Abstract
Purpose
Wear of ceramic orthopedic devices generates nanoparticles in vivo that may present a different biological character from the monolithic ceramic from which they are formed. The current work investigated protein adsorption from human plasma on alumina nanoparticles and monolithic samples representative of both wear particles and the ceramic components as implanted.
Materials and Methods
A physicochemical characterization of the particles and their dispersion state was carried out, and the protein adsorption profiles were analyzed using 1D SDS-PAGE and mass spectrometry.
Results
Significant differences in protein-binding profiles were identified where the nanoparticles selectively bound known transporter proteins rather than the more highly abundant serum proteins that were observed on the monoliths.
Conclusions
Proteins associated with opsonization of particles were seen to be present in the protein corona of the nanoparticles, which raises questions regarding the role of wear particles in periprosthetic tissue inflammation and aseptic loosening.
Introduction
In orthopedic devices, wear particles generated at the contact surfaces of the implant bearing are the primary cause of aseptic loosening (1). This is often the limiting factor for the life of the implant. With the standard hip implant bearing couple, cobalt chrome alloy on ultra-high-molecular-weight polyethylene (UHMWPE), it has been estimated that as many as 500,000 wear particles, with mean diameter of 0.5 μm, are generated in every step (2). Recent research has focused on “hard-on-hard” bearings – i.e., metal-on-metal (MoM) and ceramic-on-ceramic (CoC) – as their volumetric wear rates are 1 to 2 orders of magnitude less than that for the “hard-on-soft” bearings described above. However, the size distribution of their wear particles is also shifted 1 to 2 orders of magnitude into the nanoscale range, giving the wear particles the potential to enter cells via receptor-mediated processes. Laboratory simulations estimate that as much as 98% of the wear particles are between 5 and 20 nm, with polygonal and oval morphologies both being reported (3, 4). Taking the average volumetric wear for a CoC bearing as 0.1 mm3/year (5) and considering each particle to be the maximum of 20 nm in diameter, then in each year more than 2.3 × 107 particles are released. These nanoparticles are known to disseminate to the liver, spleen and abdominal lymph nodes (6), but few studies have gone beyond cytotoxicity testing to really estimate their biological activity and the effect of chronic exposure to as many as 20 million particles per year. It is vital to consider the long-term effects of these particles being deposited in the body of a patient, given the fact that life expectancy is increasing in the Western world and the average age for total joint replacement (TJR) is decreasing.
α-Alumina is generally seen as a bioinert material (7), although studies of the biological response to its wear particles suggest that exposure to relatively high volumetric doses of this “bioinert” material reduces cell viability (3, 8). It may be naïve to use cytotoxicity tests in isolation in evaluating biocompatibility especially in the case of nanoparticles, which uniquely have access to subcellular machinery including the nuclear machinery and as such could potentially induce some very subtle effects that would not manifest in cytotoxicity tests. Cytokine release studies have shown that exposure of cells to both microparticles and nanoparticles of alumina causes stimulation of cytokines at particle concentrations that have been seen to have a relatively small effect on the cell viability (9-10-11). Histological evidence from retrieved periprosthetic tissue shows that single cell line cytotoxicity assays do not stimulate the same biological response as that seen in vivo (12).
The role of proteins adsorbed to biomaterials as mediators of the biological response to implanted devices has long been accepted. However, in the case of wear-induced nanoparticles generated in vivo, this role of biomolecule mediation with the cells has not been documented or considered fully as yet (13, 14). It is well-known that nanoparticles, due to their size, possess physical and chemical characteristics quite distinct from those of the chemically equivalent bulk material (15). Furthermore, it is becoming increasingly accepted that the layer of adsorbed proteins influence nanoparticle fate and behavior in the body (16-17-18-19-20-21-22-23), and as such, protein adsorption and the nanoparticle protein corona have been put forward as an important determinant in nanotoxicity (14, 16, 24, 25). Long-term effects of nanoparticles are very hard to simulate using in vitro tests; however, characterization of the proteins adsorbed is much more accessible and may provide one way of predicting long-term effects.
Cedervall et al suggested that the effective unit of interest is the particle-protein complex, the “corona” of more or less strongly bound proteins, rather than the nanoparticle itself, and that an epitope map of the outer surface that interacts with cells is needed for a nanoparticle in order to be able to assess its true bioactivity (15). This work has been further confirmed by Casals et al, who observed the dynamic nature of the protein corona evolving from a soft lightly bound transient layer to a “hard,” dense corona that is more stable and will ultimately confer on the nanoparticles their effective biological character (20, 26). Studies have tried to theoretically model the solid–liquid interface to understand the adsorption behavior of inorganic nanoparticles (26, 27), and the present work aimed to provide empirical data for the determination of the protein corona for nanoparticles that are commonly introduced into the body over the lifetime of an arthroplastic implant.
Studies to investigate the protein adsorption patterns for alumina particles exposed to complex fluids such as human serum/plasma have, until now, used micron-sized particles, which cannot be said to be representative of the wear debris in vivo from hard-on-hard implants. In 1998, a study by Takami et al found that alumina had a prominent adsorption band corresponding to albumin (28). Rosengren et al found that 40-80-μm alumina and zirconia displayed low protein binding capacities in general, and almost the same adsorption affinities for albumin (29). Sun et al revealed a more complex protein adsorption pattern using 1-10 μm particles and also found that the adsorption of albumin directly enhanced the production of inflammatory cytokines TNF-α, IL-1β and IL-6 (30). Milleding et al studied the protein adsorption from plasma biofilms onto dental ceramics and found that for all materials, albumin was adsorbed in clearly detectable amounts (31).
These studies all employed particles of sizes of a micron and larger. Nanotoxicity studies show that one must consider nanoparticles of a material as different entities from their bulk counterpart (32, 33). Therefore, given the evidence that the wear particles produced in vivo are nanoparticles, there is an onus on the research community to assess size-dependent properties, and to use particle size as a primary criterion for determining implant safety (32). The present study constituted a step toward this goal, and we assessed the differences in plasma protein adsorption for clinically relevant nanoparticulate and monolithic alumina.
Materials and Methods
Materials
Alumina Nanoparticles
The materials used for this study were purchased from Nanostructured and Amorphous Materials Inc. (Los Alamos, NM, USA). The materials used were 99.95% 11-nm α-alumina, which is representative of wear particles seen in vivo and in wear simulators (4), and 99% 30-40-nm γ-alumina. γ-Alumina was used in this study to compare the protein adsorption patterns for different polymorphs of chemically identical species, to examine if the protein adsorption patterns would be similar for similar-sized particles with different internal crystal structures – i.e., if size and the resultant relative geometries of the particles are the dominant determinant of the specificity of protein binding. For all tests performed, the materials were first heated to 500°C for 4 hours to remove any possible organic contaminants from the surface of the materials.
Monolithic Sample
A monolithic sample of α-alumina was fabricated from the nanoparticles, using traditional pressing and sintering routes, without the use of a binder or sintering aid in order to ensure that the composition of the monolith was identical to that of the nanoparticles, to ensure comparability of samples was not affected by additives needed to improve monolithic mechanical strength. The final sintering temperature was 1600°C. The α-alumina sample was pressed using a hydraulic press, bringing the die pressure to 2.1 GPa, and was sintered using the UAF 17/4 furnace (Lenton, London, UK) with a ramp rate of 10°C/min to 1600°C and held for 480 minutes, and then cooled at a ramp rate of 10°C/min to 500°C, and finally air cooled. The Archimedes principle was used to determine the sintered density and was found to be 83% ± 1.5% of theoretical density. To allow the monolithic sample to fit into a 2-mL microcentrifuge tube, the samples were then cut using a diamond-coated blade using a precision saw Isomet 2000 (Buehler Ltd., USA). It was the cut monolith that was used for all relevant preparation and testing in the studies described below.
Monolithic samples were then ground using silicon carbide paper (Struers Ltd., UK) of 1200 and 1200/4000 grit followed by oxide grinding using 0.4-μm colloidal silica (MetPrep, UK), they were then polished by the α-alumina nanoparticles in suspension and finally finished by rubbing on cartridge paper. The samples were cleaned with water followed by iso-propanol, and finally acetone, and then subjected to heat treatment at 500°C for 4 hours to ensure that any organics were removed.
A monolithic sample of γ-alumina was not fabricated given that it is not used as a material in implant design, the sintering routes are complicated and expensive, and the nanoparticles are only of interest as a means of comparison of the proteins adsorbed to this crystallographic polymorph of alumina.
Characterization of the Nanoparticles
The nanoparticle sizes were characterized by the following complementary methods.
Electron Microscopy
Transmission electron microscopy (TEM; JEOL-2000; JEOL Ltd.), was used to confirm that the sizes of the individual particles were as the manufacturer stated. The particles were prepared as solutions of 2.5 wt% powder in distilled and filtered water, which were dispersed using the adaptive focused acoustic (AFA) dispersion technique (Covaris S220; Covaris Inc., USA). The sample was subsequently further diluted to 1,000th part solution using distilled and filtered water. Ten microliters of the solution was then placed on a formvar-coated copper TEM grid (Electron Microscopy Sciences, Hatfield PA, USA) and allowed to dry overnight.
Scanning electron microscopy was performed to image the monolithic sample of alumina using a JEOL-5410 (JEOL Ltd.). Samples were sputter coated with gold prior to examination.
N2-BET Adsorption
Nitrogen physisorption (Quantachrome, Nova 2200e) was used to measure the surface area of both the monolithic and nanoparticle samples and to confirm that the monolithic sample did not present nanofeatures due to imperfect densification of the monolith as a result of not using a binder or sintering aid (in order not to introduce additional substances which would affect the protein binding). Samples were degassed for 3 hours at 300°C to ensure that all water and organic contaminants were removed before measurements were taken. Adsorption isotherms were then obtained and the BET method was used to calculate the surface area of the samples.
The measurement was repeated for each sample to achieve statistically significant results.
Size Analysis of Particles in Media
The particle sizes were also investigated in sample buffer (30 mM phosphate buffer; pH 7.45; containing 1 mM ethylenediaminetetraacetic acid [EDTA]) by different methods: light-scattering nanoparticle tracking analysis (LSNTA; NanoSight Inc., UK), dynamic light scattering (DLS; Zetasizer Nano Series; Malvern Instruments) and an analytical centrifugation system (differential centrifugal sedimentation [DCS]; DC 24000; CPS Instruments Inc., USA). Initially, the particles were dispersed in phosphate buffer saline to mimic the test media for the adsorption studies – i.e., human plasma. However, the particles were seen to flocculate very quickly on visual inspection, even following intense dispersion using AFA to disperse the particles. The phosphate buffer saline was altered, by increasing the phosphate concentration and removing the NaCl, forming the sample buffer: 30 mM of phosphate buffer, pH 7.45, containing 1 mM EDTA. EDTA was added to the buffer to prevent the clotting mechanism in plasma. The sample buffer was filtered through 0.2-μm filters (Whatman Ltd., UK) before use.
Light-Scattering Nanoparticle Tracking Analysis
LSNTA was performed on the nanoparticles in ultrapure water. The concentration of the particles in the solutions was initially 5 mg/mL prior to AFA dispersion, and the solution was then diluted by 102 before being injected into the measurement instrument. Every sample was measured (n = 10), and the averaged 95% range is shown in results.
Dynamic Light Scattering
DLS was performed on the nanoparticles in both ultrapure water and the sample buffer (30 mM phosphate buffer, pH 7.45, containing 1 mM EDTA). The concentration of the particles in the solutions was initially 5 mg/mL for AFA dispersion, and the solution was diluted by placing 50 μL of the solution in 1.5 mL of sample buffer. This solution was then injected into a measurement cuvette, and DLS was performed.
Differential Centrifugal Sedimentation
The alumina nanoparticles were characterized regarding their size distribution in sample buffer using an analytical centrifugation system for DCS (DC 24000, CPS Instruments Inc., USA). Sucrose gradient, 8-24 w/w%, in sample buffer was used. The centrifuge was run at 4.5 kRPM. The particle concentration was 1 mg/mL.
ζ-Potential
The particles were also characterized in terms of their surface charge (ζ-potential) in sample buffer using a Malvern Zetasizer Nano (Malvern Instruments Ltd., UK). The samples were the same as those used for DLS.
Cytotoxicity
Cytotoxicity tests were performed using the lactate dehydrogenase (LDH) assay (cell membrane viability) and the adenosine triphosphate (ATP) assays (cell viability) on A549 carcinomic human alveolar basal epithelial cells (LGC Prochem, UK). This cell line was chosen as they are being used as a standard cell line in our group's other investigations into nanoparticle interactions and allowed comparison beyond the present study. The cells were seeded in a 96-well plate with minimal essential media, and allowed to reach approximately 70% confluence for 24 hours at 37°C. The particle concentrations investigated were approximately 10-600 μg/mL for the γ-alumina and 8-500 µg/mL for the α-alumina (with each step halving the previous concentration). Before the assays were carried out, the particles were subjected to dialysis 4 times for 4 hours each in 5 L of distilled and filtered water to remove any potential surface contaminants that could interfere with the accuracy of the results. The concentration of particles was then determined by lyophilizing a sample of the particle dispersion. Cells were incubated with the particles for a further 24 hours at 37°C. LDH assay was performed using the Promega CytoTox-ONE Homogeneous Membrane Integrity Assay kit. Cell membrane permeability after particle treatment was assessed in triplicate in 96-well plates by measurement of the activity of LDH released from the cytosol of damaged cells into the medium. Data were expressed as percentage of LDH released from cells versus LDH released from cells lysed using a detergent treatment. The LDH was detected using fluorescence, with the reaction taking place in black 96-well plates. The ATP assay was performed using the Promega CellTiterGLO Luminescent Cell Viability Assay kit. Cellular metabolism, which is directly correlated to cell proliferation, after particle treatment was assessed in triplicate in 96-well plates by a luciferase reaction with ATP present in the media of lysed cells. Data were expressed as percentage of ATP present in the media of treated cells versus ATP present in the media of untreated cells. The luminescence was detected in opaque 96-well plates, using a high-throughput microplate spectrophotometer (SpectraMax Plus384; Molecular Devices, USA). For both assays, potential interference of the nanoparticles with the reporter dyes was assessed by determining the fluorescence in the presence of particles in the absence of cells, and could be ruled out as a contributing factor.
Protein Adsorption
The hard protein coronas that formed around the nanoparticle and monolith samples were determined by mixing particles (suspended in sample buffer) with human plasma.
Nanoparticles (0.4 mL) dispersed in sample buffer were mixed with 0.4 mL human plasma. The concentrations of the nanoparticle solutions were 10 mg/mL for the α-alumina and 3.75 mg/mL for the γ-alumina (the difference in concentration was made to normalize for the surface areas of the samples given by BET due to the difference in nominal particle size as in the section ”Alumina nanoparticles”), the monolithic sample was put in 0.4 mL of the sample buffer mixed with 0.4 mL human plasma. All samples were incubated on ice for 1 hour, centrifuged for 3 minutes at 16.1 kRCF (at 4°C) followed by aspiration of the supernatant (discarded); the pellet was washed with 1 mL of the sample buffer and transferred to a new vessel. The washing step was repeated 3 times. The completeness of the washing steps was checked by running the supernatants from the washing step on a sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) gel, and ensuring that the final wash contained no detectible protein. Twenty microliters (25 μL) of SDS-loading buffer (10 w/v% SDS; 10 v/v% β-mercaptoethanol; 20 w/v% glycerol; in 125 mM Tris-HCl, pH 6.8) was added to the nanoparticle pellet and monolithic samples and heated to 99°C for 5 minutes to elute the bound proteins from the alumina surface and to denature the proteins. The proteins constituting the hard protein corona around the 3 different samples were then separated using 10% 1-dimensional SDS-PAGE (1-D SDS-PAGE). To visualize the proteins, the gel was Coomassie stained (using filtered 2.5 g Coomassie Brilliant Blue G250 in 100 mL acetic acid, 400 mL ethanol and 500 mL MilliQ H2O) and distained to remove the excess dye (using 100 mL acetic acid, 400 mL ethanol and 500 mL MilliQ H2O). SDS-PAGE gels were repeated 5 times. Gels were scanned using a GS-800 Biorad scanner, and gel bands densitometry was obtained by Image J.
Relevant bands of interest, indicated in Figure 3a, have been excised and analyzed by mass spectrometry, followed by in-gel trypsin digestion. Trypsinized peptides were re-suspended in 0.1% formic acid and analyzed by electrospray liquid chromatography mass spectrometry (LC MS/MS) using high-performance liquid chromatography (HPLC; Surveyor; ThermoFinnigan, CA, USA) interfaced with a linear trap quadrupole (LTQ) ion trap mass spectrometer (ThermoFinnigan, CA, USA). Chromatography buffer solutions (buffer A, 0.1% formic acid; buffer B, 100% acetonitrile and 0.1% formic acid) were used to deliver a 72-minute gradient (5-minute sample loading, 32 minutes to 40% buffer B, 2 minutes to 80%, hold 11 minutes, 1 minute to 0%, hold for 20 minutes, 1 minutes of flow adjusting).
A flow rate of 150 μl/min was used at the electrospray source. Spectra were searched with Sequest using the Uniprot/Swiss-Prot database (http://www.expasy.org) automated by the Proline software platform (http://www.biontrack.com). Peptide and ProteinProphet where used to validate protein hits, and a probability cutoff of 0.9 was applied to filter out false positives. If 1 protein hit was identified by a single unique peptide, the spectrum was manually interpreted to check the quality of the spectra.
Semiquantitative assessment, based on spectral counting (SpC) which represents the total number of the MS/MS spectra for all peptides attributed to a matched protein, was also performed. This semiquantitative approach, while not giving absolute values for specific protein content, does allow reasonable comparison of protein abundance between samples. The spectral count for each protein was normalized to the molecular weight (MW), to take into account the protein size, and is expressed as the normalized spectral count (NSpC): percentage of the total protein amount of the corona of each sample, as described previously (17). NSpC thus correlated to the MW and relative protein abundance. This analysis was carried out only once, which is deemed sufficient for semiquantitative reporting (17).
Results
Nanoparticle Characterization
Scanning electron microscopy (SEM) imaging was performed on the monolith sample, where significant grain growth did occur and the sample features were not on the nanoscale when compared with the particles (see Fig. 1). Even though the pictures show that the monolith was porous (as no sintering aids or binders were used in order to preserve the chemical purity of the samples), the grain size was greater than 1 μm and could be considered as representative of the bulk material (see Fig. 1, panel 1). TEM images of α- and γ-alumina (Fig. 1, panels 2 and 3, respectively) confirmed that the presence of nanoparticle units as per manufacturer's specifications; however, the presence of clusters indicated particle agglomeration. Surface areas of the particles used were calculated by N2-BET adsorption. Surface area of the monolith measured 0.851 m2/g, while the surface areas of α- and γ-alumina were 3 orders of magnitudes larger (118 m2/g and 236 m2/g) due to their smaller size.

Panel 1 shows a scanning electron microscopy (SEM) image of an α-alumina monolithic sample (scale bar 2 μm). Panel 2 shows a transmission electron microscopy (TEM) image of the α-alumina sample (scale bar 100 nm). Panel 3 shows a TEM image of the γ-alumina sample (scale bar 50 nm).
Nanoparticle dispersion was investigated by LSNTA, DLS and DCS. and the results are summarized in Table I. Due to the instability of the solution, multiple population clusters were detected, indicating nanoparticle instability in the dispersant media.
Size and ζ-potential values for the α- and γ-alumina nanoparticles in sample buffer
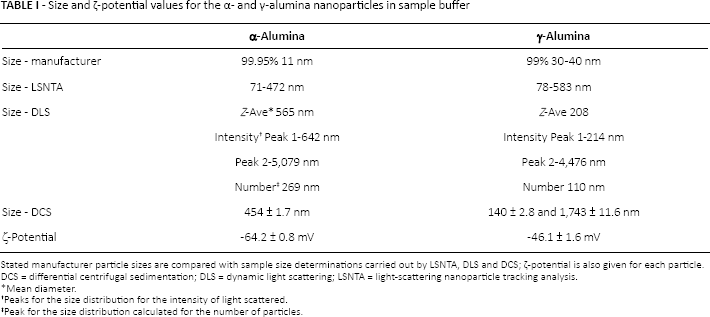

Stated manufacturer particle sizes are compared with sample size determinations carried out by LSNTA, DLS and DCS; ζ-potential is also given for each particle.
DCS = differential centrifugal sedimentation; DLS = dynamic light scattering; LSNTA = light-scattering nanoparticle tracking analysis.
Mean diameter.
Peaks for the size distribution for the intensity of light scattered.
Peak for the size distribution calculated for the number of particles.
Cytotoxicity
The results from the cytotoxicity assays are shown in Figure 2. The LDH and ATP assays showed that the alumina nanoparticles did not induce a toxic effect for concentrations up to 500 μg/mL and 600 μg/mL for the α- and γ-alumina, respectively.
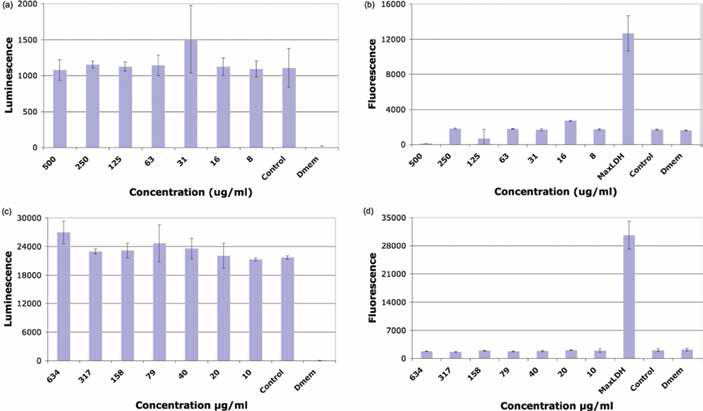
a, b) The adenosine triphosphate (ATP) and lactate dehydrogenase (LDH) cell assays for A549 cell exposed α-alumina nanoparticles, respectively; c, d) the ATP and LDH cell assays for A549 cell exposed γ-alumina nanoparticles, respectively. Numbers on the
Protein Adsorption
Figure 3a shows a representative 1-D SDS-PAGE gel with the marked sections that were excised for proteomic analysis. Figure 3b shows analysis of the pattern in Figure 3a using ImageJ analysis to highlight the clear differences in the patterns in the gels.
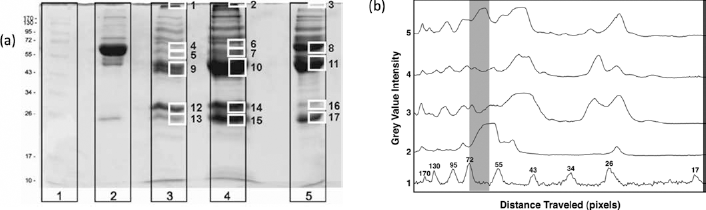
a) 1-D SDS-PAGE for nanoparticles of α- and γ-alumina and monolithic α-alumina exposed to human plasma with numbered markers of those parts of the gel excised for proteomic analysis. Lanes: 1. molecular weight standard, 2. plasma (diluted 10 times in phosphate-buffered saline), 3. α-alumina nanoparticles, 4. γ -alumina nanoparticles and 5. α-alumina monolith. b) Analysis with ImageJ of the gel in panel (a), graphically illustrating the difference in the protein adsorption patterns. Boxed gray area approximately outlines the expected band for albumin. Curves: 1. molecula weight standard, 2. plasma, 3. γ-alumina nanoparticles, 4. α-alumina nanoparticles and 5. α-alumina monolith.
TABLES II and III show the most pertinent results from the proteomic analysis (the full proteomic information can be requested through correspondence with the authors).
Discussion
The TEM images show that the nanoparticles formed aggregates. These aggregates could have been formed when water evaporated during the drying of the sample on the TEM grid, which is common when preparing TEM samples of particles. However, based on comparison of the size measurements using other techniques (see the section “Size analysis of particles in media” and Tab. I), there is obviously a significant aggregation tendency for the particles, likely as a result of the initial sample preparation which resulted in a dried powder that is difficult to re-disperse into a monodispersed solution. The fact that the particles do not act as individual particles once suspended in media raises the question as to whether the wear-induced nanoparticles form similar clusters in vivo. This possibility is reinforced by the identification of aggregates in histology from wear particle studies, as described in the “Introduction” (6, 10, 12).
The cytotoxicity studies demonstrate that the alumina nanoparticles do not induce a cytotoxic response. It is clear that we have used a cell line that would not typically be used in device-related cytotoxicity assays. However, this cell line has been used extensively in other studies evaluating cellular toxicity in response to exposure to nanoparticles (34, 35). The current study was one of the first studies to evaluate the protein adsorption profiles for these materials. For this reason, it was decided that the A549 would be used, to enable comparison of these novel findings with other nanoparticles studies currently available (34, 35).
The mass spectrometry analysis revealed striking differences in the coronas formed on the 3 different test samples (γ- and α-alumina nanoparticles and the α-alumina monolith), suggesting that surface charge, crystal form and size all have a fundamental role in determining the protein binding. The identities and abundances of the proteins determined to be in the various coronas are shown in Table II, and these proteins have been grouped according to protein function as shown in Table III. The first striking difference in the hard protein coronas formed relates to the lipoprotein and in particular to apolipoprotein B-100. Apolipoprotein B-100 is the major protein component in low-density lipoprotein (LDL) complexes. This protein was detected with over 3.15% and 3.88% NSpC in the corona of the γ-alumina nanoparticles and of the α-alumina monolith, while its detection was minimal in the corona of the α-alumina nanoparticles, as shown in Table II. This result indicates that LDL has a lower affinity for the α-alumina nanoparticles compared with the other 2 samples. The observed difference between the 2 α-alumina samples (nanoparticle and monolith) is intriguing. One possible explanation is that the different surface curvature presented by the α-alumina nanoparticles when compared with the α-alumina monolith, directly affect LDL's interaction with the surface. The influence of surface curvature on protein adsorption and protein corona composition has been highlighted before (36, 37). The α-alumina monolith will appear as an almost flat surface at the nanoscale, according to the SEM and N2-BET data for the LDL body (see Fig. 1, panel 1), while the α-alumina nanoparticles will display high surface area aggregates of ∼400 nm (see Tab. I) constructed from particles of 30-40 nm. The aggregates of the α-alumina nanoparticles will display a completely different surface geometrically compared with the α-alumina monolith, with cavities of the same order of size as the LDL complexes (these are particles which themselves have diameters of 22-29 nm (38)) and local surface curvature that is comparable to the curvature of LDL (see Fig. 1). This suggests that there may be some steric or packing hindrance to LDL interaction with the α-alumina nanoparticles. The difference observed between the LDL interaction with α-alumina and γ-alumina nanoparticles may arise from the difference in crystallographic phase or from the difference of the surface presented by the aggregates. The γ-alumina aggregates, since they are made up of 11-nm particles, will present a much smoother surface with cavities that are less deep and wide compared with those of the α-alumina nanoparticles.
Proteins identified using mass spectrometry in the coronas associated with the various alumina samples studied
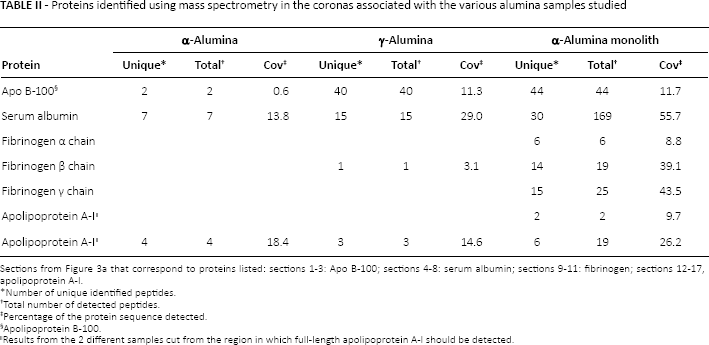
Sections from Figure 3a that correspond to proteins listed: sections 1-3: Apo B-100; sections 4-8: serum albumin; sections 9-11: fibrinogen; sections 12-17, apolipoprotein A-I.
Number of unique identified peptides.
Total number of detected peptides.
Percentage of the protein sequence detected.
Apolipoprotein B-100.
Results from the 2 different samples cut from the region in which full-length apolipoprotein A-I should be detected.
Proteins identified in coronas of the 3 samples, grouped by function and reported as normalized spectral count (NSpC)
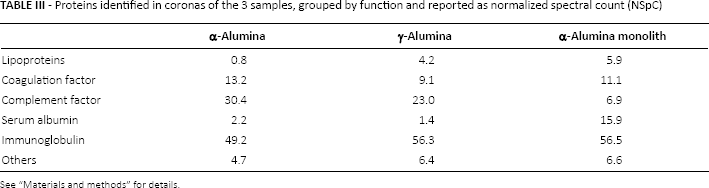
See “Materials and methods” for details.
The observed differences when it comes to detected apolipoprotein B-100 should be further investigated, as this protein is a known transporter protein, and depending on the details of the surface structure of wear-induced particles in vivo, it could potentially result in transport of these particles across biological barriers, resulting in bioaccumulation (39).
A second major difference between the nanoscale and bulk samples was in the amount of adsorbed serum albumin. As already mentioned, serum albumin is by far the most abundant protein in plasma, with a concentration of ∼40 mg/mL. The data in Figure 3 indicate that the α-alumina monolith has a significantly stronger adsorption band in the section of the gel in which serum albumin should be found, compared with either of the nanoscale samples. The proteomic data corroborate this observation. For the monolith, serum albumin was identified with 15% of the NSpC. For α- and γ-alumina nanoparticles serum albumin was only identified with 2.2% and 1.4% peptides, respectively. To be certain that the whole gel area, in which full-length serum albumin should be found, was investigated, a larger gel area was excised (2 samples were excised edge to edge) for both the α- and γ-alumina nanoparticles (see Fig. 3a). The serum albumin adsorption to comparatively flat alumina surfaces has been reported before (40). However, these results showed that there was a significant difference for the adsorption of serum albumin to macro surfaces (the monolith) compared with nanosurfaces.
A third region investigated by proteomics is the region in which fibrinogen binds differently to the alumina nanoparticles relative to the monolith sample. Fibrinogen is detected as part of the hard corona around the α-alumina monolith where the sum of the NSpCs of the α, β, and γ subunits was 6.96%, while in α- and γ-alumina nanoparticles, it represented 0.38% and 0.31%, respectively, suggesting an influence of the surface geometry and curvature on fibrinogen binding (see Tab. II). The results again show significant differences between the monolith and the nanoparticles regarding the composition of the major proteins in the hard protein corona formed following incubation with plasma.
The last site investigated with mass spectrometry was the region in which the presence of full-length apolipoprotein A-I would be detected. Apolipoprotein A-I is the major protein in high-density lipoprotein (HDL) and has been detected as one of the more common proteins found in the hard protein corona around several tested nanoparticles of significantly different composition (15, 37, 41-42-43-44-45-46-47-48). When examining this region, a larger part of the gel was excised into 2 samples for each lane (see Fig. 3a) to be certain not to miss the right MW region in which apolipoprotein A-I would be detected. As can be seen in Figure 3a, this region has a dense and a less dense band for the α-alumina nanoparticles, 2 dense bands for the γ-alumina, and a less dense and a dense band for the α-alumina monolith (with relative densities of the higher MW mentioned first in this comparison). The appearance of the gel indicates that there should be some significant differences here. However, there are no major distinctions when it comes to apolipoprotein A-I, based on the mass spectrometry data in Table II. Apolipoprotein A-I was detected in similar quantities in all of the lower MW samples for the α- and γ-alumina nanoparticles and the α-alumina monolith. It was also detected in the higher MW sample for the monolith. These numbers could indicate a difference again between the monolith and the 2 nanoparticles, however, the detected sequence coverage did not differ much between the different samples, making it hard to definitively interpret the data.
Complement factor proteins, a family of proteins involved in the opsonization (recognition of foreign objects by macrophages) and innate immune system, were also identified in the protein coronas of the nanoscale particles used in the study. In particular, while the α-alumina monolith showed low protein affinity (6% NSpC), α- and γ-alumina nanoparticles both showed a significantly higher affinity for complement factor proteins, with 30.2% and 22.9% NSpC, respectively.
It is difficult to definitively say what the difference in the affinity of complement factor proteins between the nanoscale and monolithic alumina samples means, given the numerous biological pathways that these proteins are involved in. Furthermore, we are dealing here with wear particles in human plasma in vitro and extrapolating the results to what may happen in vivo in the periprosthetic tissue following generation of wear particles. However, it is not unreasonable to infer that, given that previous researchers have shown opsonization of wear particles leading to phagocytosis and subsequent cell lysis, the absorption of complement proteins on the alumina nanoparticles may contribute to the overall aseptic inflammatory phenomenon seen in periprosthetic tissue (10, 12, 30, 49-50-51). The results shown here may reveal the reason a bioinert material in its bulk form does not result in an inflammatory reaction, whereas nanoparticles released from this bulk material as a result of wear may. The reason for this difference is not size alone but rather the nature of the protein corona that forms around the nanoparticle, which, in turn, will be a result of the size. The differing protein corona may be the underlying reason as to why wear particulates cause the inflammatory reaction that has been observed in vivo.
It is always difficult to draw conclusions for in vivo performance on the basis of in vitro results, and there are, of course, some clear factors that could have affected the outcome, such as the fact that the plasma proteins were incubated on ice, at 4°C, rather than at the more clinically relevant temperature of 37°C. However, the profile of the proteins adsorbed to the monolith was comparable to that found in the literature. Also, given that the biological fluid used in this study was human plasma, incubation at the lower temperature reduced to a minimum any enzymatic activity that could result in blood coagulation (52). Additionally previous experiments have been performed at 4°C where the protein corona evolution has been evaluated over time, and the present results may be directly compared with those data in order to obtain a more complete picture of nanoparticle proteomics across related systems (53). A desirable extension to the current work would be a time-resolved study with protein inclubation at room temperature and at 37°C and with other biological media.
The variations in the compositions of the hard protein corona for the limited MW regions that have been investigated by proteomics (mentioned in Section Protein Adsorption) are illustrative of the main differences seen in the protein binding patterns. In all of the samples cut from the 1D SDS-PAGE gels, immunoglobulin was detected, indicating that immunoglobulin is a major component of the corona for both the nanoscale and monolithic α-alumina materials.
Conclusions
The main findings from this study are simple: there are clear differences regarding which proteins adsorb to nanosurfaces compared with macrosurfaces for the same type of material. There are a range of possible explanations for this; some of which have been suggested in previous discussions (14, 15, 54), and some of which are still the subject of much debate. What is clear is that the size of a foreign object such as a wear particle from an orthopedic device is a critical factor in determining its apparent biological character, in addition to the chemical composition and the polymorph of the material. The difference in character between the bulk material and the nanoparticle is pertinent for all aspects of bioengineering and drug delivery, since if the material changes physical form during its life in vivo, it will inherently change its biological character, and previous predictions of its trajectory and interaction in vivo will be compromised. In the case of alumina, this appears to be very important when trying to understand the biological effect of the clinically relevant wear particles as opposed to alumina in its bulk implant form. From the results shown here, one could predict the behavior of the nanoparticle in vivo based on the relatively greater quantity of opsonins adsorbed in the nanoparticulate form. It would be reasonable to infer that, given the differences in the proteins adsorbed between the nanoparticles and the monolithic sample, the nanoparticles are more likely to induce an inflammatory reaction, including release of cytokines, which has been reported by several authors for wear-induced particles (30, 51). It is possible that this nanoparticle-induced inflammation may be what leads to aseptic loosening, since phagocytosis would be initiated, which ultimately can lead to cell lysis, as the phagocytes are unable to digest the nanoparticles, and a release of cytokines resulting in a cascade of cell signaling will lead to osteolysis and thus aseptic loosening (49, 50).
The findings here suggest that all materials that have the potential of presenting as nanoparticles should have their hard protein corona determined, as we cannot rely solely on the previously characterized protein adsorption profile determined for macrosurfaces. Finally, these results indicate that simple cytotoxicity tests may not be sufficient to detect subtle biological effects, such as an increased tendency for phagocytosis as a result of adsorption of opsonins at the wear particle surface. This leads to a complicated process resulting ultimately in aseptic loosening, due to a change in the geometry of the biomaterial presented to the body as materials become nanoscale. Potentially understanding the impacts of in situ protein binding could lead to improved predictions of long-term effects of wear-induced nanoparticles, allowing a better understanding of the role of nanoparticles in the body and thus enabling the best material choice during implant design, via the safety-by-design approach.
Footnotes
Financial support: This work was supported by the European Union Sixth Framework Program project NanoInteract (grant no. NMP4-CT-2006-033231), the Irish Research Council for Science, Engineering and Technology (M.L.), SFI RFP 06/RFP/ CHP031 (I.M.B.) and an UCD Ad Astra scholarship (J.P.G.). Part of this work was conducted under the framework of the INSPIRE programme, funded by the Irish Government̕s Programme for Research in Third Level Institutions, Cycle 4, National Development Plan 2007-2013 (Éilis McGrath, M.P.M.).
Conflict of interest: The authors certify that they have no affiliations with, or involvement in, any organization or entity with any financial interest (such as honoraria; educational grants; participation in speakers’ bureaus; membership, employment, consultancies, stock ownership, or other equity interest; and expert testimony or patent-licensing arrangements), or any nonfinancial interest (such as personal or professional relationships, affiliations, knowledge or beliefs) in the subject matter or materials discussed in this manuscript.
