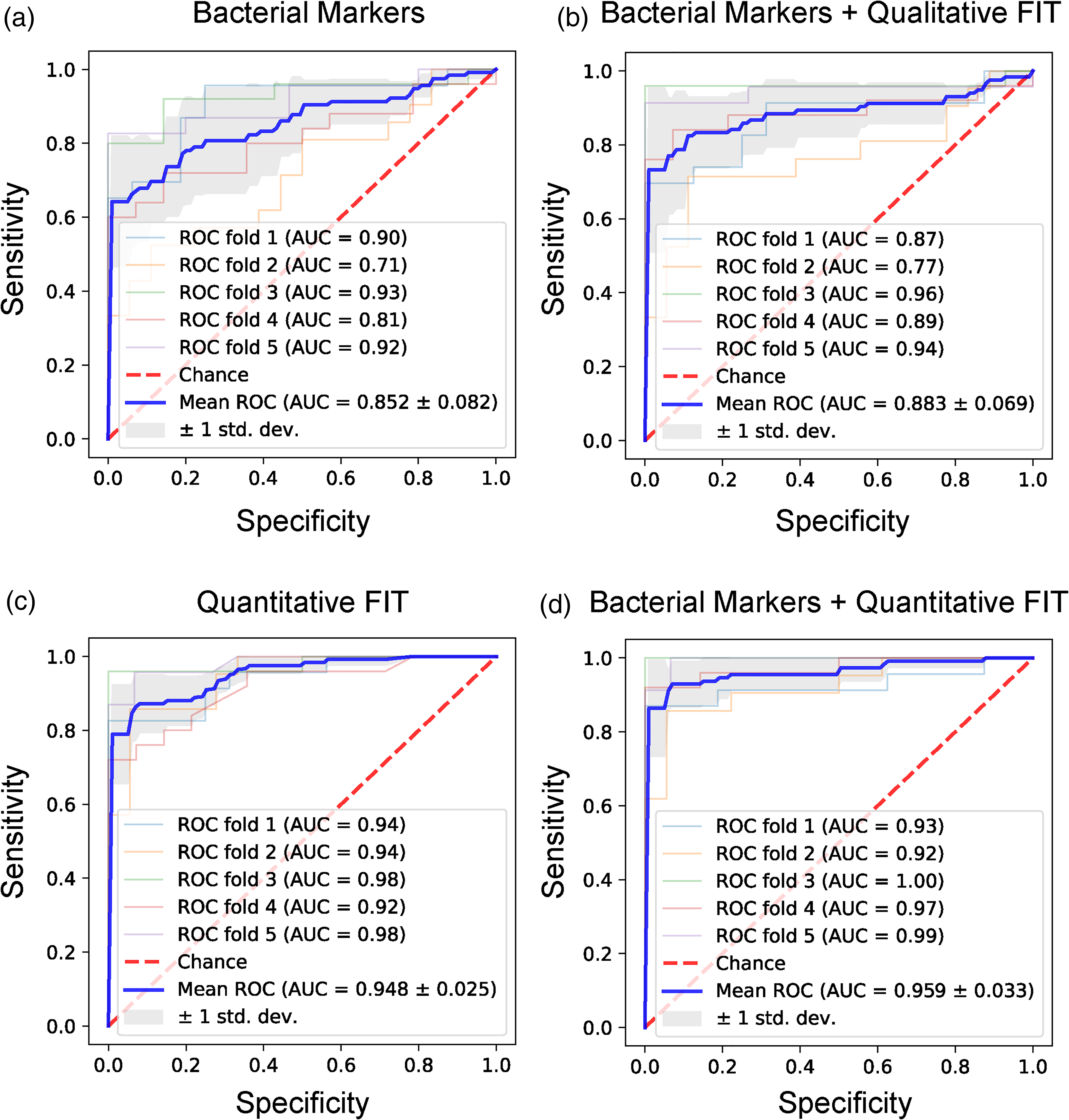
Performance of bacteria markers on CRC detection in test cohort. (a) ROC curve of five bacteria markers. (b) ROC curve of five bacteria markers combined with qualitative FIT. (c) ROC curve of quantitation of hemoglobin and transferrin. (d) ROC curve of five bacteria markers combined with quantitation of hemoglobin and transferrin.
Stool bacterial markers might be complemented with FIT
A FIT test for small amounts of blood in the stool detects hemoglobin and transferrin proteins, and is the most used method for CRC screening. However, the sensitivity of FIT decreases when the tumor is in the upper intestines, likely due in part to degradation by digestive enzymes.3,37 The bacterial markers are not as affected by digestive processes and therefore might complement FIT. To verify this assumption, we compared the abundance of bacterial markers among patients with different tumor locations, and no significant difference was found (Figure S8). Additionally, none of these markers was highly correlated with the abundance of hemoglobin or transferrin (Figure S7).
Stool bacterial markers improved the performance of FIT
Next, we built classifiers by combining bacterial markers and FIT. FIT alone achieved a sensitivity of 0.709 and a specificity of 0.961, which is consistent with other reports. 5 A combination of FIT and bacterial markers achieves an AUC of 0.883, with sensitivity = 0.803 and specificity = 0.948 (Figure 4(b)). Moreover, we also built classifiers directly using the quantitation of hemoglobin and transferrin, and achieved an AUC of 0.948 (sensitivity = 0.803, and specificity = 0.961, Figure 4(c)). We next combined our bacterial markers with the quantitation of hemoglobin and transferrin, and re-trained the classifier. Although the bacterial markers did not increase the AUC much (0.959 vs. 0.948), the classifiers with the quantitative FIT and bacterial markers achieved a higher sensitivity (0.929 vs. 0.872) at a specificity of 0.900 (Figure S9), suggesting that the bacterial markers enabled more sensitive identification of CRC patients. This is consistent with our observation that bacterial markers contrary to FIT did not decrease in signal comparing proximal with distal tumors.
Discussion
The NLP-based biomedical text-mining methods can digest research articles more efficiently and comprehensively compared to human researchers, so such methods have been widely used to identify associations between biomarkers and diseases.16–19 However, few studies directly use such methods to develop cancer screening and early detection assays, and no “best practice” workflow or validation has been provided on NLP-based assay development. In this study, using NLP-based biomedical text-mining software, we searched enterobacteria markers for CRC screening and early detection. After manually checking the search results, we validated the performance of these markers using ddPCR and built classifiers for CRC detection in clinical samples. Our results provide a recommendation pipeline for NLP-based marker screening and filtering.
Screening of effective biomarkers is complicated and difficult. Although NLP-based methods could automatically generate the relationships between biomarkers and diseases, a manual check is still necessary before experimental validation. For example, some markers might be reported as pathogenic in some studies, but be reported as disease repressors in other studies. Some markers reported an association with a disease, but sometimes such associations only exist in small subtypes of the disease. Additionally, some abbreviations might mislead the NLP-based methods, which would lead to a wrong connection between a marker and a disease. Such markers reported by NLP-based methods need to be excluded.
Recent evidence suggests that large language models (LLMs), such as ChatGPT and Bard, have the potential to assist in the discovery of biomarkers. We utilized ChatGPT to search for microbial biomarkers of CRC by repeating the search prompt “Provide 15 microbial biomarkers on colorectal cancer” five times. Although the biomarker lists provided by ChatGPT varied across the repetitions, we found that most of the markers mentioned by ChatGPT were also covered by MarkerGenie (Table S2). However, it is important to note that LLMs cannot provide the specific references for each biomarker. Therefore, it is not possible to manually trace or verify the biomarkers provided by LLMs. Nevertheless, LLMs can quickly review and summarize a vast number of sentences retrieved by MarkerGenie. The integration of MarkerGenie and LLMs has the potential to enhance biomarker identification efficiency and to reduce manual work in future research.
Fortunately, after manual filtering, 7 of the 12 bacteria species in our study showed significantly increased abundance in CRC patients, suggesting that at least half of the markers identified by our pipeline could be verified. Moreover, 5 bacteria markers were discriminative between CRC patients and healthy donors, implying that an NLP-based marker screening pipeline could provide a reasonable marker list even without re-analyzing any omics datasets. Therefore, our NLP-based pipeline could provide a hint for assay development, even for the research groups without experienced data scientists.
It is worth noting that the bacteria markers identified in our study were also discovered and validated by several metagenomic studies.20–22 Furthermore, four of the five markers, including Peptostreptococcus stomatis, Parvimonas micra, Fusobacterium nucleatum, and Gemella morbillorum, were also included in predictive models built in a meta-analysis study, 22 suggesting the markers identified in our study are credible and reproducible. Also, we found two markers, Parvimonas micra 38 and Fusobacterium nucleatum,38–42 were not only mentioned in studies of 16S rRNA gene sequencing or metagenomics sequencing, suggesting that even the markers not reported in the research articles on high throughput data analysis could also be identified by our pipeline. For many diseases, because there are not sufficient high-quality datasets available, such an NLP-based pipeline could collect evidence from functional studies and studies with low through-put marker validation, and might help to identify useful markers.
Using several publicly available metagenomic data, we also analyzed the reason why some of the bacterial biomarkers supported in the literature were not reproducible in our study.21,22,33–36 Not surprisingly, low prevalence, low fold-change, and low abundance could explain most of the failures of validation, suggesting such omics data-based filtering steps could help to filter markers for assay development. The NLP-based method could also suggest whether the marker plays driver roles or passenger roles in the disease. Therefore, combining omics data analysis with our pipeline could help to identify explicable markers with high confidence.
In conclusion, our study identified CRC diagnostic markers using an NLP-based text-mining pipeline and demonstrated that such an NLP-based pipeline could identify useful markers with or without additional omics data validation. It is worth noting that such pipelines could not only be applied for disease diagnosis, but also contribute to companion diagnosis, disease stratification, and prognosis prediction. In the future, the combination of NLP-based pipelines and high-throughput experimental validation methods could increase the effectiveness of biomarker identification for translational medicine and clinical usage.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155231210881 - Supplemental material for Development of fecal microbial diagnostic marker sets of colorectal cancer using natural language processing method
Supplemental material, sj-docx-1-jbm-10.1177_03936155231210881 for Development of fecal microbial diagnostic marker sets of colorectal cancer using natural language processing method by Houcong Liu, Changpu Song, Jidong Wang, Zhufang Chen, Xiaohong Zhang, Hekai Zhou, Linhong Yao, Dan Chen, Wenhao Gu, Rui-Kun Huang, Bing-Kun Huang, Bo-Wei Han and Jihui Du in The International Journal of Biological Markers
Supplemental Material
sj-docx-2-jbm-10.1177_03936155231210881 - Supplemental material for Development of fecal microbial diagnostic marker sets of colorectal cancer using natural language processing method
Supplemental material, sj-docx-2-jbm-10.1177_03936155231210881 for Development of fecal microbial diagnostic marker sets of colorectal cancer using natural language processing method by Houcong Liu, Changpu Song, Jidong Wang, Zhufang Chen, Xiaohong Zhang, Hekai Zhou, Linhong Yao, Dan Chen, Wenhao Gu, Rui-Kun Huang, Bing-Kun Huang, Bo-Wei Han and Jihui Du in The International Journal of Biological Markers
Supplemental Material
sj-docx-3-jbm-10.1177_03936155231210881 - Supplemental material for Development of fecal microbial diagnostic marker sets of colorectal cancer using natural language processing method
Supplemental material, sj-docx-3-jbm-10.1177_03936155231210881 for Development of fecal microbial diagnostic marker sets of colorectal cancer using natural language processing method by Houcong Liu, Changpu Song, Jidong Wang, Zhufang Chen, Xiaohong Zhang, Hekai Zhou, Linhong Yao, Dan Chen, Wenhao Gu, Rui-Kun Huang, Bing-Kun Huang, Bo-Wei Han and Jihui Du in The International Journal of Biological Markers
Footnotes
Author contributions
Houcong Liu: investigation, data curation, roles/writing—original draft. Changpu Song: investigation, data analysis, roles/writing—original draft. Jidong Wang: methodology, validation. Zhufang Chen: formal analysis. Xiaohong Zhang: methodology. Hekai Zhou: resources. Linhong Yao: resources. Dan Chen: validation. Wenhao Gu: software, visualization. Rui-Kun Huang: data curation. Bing-Kun Huang: validation. Bo-Wei Han: conceptualization, writing—review and editing. Jihui Du: conceptualization, writing—review and editing, funding acquisition. Houcong Liu and Changpu Song contributed equally to this work.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Funds of Health Science and Technology Research Key Project of Nanshan District, Shenzhen [grant number NS2022001], and the Research Funds of the Shenzhen Science and Technology Innovation Commission [grant number JCYJ20190809105001757].
Ethics approval and consent to participate
This study was approved by the Ethics Committee of Huazhong University of Science and Technology Union Shenzhen Hospital (Approval No. KY-2020-045-01). Informed consent was obtained from all enrolled participants.
Data availability
The datasets used and analyzed during the current study are available from the corresponding author upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
