Abstract
Introduction
Cancer is the leading cause of death in economically developed countries and the second leading cause of death in developing countries, and is caused by the complex interplay of genetic and environmental risk factors (1, 2). Digestive tract cancer (DTC), which refers to a group of malignancies, is the most common type of cancer worldwide. Especially, gastric and esophageal cancers are global epidemiological health concerns (3). Gastric and colorectal cancers were respectively the second and third most common causes of cancer-related mortality worldwide in 2008, accounting for over 1 million deaths (4). In China, gastric cardia adenocarcinoma (GCA) remains a major cancer burden: two thirds of the incident cases take place in rural areas, thus ranking the third most common cancer (5); adding to this, the incidence of colorectal cancer (CRC) is dramatically increasing in developing countries (including China). Furthermore, esophageal squamous cell carcinoma (ESCC) is the sixth most common cause of cancer-related death worldwide (6), and is characterized by a remarkable geographical distribution with a higher prevalence in Eastern Asia (7). China lies in the ‘‘esophageal cancer belt'' (8): 90% of the cases recorded are squamous cell carcinomas, which contribute to about half of the world's cases (9-10-11).
In recent years, the genetic predisposition has been considered as a combined influence factor. Genome-wide association studies (GWAS) play important roles in the identification of potential candidates for single nucleotide polymorphisms (SNPs) that may be involved in the disease pathogenesis. In 2010, Abnet et al (12) and Wang et al (13) have simultaneously identified the rs2274223 A>G polymorphism in phospholipase C epsilon 1 (PLCE1) as a novel susceptibility locus for ESCC and GCA in the Chinese population in 2 separate comprehensive GWAS. Later, in another GWAS, Wu et al (14) showed the significant association of PLCE1 rs2274223 polymorphism in ESCC, pointing to an intricate role of PLCE1 in malignancy.
The PLCE1 gene is located on chromosome 10q23 and is a unique member of the phospholipase family. PLCE1 encodes the phospholipase C epsilon 1 (PLCE1) that catalyzes the hydrolysis of phosphatidylinositol-4,5-bisphosphate (PIP2) into the secondary messengers inositol 1,4,5-trisphosphate (IP3) and diacylglycerol (DAG). These participate in cell growth, differentiation, and gene expression (15, 16). Recently, studies have reported that PLCE1 plays a fundamental role in carcinogenesis and progression of a series of human cancers, including cancers of the intestine, skin, bladder, colorectal, and head and neck (17-18-19-20-21). All these findings further support the biological plausibility that genetic variations affect the gene expression or protein functions and may be a potential factor for carcinogenesis.
To date, the association between the PLCE1 rs2274223 (A>G) and the susceptibility to DTC remains unclear. This study is aimed at identifying the genetic factors that predispose individuals to DTC in the Chinese population. Thus, we conducted a meta-analysis on all eligible case-control studies to estimate the overall DTC risk of the PLCE1 rs2274223 (A>G) polymorphism.
Materials and Methods
Identification and Eligibility of Relevant Studies
This meta-analysis was performed by searching PubMed, Springer, Elsevier, CNKI (Chinese), and Wanfang (Chinese) Digital Dissertations Databases for relevant articles published in English or Chinese up to November 2013, with key words ‘rs2274223', ‘PLCE1 polymorphism', and ‘cancer'. We also hand searched for references of original studies on this topic in order to identify additional studies.
All the candidate articles were carefully examined to determine whether they met the inclusion criteria for the meta-analysis. The inclusion criteria were as follows: (i) testing the influence of rs2274223 polymorphisms on cancer risk in the Chinese population; (ii) case-control study designed; (iii) sufficient published data for estimating odds ratio (OR) with 95% confidence interval (CI); and (iv) genotype frequencies available.
The articles including controls with a genotype that did not meet the assumptions of the Hardy-Weinberg equilibrium (HWE) were excluded from the analysis of this polymorphism.
Data Extraction
The data were obtained according to a standard protocol. Two investigators independently extracted the data and reached a consensus on all the items in case of discordance. All the studies included in this meta-analysis meet the inclusion criteria. For each study, the following characteristics and data were collected: first author's name, publication year, cancer type, country of origin, ethnicity, study design, genotyping method, numbers of genotyped cases and controls, genotype distributions and value of the HWE. The study design was stratified into population-based and hospital-based studies. If original genotype frequency data were unavailable in relevant articles, a request for additional data was sent to the corresponding author.
Statistical Analyses
The analyses were conducted with Review Manager 5.0. The risk of DTC associated with the PLCE1 rs2274223 variant was indicated by the pooled OR with 95% CI. Pooled ORs were obtained from the combination of individual studies by allele comparison (G vs A), heterozygote comparison (GA vs AA), homozygote comparison (GG vs AA), a dominant model (GG/GA vs AA), and a recessive model (GG vs GA/AA). Furthermore, the analyses were stratified by sub-grouping the studies based on cancer types (ESCC, GCA, and others) and source of controls.
Heterogeneity across studies was evaluated by the Cochran's Q test and I2, which represents the percentage of total variation across the studies that is attributable to heterogeneity. Heterogeneity was considered statistically significant when p<0.05, in which case the pooled OR estimate of each study was calculated by the random-effects model (DerSimonian and Laird method) (22). Otherwise, the fixed-effect model (Mantel-Haenszel method) was used (23). The significance of the overall OR was determined by the Z-test. If there was significant heterogeneity among the included studies, the sources of heterogeneity would be explored using meta-regression in Stata version 12.0 (http://www.stata.com).
To assess the stability of the results, one-way sensitivity analyses were carried out as to evaluate the influence of each study on the overall estimate from the meta-analysis. Publication bias was assessed using funnel plots, Begg's test (rank correlation test), and Egger's test (weighted linear regression test) for funnel plot symmetry with Stata 12.0. Statistical tests performed in the present analysis were considered significant whenever the corresponding null-hypothesis probability was p<0.05.
Results
Study characteristics
The main characteristics of the 13 articles included in the meta-analysis (15 studies) (12-13-14, 24-25-26-27-28-29-30-31-32-33) are summarized in Table I. The study selection process is shown in Figure 1, which also indicates that two articles (12, 13) included 2 tumor types each. A total of 29,805 cases and 32,225 controls were included; 2 articles (12, 13) exploring ESCC and GCA included the same controls. Among the selected articles, 8 studies were on ESCC, 5 on GCA, and one each on CRC and head and neck cancers (HNC). Overall, 12 studies recruited population-based controls and 3 studies were performed on hospital-based controls. In 14 studies the analysis of a portion of the samples was randomly repeated as quality control of genotyping.
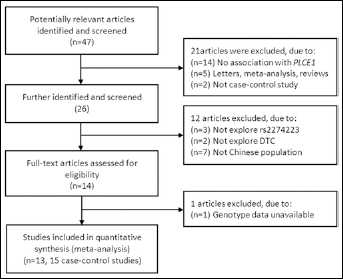
Flow chart of literature search and study selection.
CHARACTERISTICS OF THE STUDIES INCLUDED IN THE META-ANALYSIS
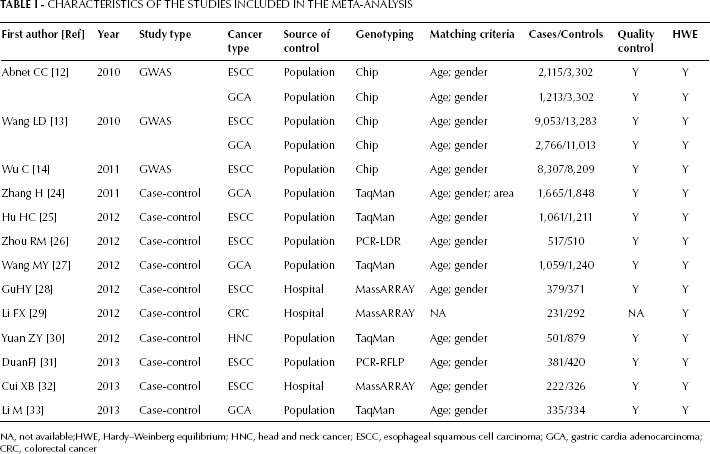
NA, not available;HWE, Hardy–Weinberg equilibrium; HNC, head and neck cancer; ESCC, esophageal squamous cell carcinoma; GCA, gastric cardia adenocarcinoma; CRC, colorectal cancer
Quantitative Synthesis
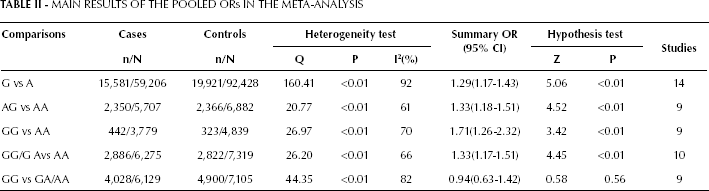
The evaluation of the association between the PLCE1 rs2274223 polymorphism and the susceptibility to DTC is presented in Table II and Figure 2. In the overall analysis, the variant G allele of rs2274223 A>G significantly increased the risk of cancer in all genetic models (allele comparison, G vs A: OR=1.29, 95% CI: 1.17-1.43, p<0.01; heterozygote comparison, GA vs AA: OR=1.33, 95% CI: 1.18-1.51, p<0.01; homozygote comparison, GG vs AA: OR=1.71, 95% CI: 1.26-2.32, p<0.01; dominant model, GG/GA vs AA: OR=1.33, 95% CI: 1.17-1.51, p<0.01), except for the recessive model (GG vs GA/AA: OR=0.94, 95%CI: 0.63-1.42, p=0.56).
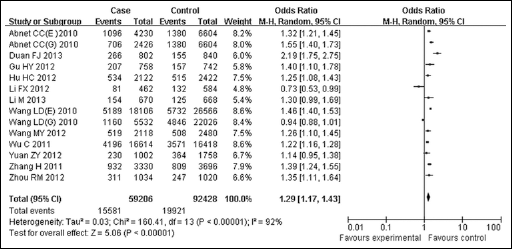
Forest plot of the cancer risk associated with PLCE1 rs2274223 for the allele comparison (G vs A). Squares and horizontal lines respectively correspond to the study-specific OR and 95% CI. The area of the squares reflects the study-specific weight. The diamond represents the pooled OR and 95% CI.
MAIN RESULTS OF THE POOLED ORs IN THE META-ANALYSIS
We performed a further stratification analysis by cancer types (Tab. III, Fig. 3). For the ESCC, the results showed that the individuals carrying the rs2274223 AG or GG genotype had a significantly higher cancer risk than those with the rs2274223 AA genotype (AG vs AA: OR=1.56, 95% CI: 1.25-1.93, p<0.01 and GG vs AA: OR=2.10, 95% CI: 1.30-3.38, p<0.01). Also, the PLCE1 rs2274223 G allele and GG/GA genotype were found to be associated with an increased risk of cancer compared to the A allele and AA genotype (G vs A allele: OR=1.39, 95% CI: 1.26-1.55, p<0.01 and GG/GA vs AA: OR=1.46, 95% CI: 1.31-1.62, p<0.01). For GCA, we found a significant association between the PLCE1 rs2274223 polymorphism and DTC risk in the allele comparison, heterozygote comparison, homozygote comparison and dominant model (G vs A: OR=1.27, 95% CI: 1.01-1.59, p<0.01; AG vs AA: OR=1.30, 95% CI: 1.20-1.50, p<0.01; GG vs AA: OR=1.78, 95% CI: 1.42-2.23, p<0.01 and GG/GA vs AA: OR=1.39, 95% CI: 1.25-1.55, p<0.01). None of the associations was significant in the subgroup of ‘‘other cancers'' (including CRC and HNC).
STRATIFIED ANALYSES OF THE PLCE1 rs2274223 POLYMORPHISM ON CANCER RISK
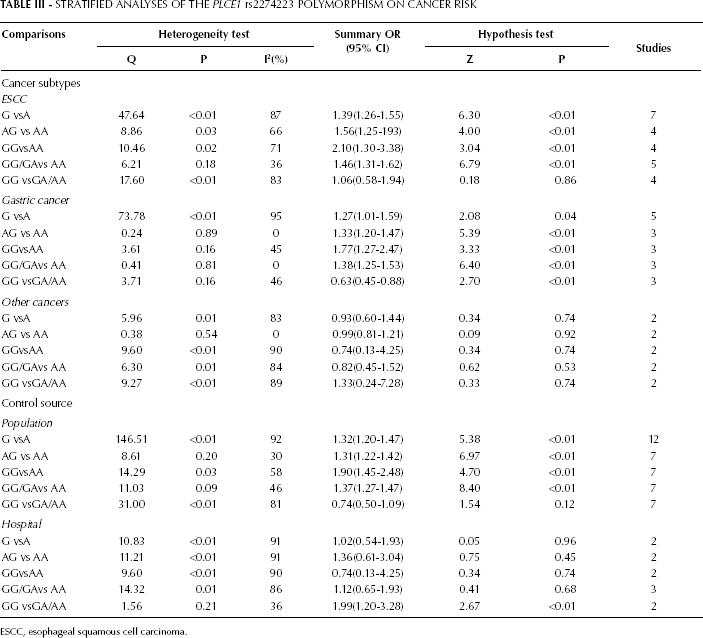
ESCC, esophageal squamous cell carcinoma.
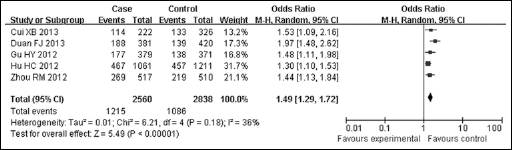
Forest plot of the cancer risk associated with PLCE1 rs2274223 for the dominant model (GG/GA vs AA) for ESCC. Squares and horizontal lines correspond to the study-specific OR and 95% CI. The area of the squares reflects the study-specific weight. The diamond represents the pooled OR and 95% CI.
By subgroup analyses based on the source of controls (Tab. III) we found that the PLCE1 rs2274223 polymorphism might increase the risk of DTC in all genetic models in a population-based subgroup (allele comparison, G vs A: OR=1.32, 95% CI: 1.20-1.47, p<0.01; heterozygote comparison, GA vs AA: OR=1.31, 95% CI: 1.22-1.42, p<0.01; homozygote comparison, GG vs AA: OR=1.90, 95% CI: 1.45-2.48, p<0.01; dominant model, GG/GA vs AA: OR=1.37, 95% CI: 1.27-1.47, p<0.01), except for the recessive model (GG vs GA/AA: OR=0.74, 95% CI: 0.50-1.09, p=0.78). No significant associations were found in the hospital-based subgroup, except for the recessive model (GG vs GA/AA: OR=1.99, 95% CI: 1.20-3.28, p=0.78).
Test of Heterogeneity
When evaluating the association between the PLCE1 rs2274223 polymorphism and the susceptibility to DTC, we found that a significant heterogeneity in all genetic models in the overall analysis (Pheterpgeneity<0.05). Therefore, meta-regression in Stata 12.0 was used to assess the source of heterogeneity for rs2274223 including the following parameters: publication year, cancer type, matched controls (yes or not), language (English or Chinese), source of control (population or hospital), assay, sample size (500 as the cutoff), and quality control (with or without). We found that the systemic results were not affected by these characteristics (Tab. IV).
RESULTS OF THE HETEROGENEITY TEST FOR PLCE1 rs2274223
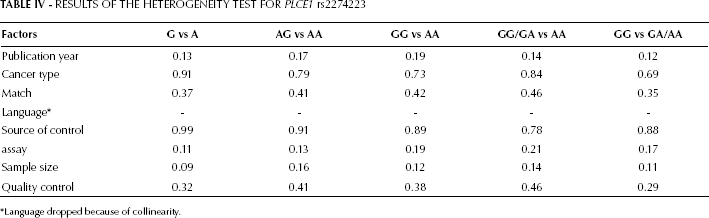
Language dropped because of collinearity.
Sensitivity Analysis
Sensitivity analysis was performed to assess the influence of each individual study on the pooled ORs by omission of individual studies: the corresponding pooled ORs were not altered by this analysis (data not shown), suggesting that our results were statistically robust.
Publication Bias
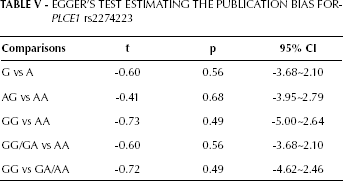
The Begg's funnel plots and Egger's tests were performed to assess the publication bias of the currently available literature. The shape of the funnel plots did not reveal any evidence of obvious asymmetry in all comparison models (Fig. 4). Then, the Egger's test was used to provide statistical evidence for funnel plot symmetry, and once again, the results did not show any clear evidence of publication bias (Tab. V).
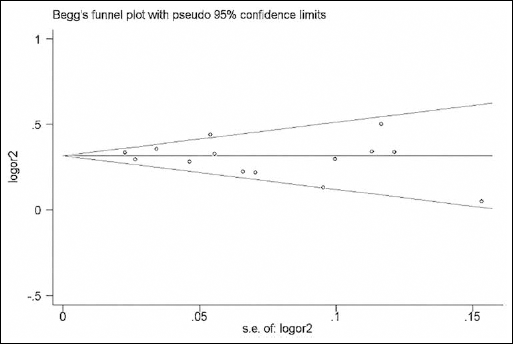
Funnel plot of the PLCE1 rs2274223 polymorphism and DTC risk for allele comparison (G vs A).
EGGER'S TEST ESTIMATING THE PUBLICATION BIAS FOR PLCE1 rs2274223
Discussion
GWASs represent an unbiased and fairly comprehensive approach to exploring the genetic variation related to cancer. Three independent GWASs, recently performed in the Chinese population, identified the PLCE1 rs2274223 variation as a susceptibility locus for both ESCC and GCA (12-13-14). However, the replications of this finding in various types of malignancies are inconsistent. Therefore, we carried out a meta-analysis of the replication studies incorporating 15 case-control studies. To our knowledge, this is the first study investigating the association between the PLCE1 rs2274223 A>G polymorphism and DTC risk in the Chinese population.
PLCE1 is a member of the phospholipase C family of proteins. It differs from other members of its family and can uniquely interact with the RAS proto-oncogene. The protein functions as an effector of guanosine triphosphatases (GTPases) such as Ras, Rap1, and Rap2, involving the regulation of cell growth, differentiation, and apoptosis (15, 21). Mutations in the RAS gene family are associated with approximately 30% of all human cancers, in terms of both cancer development and progression (21, 34). So far, PLCE1 has been reported to have multiple mutation sites such as rs2274223, rs11187870, rs17417407, rs2274224, and rs3765524 (13, 25, 31, 35). All the abovementioned studies discussed the relationship between the rs2274223 gene variant and ESCC or GCA risk. However, the frequencies of the rs11187870, rs2274224, and rs3765524 gene variants were insufficient to perform a pooled analysis. This meta-analysis only examines the association between the PLCE1 rs2274223 gene variant and DTC risk.
In the present study, the results of our meta-analysis indicated that the G allele of the PLCE1 rs2274223 variant is significantly associated with an increased risk of DTC (in terms of allele comparison, heterozygote comparison, homozygote comparison, and dominant model), and especially with ESCC and GCA in homozygote comparisons (OR=2.10, 95% CI: 1.30-3.38, p<0.01 and OR=1.77, 95% CI: 1.27-2.47, p<0.01). Interestingly, similar results have been obtained by Zhang et al (36) in other Asian populations. However, no significant associations were found in the subgroup of “other cancers”. The results are consistent with the conclusions of previous studies (37), despite the association of the rs2274223 was not confirmed in the other published study (38). Thus, it appears that PLCE1 may not be a risk factor for the subgroup of ‘‘other cancers'' (CRC and HNC).
Stratification based on the source of controls showed that the PLCE1 rs2274223 polymorphism is associated with population-based controls in every genetic model, but a lack of associations was found in hospital-based controls (except for the recessive model). In contrast to the published results, this study reveals the different associations (39). Possible reasons for these discrepancies are the differences in genetic backgrounds, and the limited number of eligible studies with available data, which together may determine an insufficient statistical power to detect a slight difference and may also generate a fluctuated risk estimate.
The strengths of this study include the relatively large sample size, the lack of deviation from HWE of the controls in all included studies, and the low probability of publication bias. However, some limitations need to be addressed. Firstly, due to the limited availability of published results, the number of studies included in each meta-analysis was relatively small and may not provide sufficient statistical power to estimate the correlation between the PLCE1 rs2274223 polymorphism and DTC risk. Secondly, in the eligible studies, some controls were selected from hospital populations: these subjects might have benign digestive tract diseases and might determine a potentially incremental risk of DTC. Thirdly, the overall outcomes were based on unadjusted estimates, and the lack of original data in some studies limited our further evaluation of potential interactions. Nevertheless, it is well acknowledged that gene-gene or gene-environment interactions may affect the risk of DTC. Finally, although the Begg's Funnel plot and Egger's test did not show any publication bias, we cannot exclude a possible selection bias as only studies published in English language were included in the meta-analysis.
Conclusion
Our results support the association of the PLCE1 rs2274223 A>G polymorphism with increased risk of DTC in the Chinese population, especially with ESCC and GCA. Nevertheless, larger and well-designed multicenter studies, including gene-gene, and gene-environment interactions should be carried out to validate our findings.
