Abstract
Background:
Triple-negative breast cancer (TNBC) is the most aggressive subtype of breast cancer, characterized by advanced disease stage and poor prognosis. Moreover, due to the lack of therapeutic markers, TNBC patients can’t benefit fully from currently available targeted therapies.
Methods:
To fully understand the molecular basis of TNBC, we used gene set enrichment analysis (GSEA) to screen out the most altered functional module in TNBC, from publicly available microarray data and studied the association of the candidate gene with TNBC development.
Results:
We found that the proteasome was significantly activated in TNBC. As compared with other breast cancer subtypes and normal tissue, proteasome subunit beta 5 (PSMB5), the key regulator of proteasome function, was overexpressed in TNBC tissue and predictive of poor prognosis. Moreover, we also found that PSMB5 knockdown induced TNBC apoptosis and significantly enhanced cancer cell sensitivity to the chemotherapeutic agents bortezomib and paclitaxel.
Conclusions:
Our results suggest a potential role for PSMB5 as a biomarker and therapeutic target for TNBC.
Introduction
Breast cancer is the most common and life-threatening cancer among women worldwide, affecting 1.7 million women and killing 450,000 patients every year (1). Triple-negative breast cancer (TNBC) is the most aggressive subtype of breast cancer and is associated with larger tumor size, higher histological grade and positive lymph node at diagnosis (2). Due to the lack of the oncogenic markers estrogen receptor (ER), progesterone receptor (PR) and human epidermal growth factor receptor 2 (HER2), current targeted therapies are not applicable to TNBC, and chemotherapy remains the mainstay of treatment (2, 3). However, TNBC patients are prone to develop chemoresistance and are at higher risk of relapse and metastasis.
Over the last decade, great efforts have been made to understand TNBC pathophysiology and develop novel therapeutic targets. Higher mutation levels of the BRCA1/2 genes have been identified in TNBC, and inhibiting poly(ADP-ribose) polymerase (PARP) has shown promise in inducing tumor death in BRCA-defective breast cancer patients (4). Also, increased expression of vascular endothelial growth factor (VEGF) has also been observed in TNBC, and inactivating VEGF function has displayed beneficial effects for metastatic breast cancer patients (5). Moreover, EGFR, mTOR and Src, which are therapeutic targets for other malignancies, are also potential candidates for treating TNBC (6). However, most of the remarkable advances are still at an early phase of clinical trials within small patient cohorts, and their clinical efficacy remains to be determined. Moreover, TNBC is a heterogeneous disease, and a small number of therapeutic strategies would not benefit all patients (7). It is therefore necessary to get a global understanding of the complex basis of the disease and identify the master regulators that could therapeutically be targeted.
To this end, we studied the transcriptome of TNBC and identified a greatly activated proteasome function in TNBC tissue. Notably, we also revealed that highly expressed proteasome subunit beta 5 (PSMB5) was indicative of poor prognosis of TNBC patients, while silencing PSMB5 could sensitize TNBC cells to apoptosis and enhance the effect of chemotherapy. Our result confirmed the importance of the proteasome in tumor development and suggested the potential of PSMB5 as a target for TNBC therapy.
Methods
Tissue samples
The research was carried out in accordance with the World Medical Association’s Declaration of Helsinki. Fresh frozen human tissue samples were collected from surgical procedures performed in the Department of Breast Cancer, the Third Hospital of Nanchang City (Nanchang, China) from January 2000 to July 2003. All investigations described in this study were performed after written informed consent was obtained and in accordance with the guidelines of the local independent ethics committee of the Third Hospital of Nanchang City.
Quantitative real-time PCR
Total RNA was isolated using Trizol reagent (Invitrogen), and was reverse-transcribed to cDNA with a first-strand cDNA system. Synthesized cDNA was amplified with SYBR Green Supermixes (Bio-Rad) in a LightCycler system (Roche). Experiments were performed with the following PCR primers: PSMB5: forward 5’-AGGAACGCATCTCTGTAGCAG-3’, reverse 5’-AGGGCCTCTCTTATCCCAGC-3’; GAPDH: forward 5’-ACAACTTTGGTATCGTGGAAGG-3’, reverse 5’-GCCATCACGCCACAGTTTC-3’.
Relative gene expression was determined with the comparative CT method (2−ΔΔCT method).
Cell culture and PSMB5 silencing
Human TNBC cell lines HCC1143 and BT20 were cultured in RPMI-164 medium and modified Eagle’s medium (MEM), respectively, supplemented with 10% fetal bovine serum (FBS) (HyClone), and incubated at 37°C in a fully humidified incubator with 5% CO2 supply. Small interfering RNA targeting PSMB5 (siPSMB5) and nonsilencing control (siNC) were purchased from Thermo Fisher Scientific and were delivered into cell cultures using Lipofectamine 2000 (Invitrogen).
Proteasome activity assay
Breast tissue lysates were prepared by homogenizing breast samples in lysis buffer containing 50 mM HEPES (pH 7.8), 10 mM NaCl, 1.5 mM MgCl2, 1 mM EDTA, 1 mM EGTA and 250 mM sucrose. Cell line lysates were prepared by scraping cells into lysis buffer. Protein content was determined using the Bradford method (Bio-Rad). Proteasome activity was analyzed in 50-µg protein extracts with Promega Proteasome-Glo Assay.
Apoptosis assay by flow cytometry
Trypsinized cell cultures were washed 3 times and resuspended in Annexin V staining buffer. After double-staining with Annexin V-fluorescein isothiocyanate (FITC) and propidium iodide (PI), cell populations were analyzed in a FACS Calibur Flow Cytometer. Each experiment was performed with 3 replications.
Cell viability assay
Treated and untreated cell cultures were seeded in 96-well plates, and viable cells were detected with a CellTiter-Blue viability assay (Promega) following the manufacturer’s instructions. Each experiment was performed with 5 replicates.
Western blot
Cell cultures were treated with RIPA buffer. Protein extracts were separated in 12.5% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE), transferred onto nitrocellulose membranes (Sigma) and immunoblotted with the antibodies against PSMB5 (Abcam) and actin (Abcam). Protein bands were visualized with enhanced chemiluminescence (Millipore).
Results
Activation of proteasome module in TNBC
Large-scale profiling analysis (e.g., genomics, transcriptomics, proteomics, metabolomics) always provides a valuable resource to uncover the underlying mechanism of disease development and guide further research. To investigate aberrantly expressed genes and functional groups associated with TNBC, we compared the transcriptome between 165 TNBC tissues and 33 adjacent normal tissues from a publicly available microarray dataset (GSE76250) (8), and used gene set enrichment analysis (GSEA) (9) to identify a set of genes involved in the proteasome as one of the most significantly affected gene modules in TNBC (Fig. 1A). Of the 45 genes annotated as proteasome subunits, 32 (71.11%) were significantly up-regulated in TNBC (fold-change >1.2, p<0.001), while none of them were down-regulated (Fig. 1B; and see Supplementary Table I, available online at www.biological-markers.com – Expression changes of the genes coding for proteasome subunits). This result may imply a dramatically activated proteasome function in TNBC.
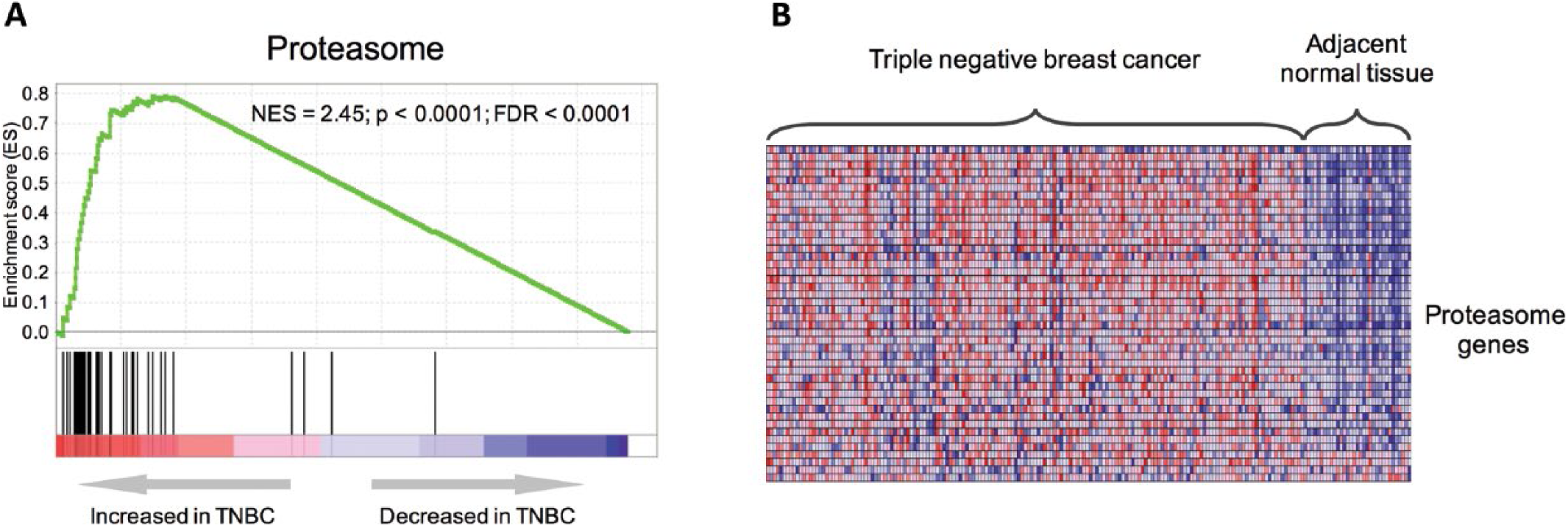
Gene set enrichment analysis (GSEA) plot for the proteasome. (
PSMB5 is up-regulated in TNBC and indicative of poor prognosis
The proteasome plays a key role in degrading 80%~90% of intracellular proteins through ubiquitin-dependent processes, and is responsible for regulating many cellular processes, such as cell cycle, transcription, DNA repair and apoptosis (10). Therefore, the abnormally activated proteasome in TNBC may affect cellular homeostasis and contribute to disease origin and development.
To validate our result, we measured the mRNA expression of PSMB5 within 96 TNBC tissues, 42 adjacent normal (ADNO) tissues as well as 67 non-TNBC tissues collected from surgery procedures (Tab. I). PSMB5 is 1 of the beta subunits in the 20S proteasome core particle, which is responsible for most of the chymotrypsin-like activities in the proteasome. As shown in Figure 2A, PSMB5 was expressed at significantly higher levels in TNBC than in normal tissue and that of other breast cancer types. We also evaluated the chymotrypsin-like activity in 15 TNBC tissues, 7 non-TNBC tissues and 5 ADNO tissues, and identified the strongest activity in TNBC (Fig. 2B). These results demonstrated a largely increased PSMB5 expression in TNBC, which is associated with greatly activated proteasome chymotrypsin-like activity.
Patient characteristics
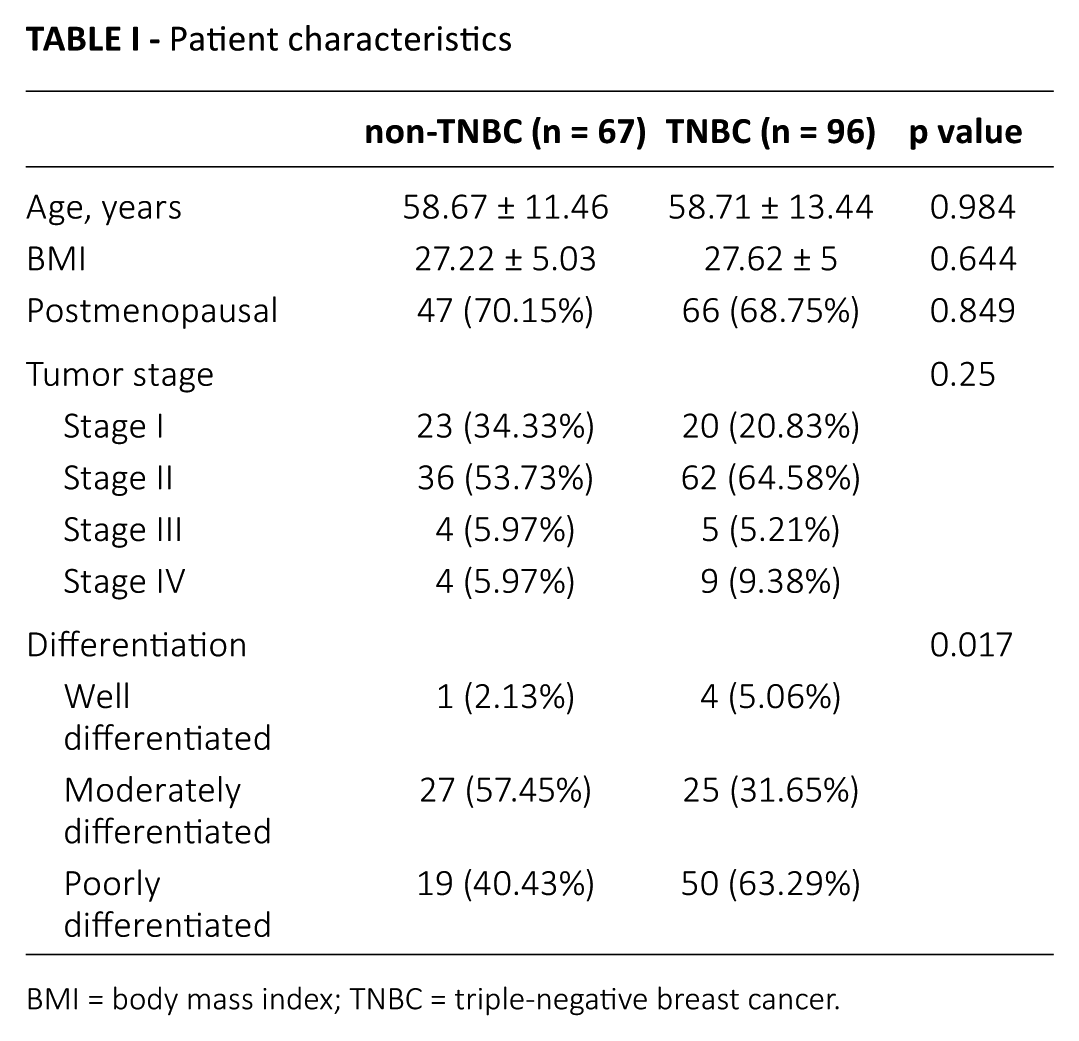
BMI = body mass index; TNBC = triple-negative breast cancer.
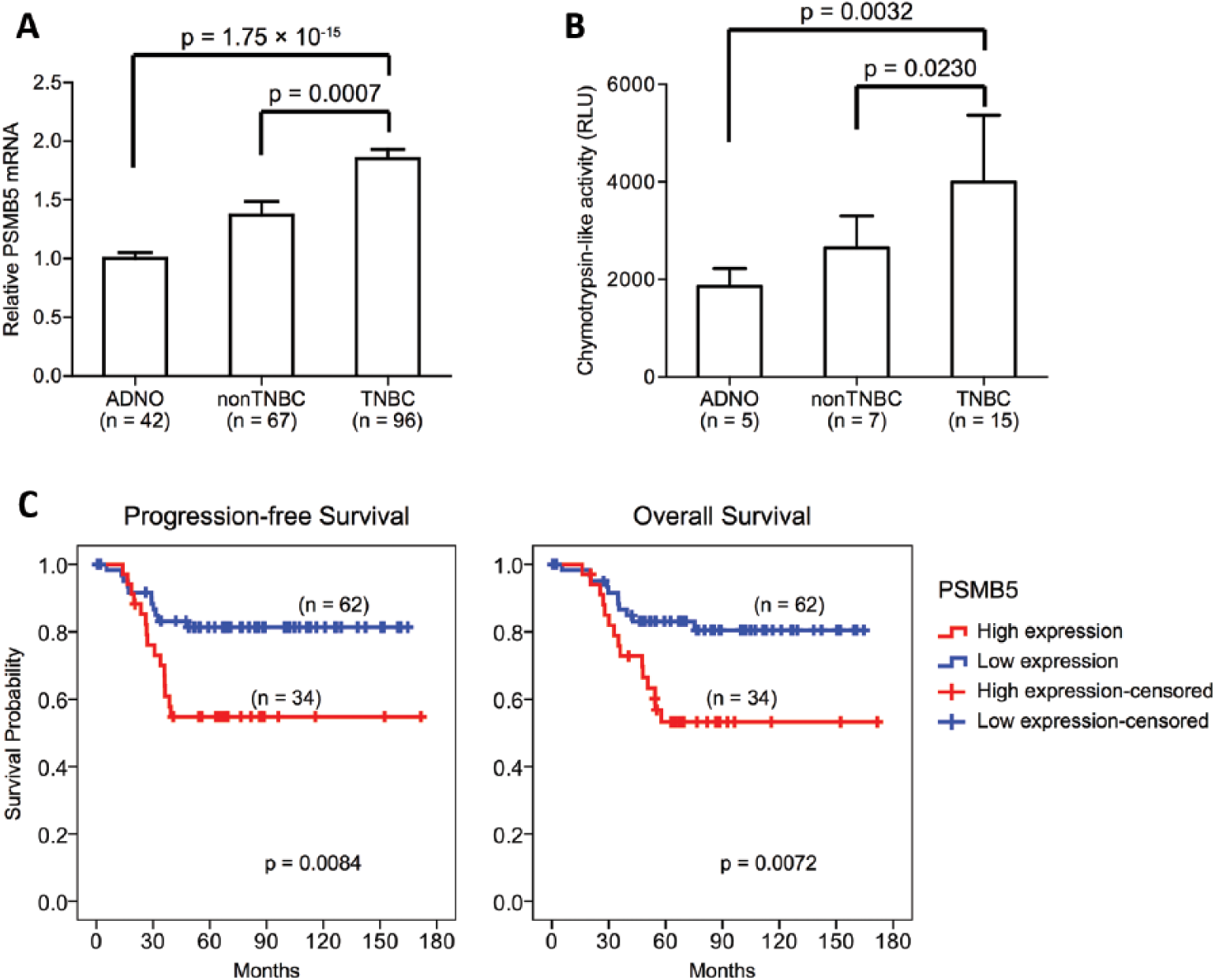
PSMB5 was highly expressed in triple-negative breast cancer (TNBC) patients and associated with increased proteasome function and poor prognosis. PSMB5 mRNA expression (
Since TNBC is recognized to have a poor prognosis, we then asked if PSMB5 could be associated with the survival of TNBC patients. We classified the TNBC patients into 2 groups based on their PSMB5 levels. Patients with PSMB5 levels higher than average were defined as the high PSMB5 group (n = 34) and the others were in the low PSMB5 group (n = 62). As shown in Figure 2C, TNBC patients with relatively lower PSMB5 levels had significantly longer progression-free and overall survival times then the high PSMB5 patients, suggesting PSMB5 was indicative of the severity of the disease.
PSMB5 silencing induces apoptosis in TNBC cells
We then asked if TNBC cell could be eliminated by inhibiting PSMB5 expression. To this end, we transfected siPSMB5 into the human TNBC cell lines HCC1143 and BT20 and identified that successful depletion of PSMB5 resulted in greatly reduced chymotrypsin-like activity, while other proteasome activities, such as trypsin-like and caspase-like activities, were not affected (Fig. 3A, B, E and F). Notably, successful PSMB5 silencing was also associated with the accumulation of cleaved caspase-3, indicating TNBC cell apoptosis (Fig. 3A and E). This result was further confirmed by FACS analysis, which showed that compared with siNC-transfected cells, the number of apoptotic cells was significantly increased in TNBC cells treated with siPSMB5 (Fig. 3C, D, G and H). These results indicated that PSMB5 might function as an oncogene in TNBC.
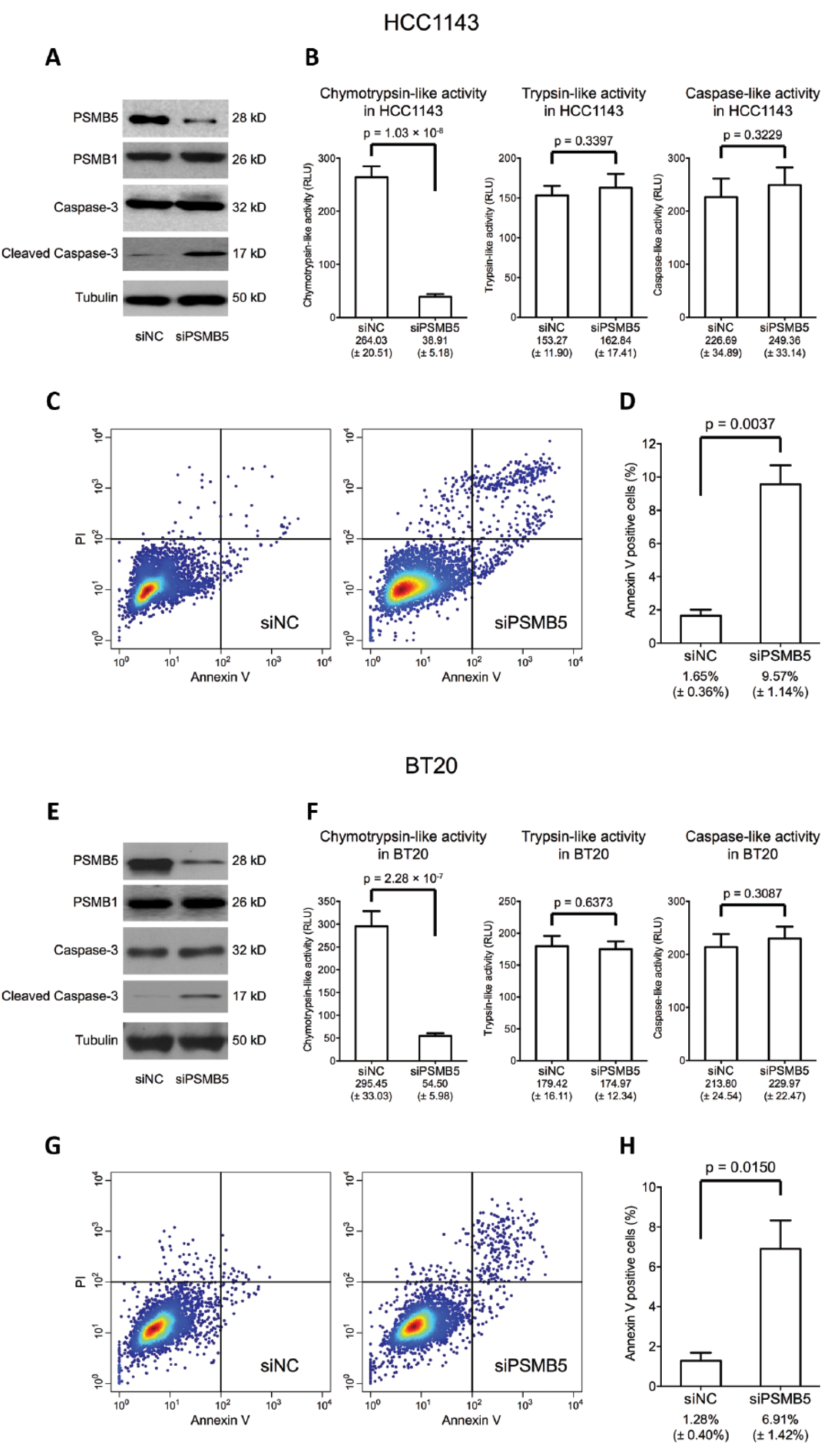
PSMB5 silencing induces apoptosis in triple-negative breast cancer (TNBC) cells. Western blot revealed that successful knockdown of PSMB5 led to caspase-3 cleavage in HCC1143 (
PSMB5 silencing sensitizes TNBC cell to proteasome inhibitor
Currently, there is a broad array of proteasome inhibitors that have been approved for treatment of various cancers in clinical practice, including bortezomib, carfilzomib and marizomib (11-13). As PSMB5 plays a key role in regulating the catalytic functions of proteasome, we thus hypothesized that high expression levels of PSMB5 might be linked to enhanced drug resistance in some TNBC cells. We then investigated the effect of PSMB5 silencing and proteasome inhibitors on TNBC cells by treating HCC1143 and BT20 with siPSMB5 and bortezomib (8 nM) alone and in combination. As shown in Figure 4A, the growth rate of HCC1143 was efficiently suppressed by both bortezomib and siPSMB5 with significant cytotoxicity from 72 to 96 hours, while BT20 had less sensitivity to bortezomib with only moderate cell death at 72 hours. However, PSMB5 silencing greatly improved the effectiveness of bortezomib in both cell lines with cytotoxicity starting as early as 24 hours.
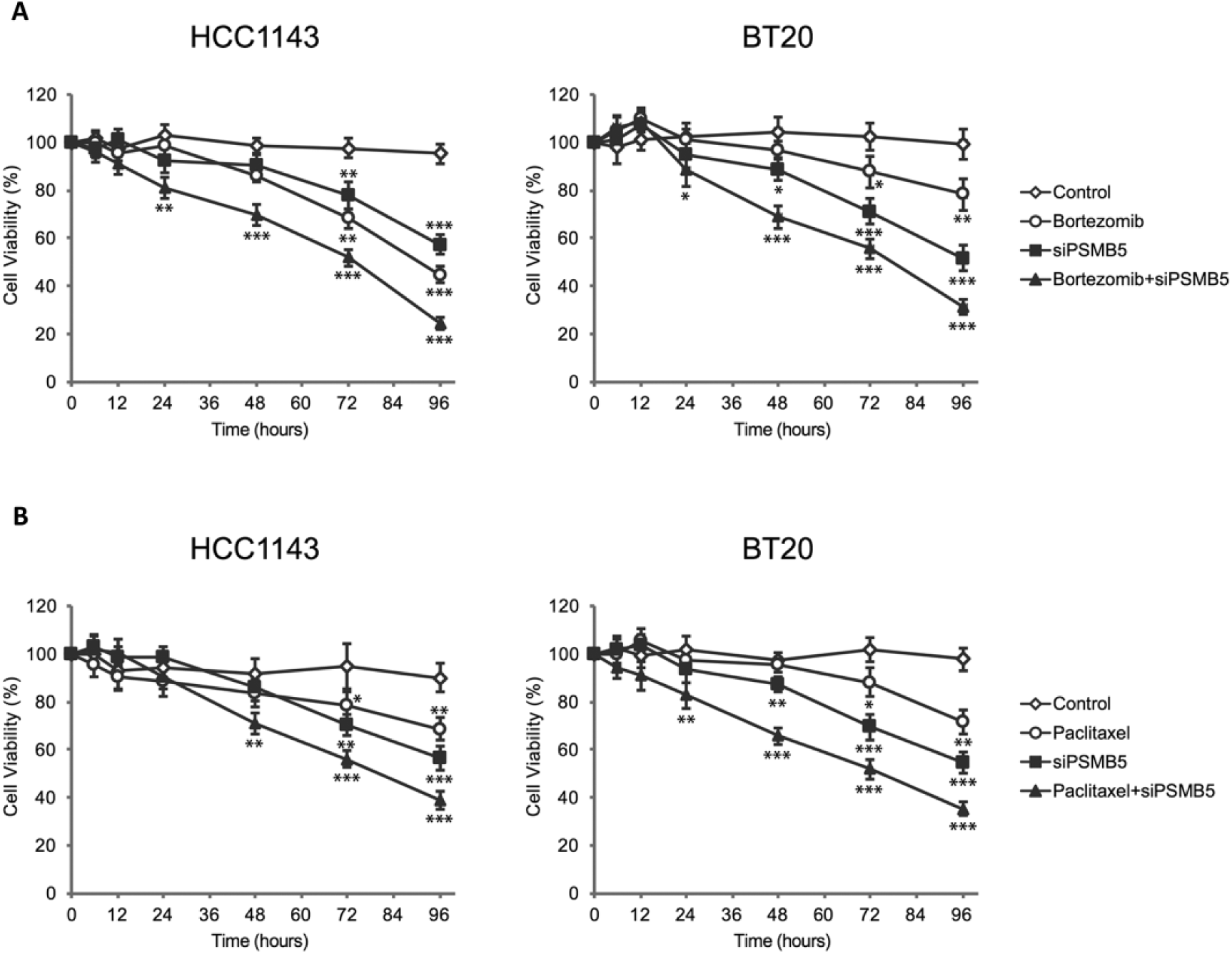
PSMB5 silencing sensitized triple-negative breast cancer (TNBC) cells to chemotherapeutic agents. (
Further, we also tested if PSMB5 could enhance chemotherapeutic agents targeting other cellular functions. Paclitaxel is a cytoskeletal drug and has been widely used for breast cancer treatment. It binds to the beta tubulin subunit of microtubules and protects spindle microtubule polymer from disassembly, which results in cell cycle arrest at the M phase and finally triggers cell apoptosis. As shown in Figure 4B, moderated paclitaxel sensitivity was observed in HCC1143 and BT20 cell lines, while PSMB5 knockdown significantly enhanced the cytotoxicity of paclitaxel with significant cell death identified at 48 hours for HCC1143 and at 24 hours for BT20.
These results provided clear evidence that PSMB5 is a potential target for TNBC therapy alone or in combination with other treatments.
Discussion
In cells, the homeostasis of proteins is determined not only by protein biogenesis, but also by protein degradation. In addition to removing misfolded and damaged proteins, protein degradation is also involved in cellular stress response and signal transduction by regulating the abundance of related proteins. Proteasome-dependent protein degradation undertakes more than 80% intracellular degradation tasks (10). Aberrant proteasome activity therefore disrupts many important cellular processes and leads to a number of diseases, including cancer (14). In the current study, we demonstrated that TNBC patients, especially those with poor prognosis, had an increased expression of proteasome subunit PSMB5. Due to the importance of PSMB5 in regulating proteasome function, the changes of PSMB5 reflected the dysfunction of the whole proteasome complex in TNBC. Since our study was performed in a south-Asian population (China), our results supported a global phenomenon of high expression of PSMB5 and overactivated proteasome function in TNBC. More importantly, we also provided evidence that knockdown of PSMB5 suppressed TNBC cell growth, and we suggest the potential of PSMB5 as a target for TNBC treatment.
Recently, PSMB5 expression and proteasome activity have been reported to be induced by STAT3-related oncogenic signaling, which also indicates an oncogenic role of PSMB5 (15). Inhibiting PSMB5 and subsequent proteasome function is believed to induce cellular stress and apoptosis by causing intracellular accumulation of poly-ubiquitinated proteins and has recently emerged as a promising strategy for anticancer therapy, and numerous proteasome inhibitors have been developed (11, 16, 17). Bortezomib is the first US Food and Drug Administration (FDA)–approved proteasome inhibitor for the treatment of relapsed and refractory multiple myeloma (18). Furthermore, it has also shown an effect in suppressing a variety of cancer cells, including those of pancreatic cancer, colorectal cancer, lung cancer, prostate cancer, breast cancer and glioblastomas (18, 19). Despite promising clinical efficacy, some patients develop bortezomib resistance during therapy (20). PSMB5 overexpression and mutations (e.g., G322A, C323T, C326T) have frequently been associated with bortezomib resistance (21-23), while direct and indirect inhibiting of PSMB5 expression has been reported to restore bortezomib sensitivity by exacerbating proteasome dysfunction and cellular apoptosis (15, 23).
Our results provided the first evidence that PSMB5 knockdown increased TNBC cell sensitivity to bortezomib. Notably, in addition to facilitating proteasome inhibition, we also found that PSMB5 silencing enhanced the efficacy of chemotherapeutic agents targeting other cancer cell functions. All of this evidence suggests the potency of PSMB5 depletion as an adjuvant therapy for TNBC, benefiting more patients in clinical practice.
Recently, another proteasome subunit, that of beta 1 (PSMB1), has also been reported to be inhibited by bortezomib (17). PSMB1 is a well-known proteasome structure subunit and also carries catalytic functions. In our study, we also measured PSMB1 expression as a control and identified no significant change of PSMB1 associated with knockdown of PSMB5 (Fig. 3A and E). Constant expression of PSMB1 confirmed that the PSMB5 silencing model is successful and specific, with the changes of the model specifically associated with PSMB5. However, the relationship between PSMB5 and PSMB1 and their interaction with bortezomib deserve to be investigated in depth in a future study.
In summary, our study revealed the abnormal expression of PSMB5 in TNBC and demonstrated the promise of PSMB5 as a therapeutic target for TNBC alone or in combination with other treatments.
Footnotes
Disclosures
Financial support: No grants or funding have been received for this study.
Conflict of interest: The authors declare that they have no conflicts of interest.
