Abstract
Background:
Many studies have evaluated the accuracy of EGFR mutation status in blood against that in tumor tissues as the reference. We conducted this systematic review and meta-analysis to assess whether blood can be used as a substitute for tumor tissue in detecting EGFR mutations.
Methods:
Investigations that provided data on EGFR mutation status in blood were searched in the databases of Medline, Embase, Ovid Technologies and Web of Science. The detect efficiency of EGFR mutations in paired blood and tissues was compared using a random-effects model of meta-analysis. Pooled sensitivity and specificity and diagnostic accuracy were calculated by receiver operating characteristic curve.
Results:
A total of 19 studies with 2,922 individuals were involved in this meta-analysis. The pooled results showed the positive detection rate of EGFR mutations in lung cancer tissues was remarkably higher than that of paired blood samples (odds ratio [OR] = 1.47, p<0.001). The pooled sensitivity and specificity of blood were 0.65 and 0.91, respectively, and the area under the receiver operating characteristic curve was 0.89.
Conclusions:
Although blood had a better specificity for detecting EGFR mutations, the absence of blood positivity should not necessarily be construed as confirmed negativity. Patients with negative results for blood should decidedly undergo further biopsies to ascertain EGFR mutations.
Introduction
The detection of epidermal growth factor receptor (EGFR) mutations is critical in the treatment of non-small cell lung cancer (NSCLC) patients receiving EGFR tyrosine kinase inhibitors (TKIs). For example, gefitinib and erlotinib are specially targeted at the binding site of EGFR, and have been widely recommended for treating advanced NSCLC, but research shows that only patients with positive results for EGFR mutations respond to this strategy of treatment (1). Up to now, EGFR mutations confirmed in lung cancer tissues have been considered as the gold standard in the prediction of TKI treatment response and prognosis (2). Unfortunately, lung tissue specimens are sometimes of limited availability for testing of EGFR mutations because of various subjective and objective factors (3, 4). Research shows that the detection of EGFR mutations in cell-free DNA is a feasible option in NSCLC patients (5) because the DNA of tumor cells from the death of those tumor cells and tissues can be released from primary tumors and/or local metastatic sites into the blood (6). Based on this, blood of patients may be an alternative for detecting EGFR mutations.
Although blood has been investigated as a feasible substitute for tumor tissues in testing for EGFR mutations, not all investigations have shown a stable consistency in their results for EGFR mutations between blood and cancer tissues. So far, there is still some controversy regarding whether blood can be an alternative for tumor tissues in detecting EGFR mutations to guide EGFR TKI treatment. We believe that the key to answering this question is that those samples of blood and tissues for detecting EGFR mutations should be of paired design (blood and tissue must be from the same patient). Only in this way, can we compare the advantages and disadvantages between blood and cancer tissues for detecting EGFR mutations, which is also a basic requirement of statistics for scientific research. However, so far the meta-analyses published regarding to this question have not done this very well. To clarify this problem, we carried out this systematic literature evaluation.
Materials and methods
Identification and eligibility of relevant studies
We searched the scientific literature published in the databases of Medline, Embase, Ovid Technologies and Web of Science, using free text and the Medical Subject Headings (MeSH) terms “non-small cell lung cancer,” “NSCLC,” “EGFR,” “epidermal growth factor receptor,” “mutation,” “plasma,” “serum,” “tissues,” “cell-free DNA,” “circulating tumour DNA,” “lung cancer” and “TKIs.” We also manually checked the references from the studies included. If necessary, we also contacted the authors using e-mail and telephone to verify critical data.
Criteria for inclusion and exclusion
Inclusion criteria: literature studies (i) must have compared the EGFR mutations in paired blood and tissue samples (blood and tissues from the same patients); (ii) must have reported the number of true positives (TP), true negatives (TN), false positives (FP) and false negatives (FN); and (iii) must have confirmed patients’ results histopathologically or cytologically. Exclusion criteria were (i) nonoriginal literature, including abstracts, letters, editorials and expert opinions and reviews were excluded; (ii) patients had received anticancer therapy before collecting the samples; (iii) did not contain a distinct group of patients with NSCLC; and (iv) did not pair the samples (the number of blood and tissue samples was not consistent).
Data extraction
The data we extracted were as follows: (i) title of study, author, publication date and country; (ii) study design, case number of paired blood and tissue samples and data for follow-up; and (iii) testing methods for EGFR mutations and predictive ability of EGFR mutations for prognosis in blood and tissues when patients received EGFR TKI treatment.
Methodological quality assessment
The quality of the studies was assessed by 2 evidence-based medicine experts (Gao Wenlong and Liu Xia) according to the guidelines from the Standards for Reporting of Diagnostic Accuracy (STARD) initiative, with a maximum score of 30 (7) and the Quality Assessment for Studies of Diagnostic Accuracy (QUADAS) tool, with a maximum score of 14 (8). The discrepancy of assessment was resolved by negotiating with one of the authors.
Statistical analysis
The chi-square and Fisher’s exact tests were used to detect statistically significant heterogeneity. Heterogeneity was also quantified with the I2 statistic which describes the proportion of total variation in study estimates due to heterogeneity. A fixed-effect model was first performed, assuming the same homogeneity of the effect size across all of the studies, followed by a random-effect model under the assumption of heterogeneity between studies. The effect size was generated by the estimation of odds ratios (ORs) with a 95% confidence interval (95% CI). The overall effect was calculated by Z-scores, with significance being set at a p value of <0.05. In addition, sensitivity analysis, funnel plots, Begg’s and Egger’s tests were performed to evaluate the bias of publications. Moreover, the overall sensitivity, specificity and diagnostic odds ratio (DOR) were calculated by a summary receiver operating characteristic (SROC) curve analysis (9). Relevant analyses were performed using SPSS (version 15.0; SPSS Institute, Chicago, IL, USA), Meta DiSc statistical software (version 1.4; Madrid, Spain) and Stata version 12.0 (Stata Corp., College Station, TX, USA). All p values were defined as 2-sided, and a p value of <0.05 indicated a statistical significance.
Results
Process used to search the literature
Initially, a total of 2,568 studies regarding EGFR mutations and lung cancer were found. Of these, 2,200 references were identified to be unsuitable and were removed. Subsequently, 320 studies from the rest were excluded because they were not original literature such as review, abstract and meeting records. Of the 48 publications that remained, 29 studies had to be excluded for the following reasons: repeated data, lacking key data, too small a number of cases or poor quality of study design. Finally, a total of 19 publications with 2,922 individuals met the inclusion criteria and were accepted for further analysis (Fig. 1) (10-28).
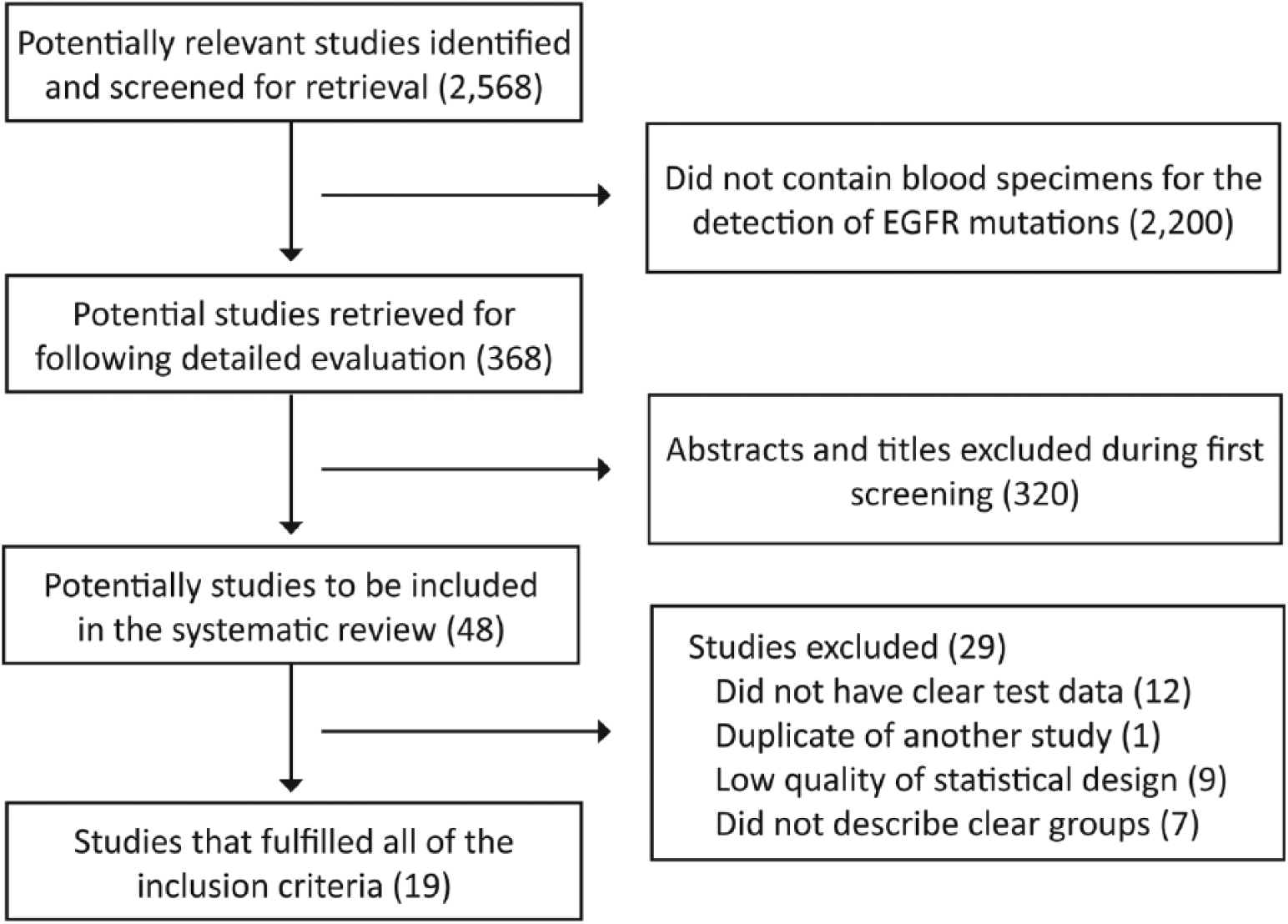
Study selection algorithm.
Description of the studies
As shown in Table I, the 19 studies included a total of 2,922 individuals, most of the studies were performed in Asian populations, and partly in Whites (10, 11). The sample sizes of studies varied between 27 (12) and 822 patients (13). Eleven studies (12-23) mainly included lung cancer patients with stages III–IV disease, and individuals in 6 studies (10, 11, 17, 24-26) had disease ranging from stage I to IV. Of the studies, 7 studies were retrospective (10, 12, 14, 15, 19, 24, 25), 3 studies were prospective (13, 16, 18) and the other 9 did not describe whether they are retrospective or prospective (11, 17, 20-23, 26-28).
Description of the studies included
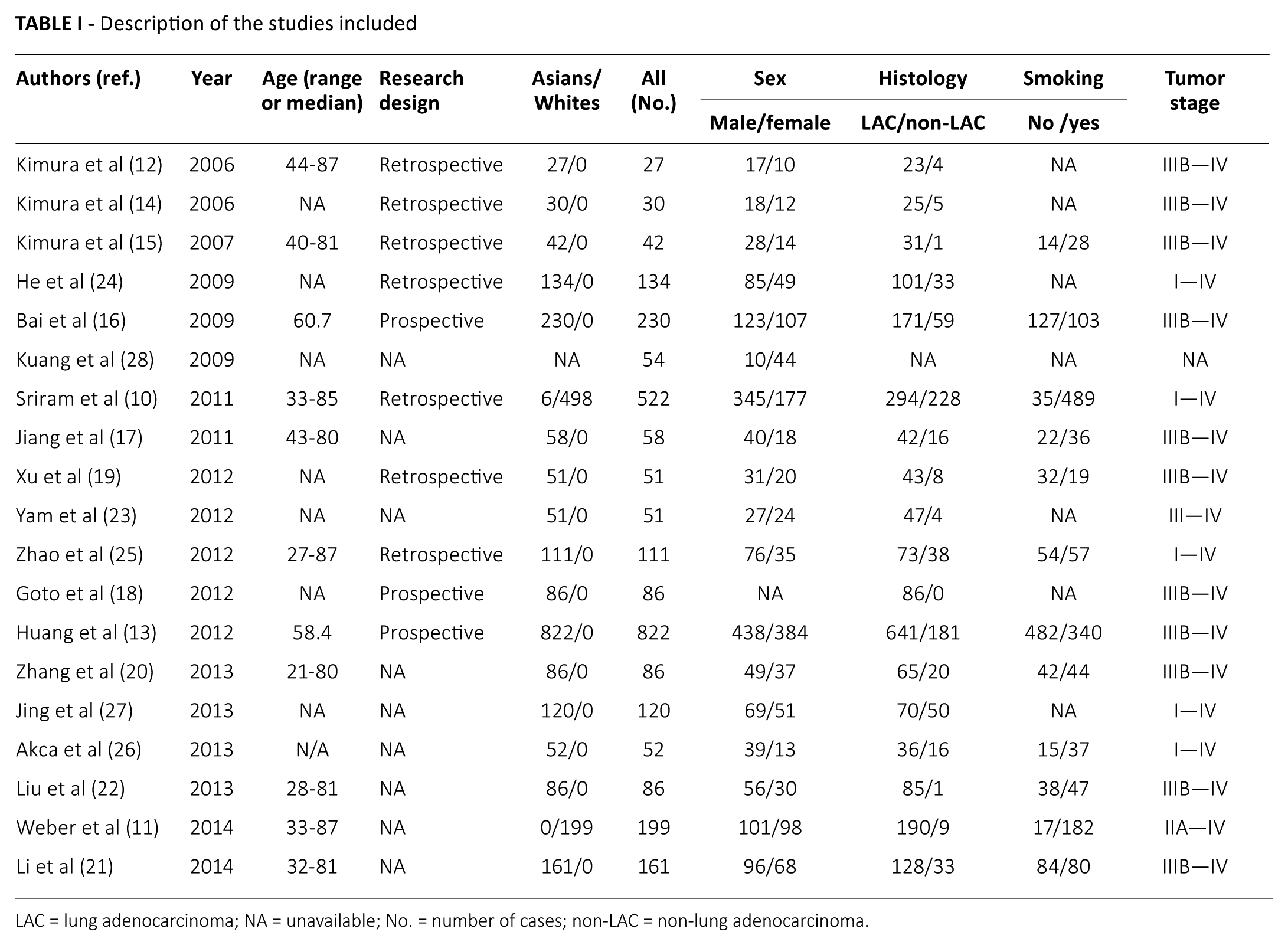
LAC = lung adenocarcinoma; NA = unavailable; No. = number of cases; non-LAC = non-lung adenocarcinoma.
Methodology and quality assessment
As shown in Table II, most of the studies included adopted direct sequencing and the strategy of Scorpion Amplification Refractory Mutation System (SARMS) to detect the status of EGFR mutations. On the whole, the studies included had a moderate (11 studies) to high (5 studies) quality. As for patient recruitment, 9 studies (10, 11, 13, 15-17, 19, 25, 28) used consecutive samples and applied the principle of randomization, and 3 studies prevented clinical design imbalances (23, 25, 28). According to the criteria of STARD and QUADAS (7, 8) the total methodological quality score ranged from 64% to 93% in the studies included, and 7 studies had QUADAS scores ≥10.
Methodology and quality of studies included
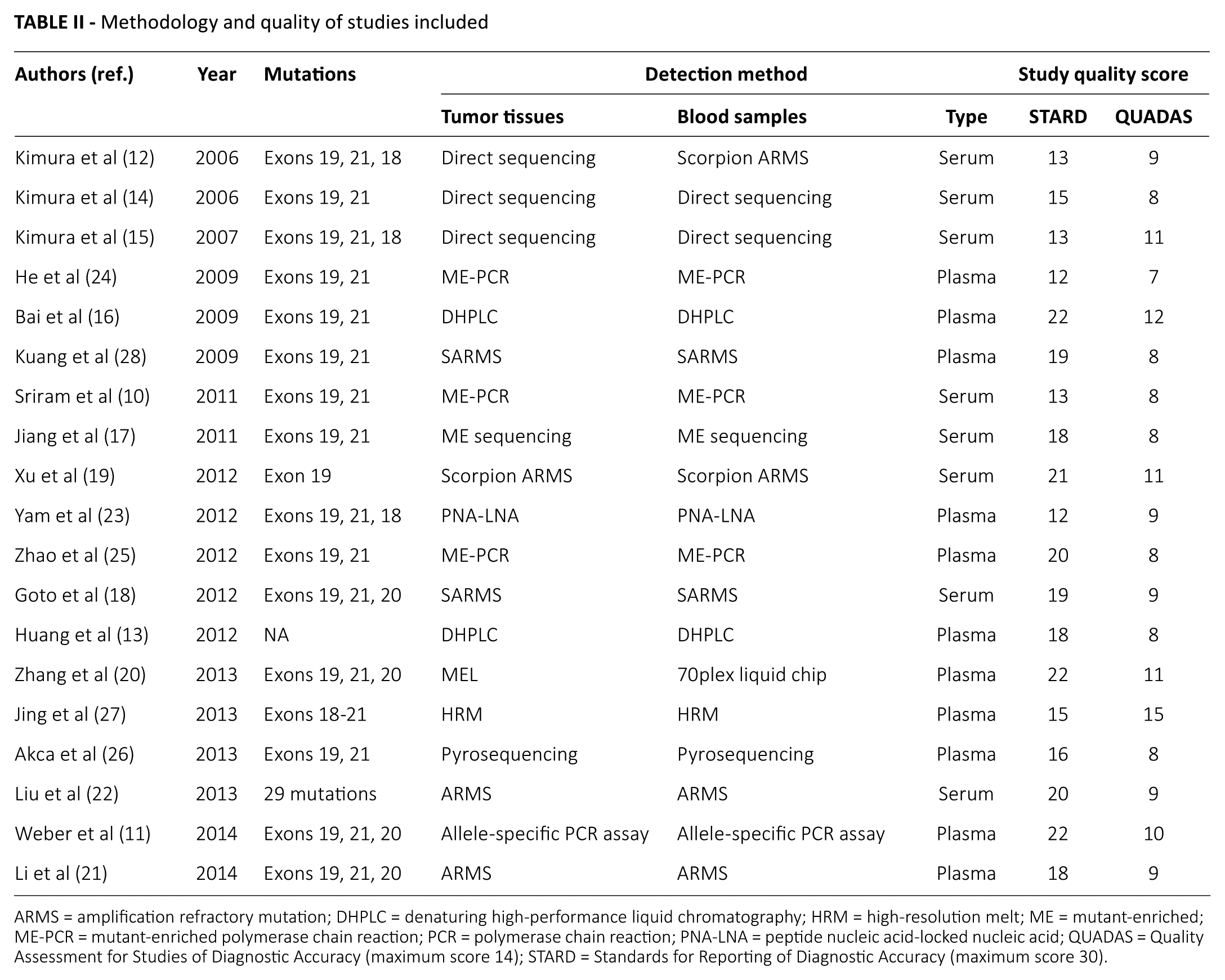
ARMS = amplification refractory mutation; DHPLC = denaturing high-performance liquid chromatography; HRM = high-resolution melt; ME = mutant-enriched; ME-PCR = mutant-enriched polymerase chain reaction; PCR = polymerase chain reaction; PNA-LNA = peptide nucleic acid-locked nucleic acid; QUADAS = Quality Assessment for Studies of Diagnostic Accuracy (maximum score 14); STARD = Standards for Reporting of Diagnostic Accuracy (maximum score 30).
Test of heterogeneity
At first, we used a fixed-effects model to perform the meta-analysis, and the chi-square value for heterogeneity was 32.6 with 18 degrees of freedom and a p value of 0.020, which indicated the possibility of heterogeneity. We inferred that the heterogeneity was mainly derived from the different test methods and sizes of samples. But we noticed that these studies had a quite good clinical homogeneity. In addition, the I2 value for the heterogeneity test was 44.2%, which suggested that the heterogeneity was basically acceptable. However, we still used the random-effects model to do the following analysis to keep the conclusions reliable (29).
Comparison of detection rates of EGFR mutations in paired blood and lung cancer tissues
As shown in Table III and Figure 2, all studies provided data on the detection rate of EGFR mutations in paired blood and tumor tissues, which showed that the weight of the included studies in the meta-analysis ranged from 1.26% to 13.03%. The OR value of comparison on detection rate of EGFR mutations in paired blood and lung cancer tissues was 1.47 (95% CI, 1.20-1.80) and the test for Z value of overall effect was 5.20 (p<0.001) (Fig. 2), indicating that the detection rate of EGFR mutations in lung cancer tissue was remarkably higher than that of the paired blood samples.
Data for EGFR mutations in paired tumor tissue and blood of patients
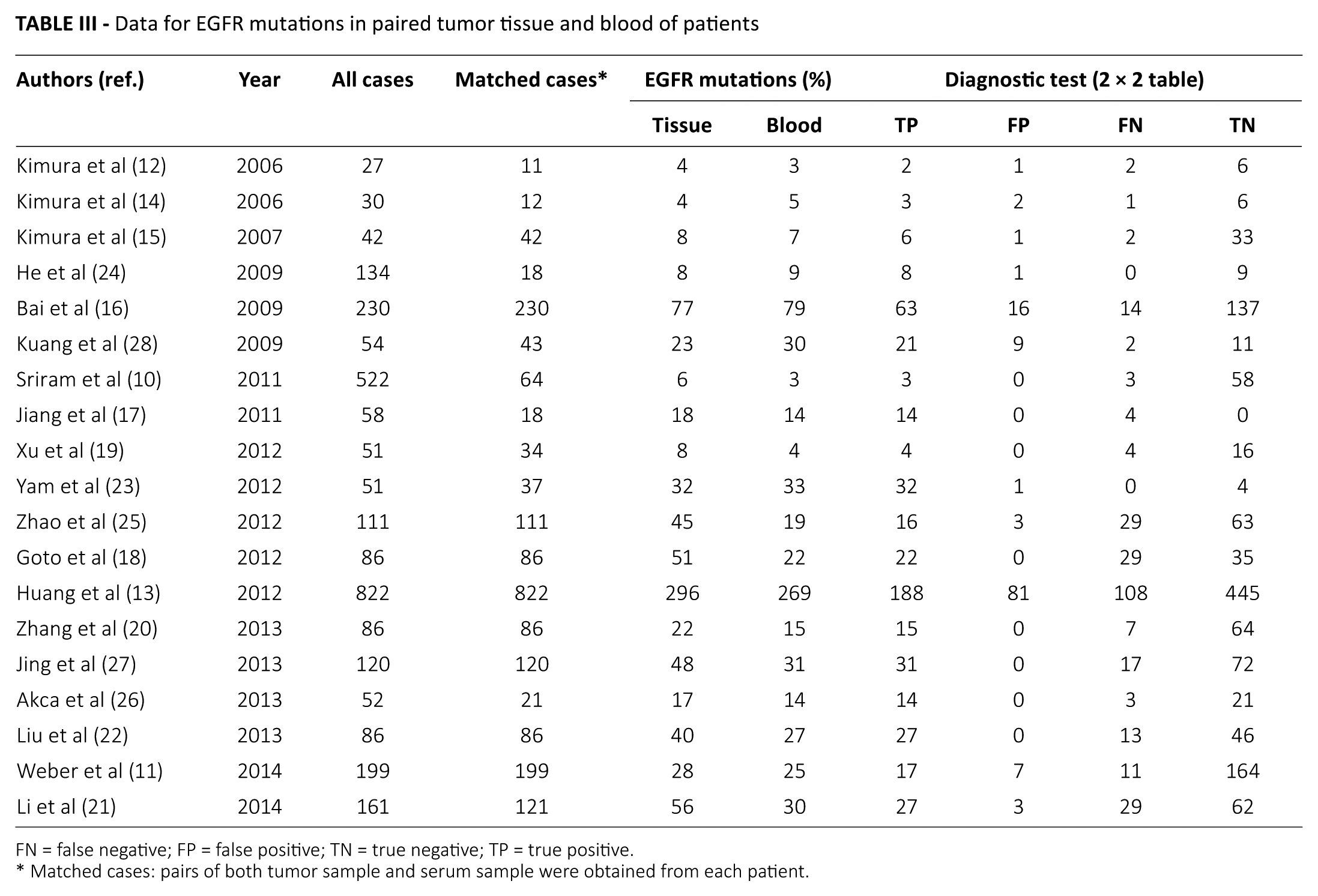
FN = false negative; FP = false positive; TN = true negative; TP = true positive.
Matched cases: pairs of both tumor sample and serum sample were obtained from each patient.
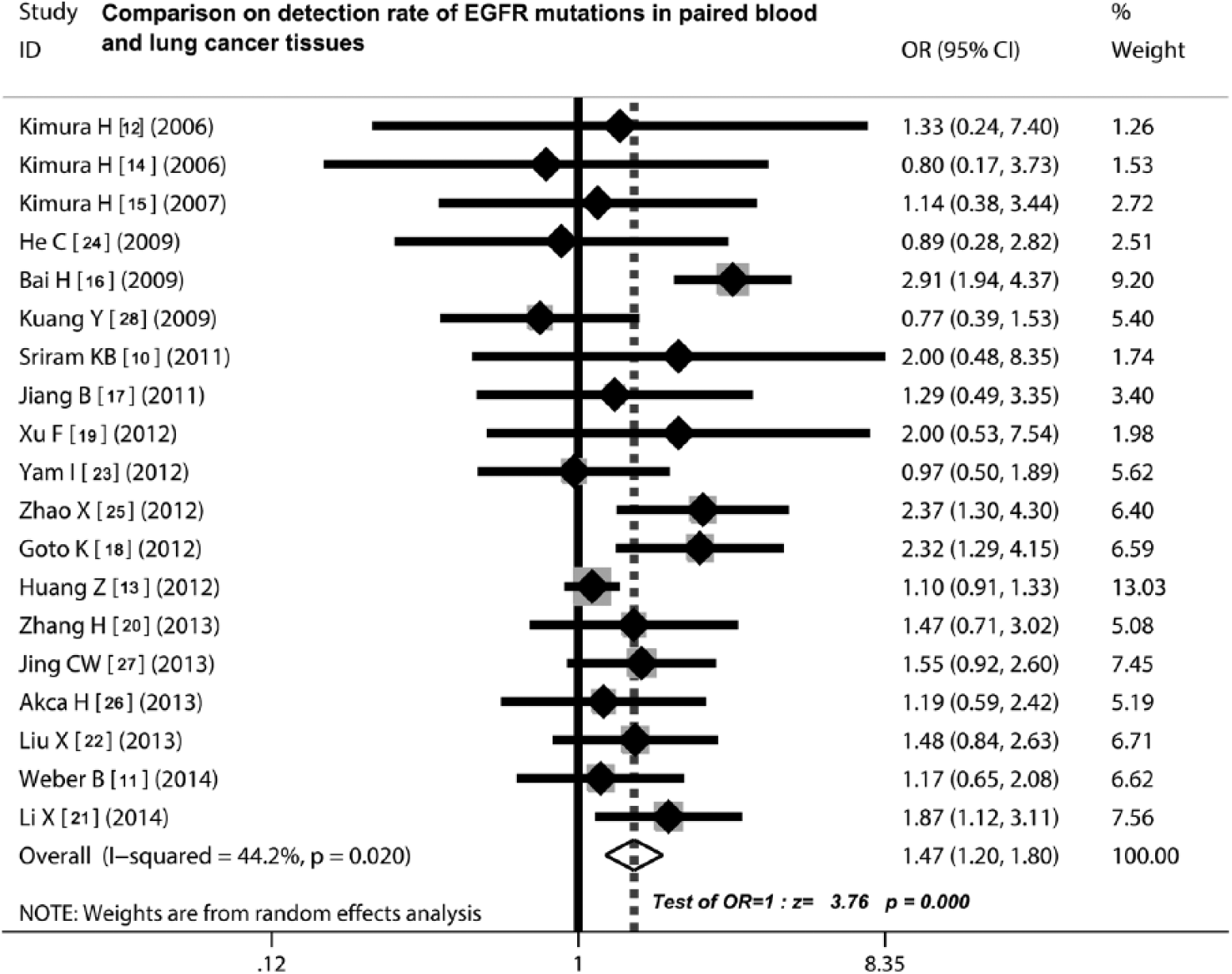
Meta-analysis using a random-effects model for EGFR mutations in blood and tissue samples. The detection rate of EGFR mutations in lung cancer tissue was significantly higher than in the paired blood samples. Test for overall effect: Z = 5.20, p<0.00001. CI = confidence interval; OR = odds ratio.
Predictive ability of EGFR mutations in different samples for treating NSCLC patients with EGFR TKIs
Five studies reported that EGFR mutations were associated with increased progression-free survival (PFS) (13, 15, 16, 19, 21), whether in paired blood or tumor tissues when the patients received TKI therapy. We found that the mean of the p value for prognosis in tumor tissues was 0.018, which was remarkably less than in blood samples (p = 0.254) (Tab. IV), suggesting that tissue EGFR mutations had a better predictive ability for PFS than blood EGFR mutations. Three studies concerned the overall survival (OS) (15, 16, 19) and showed that the patients with EGFR mutations had a longer OS than those without EGFR mutations, but the difference of predictive ability between detection in blood and tumor tissues did not show statistical significance (mean of p value in tissue: 0.163; in blood: 0.656). However, the mean of the p value in tumor tissues was still lower than that in blood, indicating that the predictive ability of EGFR mutations in tissues had a clear advantage (Tab. IV).
Predictive ability of EGFR mutations in paired blood and tumor tissue treated with EGFR-TKI
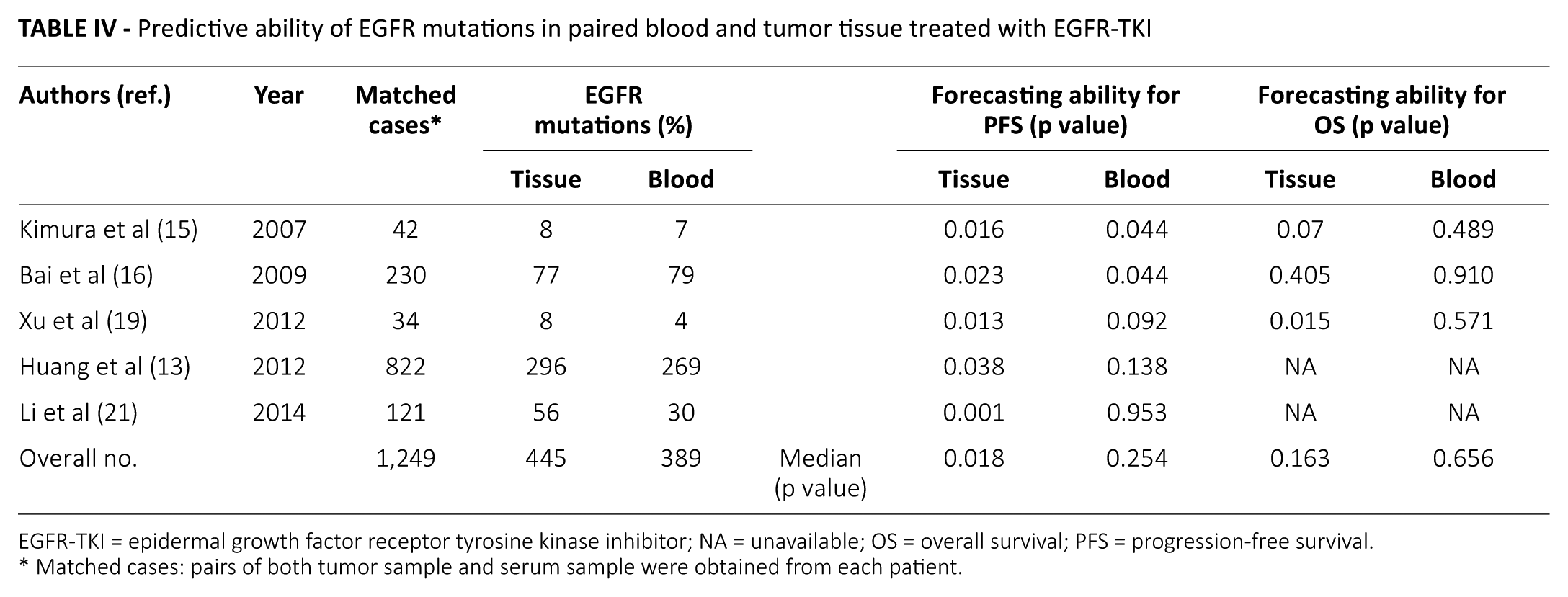
EGFR-TKI = epidermal growth factor receptor tyrosine kinase inhibitor; NA = unavailable; OS = overall survival; PFS = progression-free survival.
Matched cases: pairs of both tumor sample and serum sample were obtained from each patient.
Analysis of sensitivity and publication bias
The sensitivity analysis showed that the deletion of any 1 study did not substantially modify the estimators and affect the statistical value, and the pooled OR varied between 0.52 and 45.85 (Fig. 3A). The shape of the funnel plot derived from the studies included appeared to be approximately symmetrical (Fig. 3B). Moreover, the Begg’s test (p = 0.463) and the Egger’s test (p = 0.67) both did not indicate that a publication bias existed (Fig. 3C).
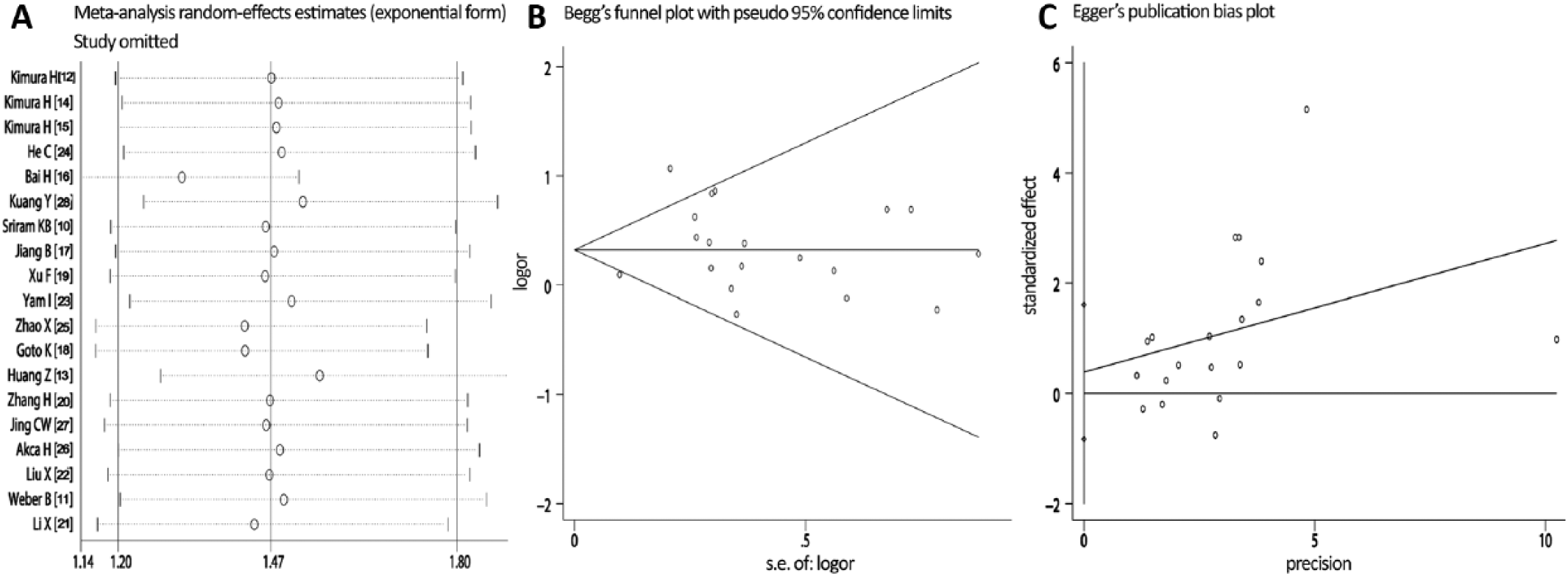
Analysis of sensitivity and publication bias. (
Sensitivity and specificity of EGFR mutation detection in blood
Figure 4 shows the forest plot of the sensitivity and specificity of 19 studies for EGFR mutations in paired blood and tumor tissues. The pooled sensitivity of EGFR mutations in blood was 0.65 (95% CI, 0.65-0.68) (Fig. 4A) and specificity was 0.91 (95% CI, 0.89-0.92) (Fig. 4B), when the detection results in lung cancer tissues was served as a gold standard. The summary positive likelihood ratio (PLR) was 8.225 (95% CI, 5.127-13.196) and summary negative likelihood ratio (NLR) was 0.398 (95% CI, 0.325-0.487).
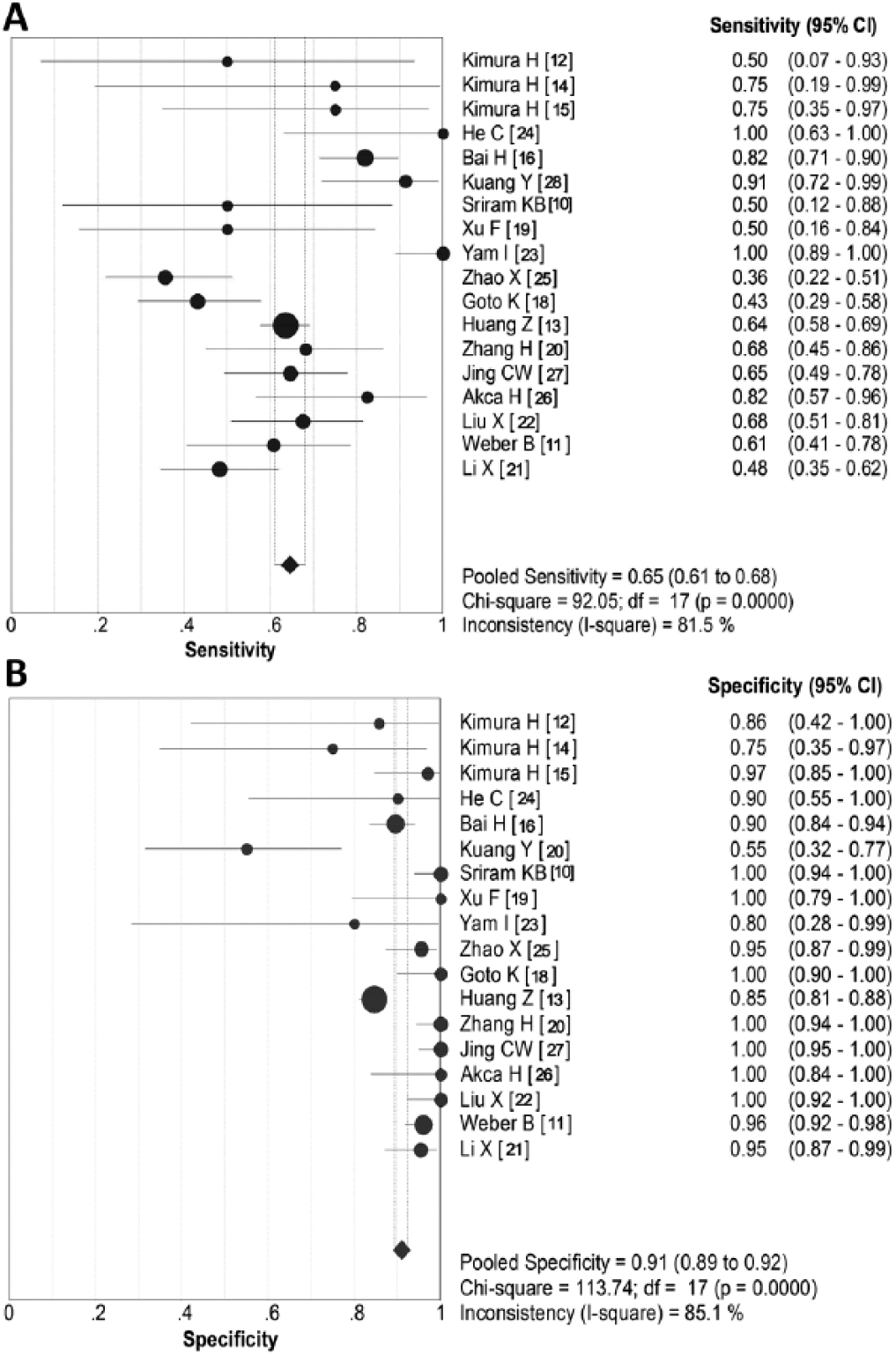
Forest plots of sensitivity and specificity for EGFR mutation detection in blood samples. (
Diagnostic accuracy for EGFR mutation detection in blood
The diagnostic odds ratio (DOR) of 19 studies on EGFR mutation detection in blood was 32.33 (95% CI, 17.6-59.4), and the chi-square value was 38.51, with all p values less than 0.05 (Fig. 5A). From the SROC curve analysis, we found most studies had a better specificity estimation close to 1.00 for blood EGFR mutations, only 1 study (28) showed a low specificity of EGFR mutations in peripheral blood (<0.60). In contrast, estimations of sensitivity were relatively lower in blood than in lung cancer tissues because the sensitivity of 6 studies being less than 0.60 (10, 12, 18, 19, 21, 25). Finally, the maximum join of sensitivity and specificity was 0.83 (standard error of the mean = 0.027) while the area under curve (AUC) was 0.894 (standard error of the mean = 0.026) (Fig. 5B).
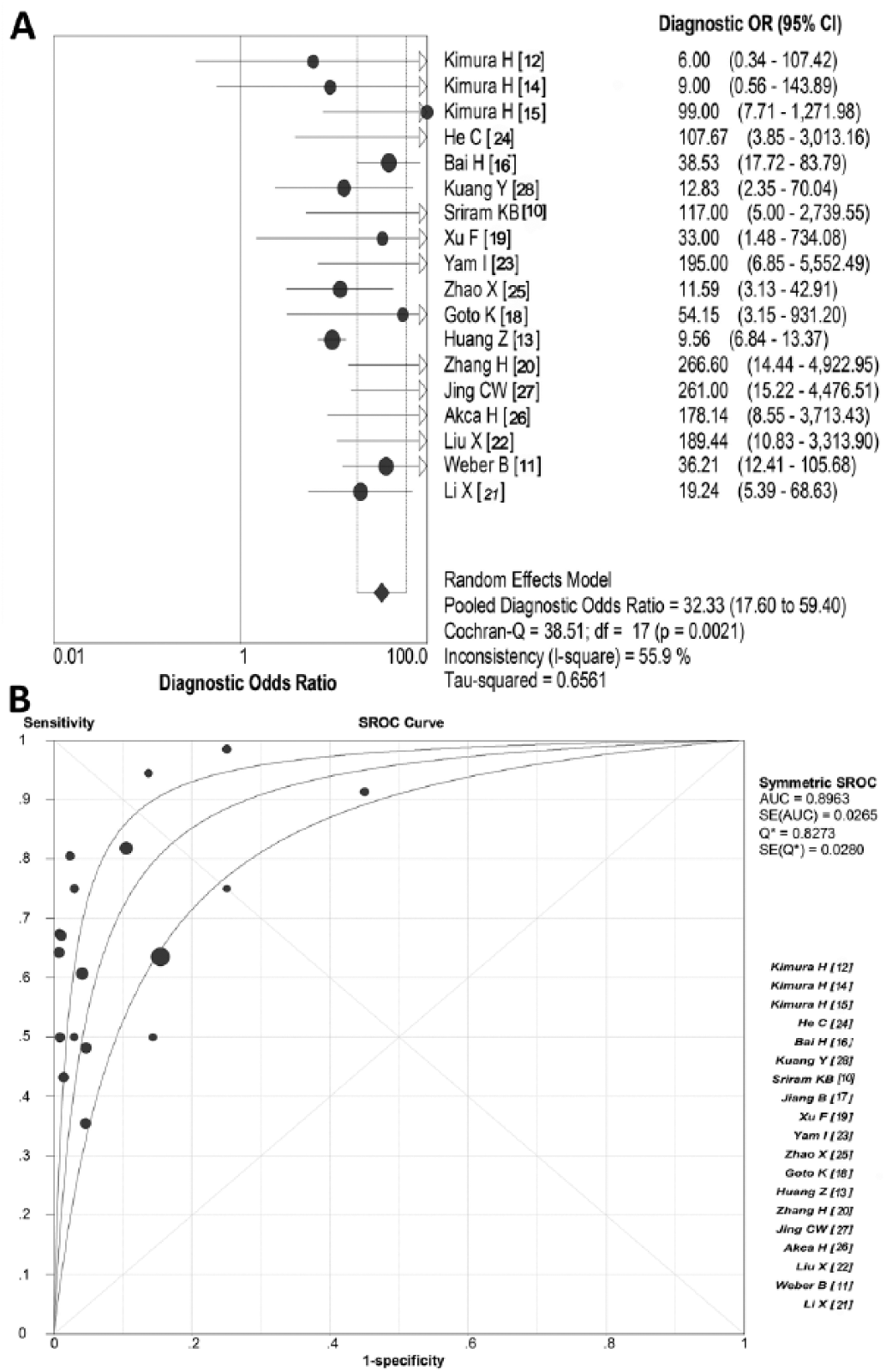
Diagnostic odds ratio (DOR) and summary receiver operating characteristic (SROC) curve for EGFR mutation detection in blood samples. (
Discussion
The outcomes of TKI treatment for NSCLC are always dependent on the status of EGFR mutations (30) because studies has been showing that NSCLC patients with mutations of EGFR have a better response to EGFR TKIs than those without mutations (31, 32). In the clinic, EGFR mutations are always identified from operative resected tumor samples, as we know. However, it is sometimes difficult to obtain tumor samples from patients with inoperable NSCLC. Moreover, repeated biopsies for tissue detection are viewed to be impractical because of the poor condition of these patients’ bodies. Thus, it is necessary to develop other more readily accessible samples to detect EGFR mutations (33). Up to now, many relevant studies into detecting EGFR mutations have focused on the blood samples, but theirs conclusion are still not certain.
In the present study, we reviewed 19 studies which detected the EGFR mutations in paired samples of lung cancer tissues and blood. Through a strict quality assessment, we found that the quality of these studies is relatively good. (7, 8). The results showed that the OR for comparisons of EGFR mutations between lung cancer tissues and blood was 1.47, suggesting that the detection rate of EGFR mutations in lung cancer tissues was remarkably higher than that of blood. This result implied that the absence of blood positivity should not necessarily be construed as a confirmation of negativity. Even so, the results still showed that blood could be a valuable alternative when lung cancer tissue is difficult to obtain or is insufficient (34). This new strategy may help us select patients who are most likely to benefit from TKIs, although the different detection methods used in the studies had an obvious impact on the reliability of any conclusions, and these partly explained the heterogeneity in our study. However, we also found a good clinical homogeneity existed in those studies, and there was not a possibility of publication bias. Altogether, the conclusion of this present meta-analysis should be more certain.
Using the positive results of EGFR mutations determined from tissues with lung cancer as a gold standard, we found that the detection of EGFR mutations in blood samples had a relatively higher specificity of 0.91 (95% CI, 0.89-0.92) and a summary PLR of 8.225 (95% CI, 5.127-13.196), which suggested that choosing blood samples to determine EGFR mutations has a potential advantage of specificity, thus indicating that blood could be a potential alternative for lung cancer tissues. However, the summary estimate of sensitivity was only 0.65 (95% CI, 0.61-0.68), which implied that some real EGFR mutations can not be found by the detection of blood samples, But the detection of EGFR mutations in blood samples is still necessary because we can confirm the EGFR mutations as long as it is positive in the blood. In addition, we noticed that the blood detection of EGFR mutations had lower NLR values, which indicated that EGFR mutations could not be excluded even if a patient’s result for EGFR mutations was negative in blood. In other words, a negative test in blood does not mean the absence of EGFR mutations, and patients with negative results in blood still have a chance of having positive results in lung cancer tissues, which is an important issue that we still have to face.
In the present study, we found that tissue that was EGFR mutation positive had a higher predictive ability in terms of survival than the results of blood detection, which means that the advantages of EGFR mutations in tissues for prognosis of lung cancer treated with TKIs far outweigh those for blood samples. The mean DOR of 38.53 further indicated that blood samples had a better specificity for the identification of EGFR mutations in spite of its low detection rate. In clinical practice, the identification of EGFR mutations usually requires biopsies of adequate size, a 38.53 DOR for blood samples would help confirm the positive result. As soon as the EGFR mutations are confirmed to be positive in blood samples of patients, the conclusion is certain, which may avoid further invasive tests on patients, which has great significance. Nevertheless, although blood samples have the advantage of specificity, this choice still has certain restrictions due to its higher false-negative rate. The SROC curve represents the performance of a diagnostic test (35): an AUC of 0.93 to 0.96 is very good, and 0.75 to 0.92 is good (29, 35). In our analysis, the AUC of blood in detecting EGFR mutations was 0.89, which showed that the choice of blood samples has a relatively good accuracy.
Based on the facts mentioned above, our present study proposes that patients who intend to accept TKI therapy should firstly undergo evaluation of blood samples for EGFR mutations, which can provide a evidence that is certain and avoid further invasive biopsy. However, patients with negative results for EGFR mutations in blood should decidedly undergo further invasive tests to ascertain EGFR mutations from tumor tissues.
Finally, there were limitations to this systematic evaluation: first, some studies had small size; second, the statistical designs of some of the studies had flaws; and third, and most critically, the detection methods for EGFR mutations in the studies were not the same.
Therefore, in future we should further investigate the differences in EGFR mutations in paired blood and lung cancer tissues in a large population involving multiple clinical centers. We should also continue to focus on the optimization of the methodology for the detection of EGFR mutations.
Conclusion
The detection rate of EGFR mutations in tissues with lung cancer was remarkably higher than that in blood, which means blood detection may miss some positive results for EGFR mutations. However, the higher specificity of blood samples suggests that their investigation could still be a valuable alternative to the use of tumor samples when tumor tissue is absent or insufficient.
Footnotes
Acknowledgements
The authors wish to thank Drs. Wu Dianlei and Wang Xinhong for their helpful discussions and critical comments provided during the preparation of this manuscript. They also wish to thank Gao Wenlong and Liu Xia for the quality assessment of the included research.
Disclosures
Financial support: No grants or funding have been received for this study.
Conflict of interest: None.
Abbreviations
AUC Area under the curve
CI Confidence interval
DOR Diagnostic odds ratio
EGFR Epidermal growth factor receptor
FN False-negative
FP False-positive
NSCLC Non-small cell lung cancer
NLR Negative likelihood ratio
OR Odds ratios
OS Overall survival
PFS Progression-free survival
PLR Positive likelihood ratio
QUADAS Quality Assessment for Studies of Diagnostic Accuracy
SARMS Scorpion Amplification Refractory Mutation System
SROC Summary receiver operating characteristic curve
STARD Standards for Reporting Diagnostic Accuracy
TN True-negative
TP True-positive
TKI Tyrosine kinase inhibitor
