Abstract
Introduction
Non-small cell lung cancer (NSCLC), which accounts for >85% of all lung cancers, is the most common solid malignant tumor, with high morbidity and mortality worldwide. Epidermal growth factor receptor tyrosine kinase inhibitor (EGFR-TKI) treatment is widely used clinically because the epidermal growth factor receptor (EGFR) is the main driver gene of lung cancer; however, the development of drug resistance is inevitable. Epigenetic abnormalities can also lead to tyrosine kinase inhibitor (TKI) resistance. However, the relationship between N6-methyladenosine (m6A)-related proteins and EGFR mutations in NSCLC and their clinical significance remain unclear. In this retrospective study, the expression of m6A-related regulatory factors in patients with EGFR mutations were analyzed to investigate their relationship with clinicopathological features and prognoses.
Methods
The association between m6A-related regulatory factors and NSCLC was analyzed using data derived from The Cancer Genome Atlas, case collection, follow-up, immunohistochemistry, and scoring.
Results
A total of 246 NSCLC specimens were examined in this study. Among these, 143 EGFR-mutant cases exhibited significantly higher expression of METTL3 and ALKBH5 compared to EGFR-wildtype specimens. The median progression-free survival time of patients with high METTL3 expression (SI >6) was 25.0 months, and that of patients with high ALKBH5 expression (SI >6) was 24.1 months in EGFR-mutant cases. High METTL3 and ALKBH5 expression levels are independent risk factors for progression-free survival in patients with EGFR mutations. The median progression-free survival time of patients with EGFR mutations was 45.7 months in those with high expression of METTL3 or ALKBH5 alone, whereas it decreased to 20.1 months in those with high simultaneous expression of METTL3 and ALKBH5.
Conclusions
METTL3 and ALKBH5 were upregulated in NSCLC tissues with EGFR mutations and significantly correlated with poor prognoses. Thus, METTL3 and ALKBH5 may serve as prognostic biomarkers in EGFR-mutant NSCLC.
Plain Language Summary
Non-small cell lung cancer (NSCLC), which accounts for >85% of all lung cancers, is the most common solid malignant tumor with high morbidity and mortality worldwide. Epidermal growth factor receptor tyrosine kinase inhibitor (EGFR-TKI) treatment is widely used clinically because the epidermal growth factor receptor (EGFR) is the main driver gene of lung cancer; however, drug resistance is inevitable. Epigenetic abnormalities can also lead to tyrosine kinase inhibitor (TKI) resistance. However, the relationship between N6-methyladenosine (m6A)-related proteins and EGFR mutations in NSCLC and their clinical significance remain unclear. In this retrospective study, the expression of m6A-related regulatory factors in patients with EGFR mutations were analyzed to explain their relationship with clinicopathological features and prognoses. A total of 246 NSCLC specimens were examined in this study. Among these, 143 EGFR-mutant cases exhibited significantly higher expression of METTL3 and ALKBH5 compared to EGFR-wildtype specimens. The median progression-free survival time of patients with high METTL3 expression (SI >6) was 25.0 months, and that of patients with high ALKBH5 expression (SI >6) was 24.1 months in EGFR-mutant cases. High METTL3 and ALKBH5 expression levels are independent risk factors for progression-free survival in patients with EGFR mutations. The median progression-free survival time of patients with EGFR mutations was 45.7 months in those with high expression of METTL3 or ALKBH5 alone, whereas it decreased to 20.1 months in those with high simultaneous expression of METTL3 and ALKBH5. METTL3 and ALKBH5 were upregulated in NSCLC tissues with EGFR mutations and significantly correlated with poor prognoses. Thus, METTL3 and ALKBH5 may serve as prognostic biomarkers in EGFR-mutant NSCLC.
Introduction
Lung cancer, a malignant tumor originating from the bronchial mucosa or glands, is the second most common type of malignant tumor worldwide, with new cases accounting for 11.4% of all cancer cases, following breast cancer (11.7%). 1 According to Cancer Statistics 2023, lung cancer has the highest death rate among all cancers, which is much higher than that of breast, prostate, and colorectal cancers. 2 In China, approximately 787 000 people die of lung cancer, accounting for the highest cancer death rate among those aged 40-80 years.3,4 According to its histopathological classification, lung cancer is mainly divided into small cell lung cancer and non-small cell lung cancer (NSCLC), of which NSCLC accounts for more than 85% of cases. 5 Platinum-based chemotherapy is the main treatment for lung cancer, which results in healthy cells being damaged while killing tumor cells, causing strong side effects owing to poor specificity. With the development of molecular diagnostics and precision medicine, targeted therapies for oncogenic mutations have gradually become common. TP53, KRAS, and epidermal growth factor receptor (EGFR) are the primary oncogenes involved in NSCLS. 6 EGFR, a transmembrane glycoprotein receptor, comprises 28 exons on the short arm of chromosome 7 (7p12-14). It belongs to the receptor tyrosine kinase erbB family and promotes tumor cell proliferation, angiogenesis, and invasion after activation. 7 EGFR mutations in NSCLC are mainly located in the tyrosine kinase exon 18-21 region, among which exon 19 deletions (19del) and 21 L858R point mutations (L858R) are the most common, accounting for 45% and 40% of the mutations, respectively. In addition, uncommon mutations, such as G719X, L861Q, S768I, and T790M, account for approximately 10-18% of cases. 8 Epidermal growth factor receptor tyrosine kinase inhibitors (EGFR-TKIs) enhance the efficacy and improve the survival of patients with NSCLC. Guidelines from the National Comprehensive Cancer Network and Chinese Society of Clinical Oncology suggest that EGFR-TKIs should be the first-line treatment for patients with EGFR mutations. First-generation EGFR-TKIs (gefitinib, icotinib, and erlotinib) prolong progression-free survival (PFS) but do not affect overall survival (OS).9,10 Second-generation TKIs (afatinib and dacomitinib) covalently and irreversibly bind to ATP and act as EGFR-TKIs. However, the drug resistance caused by the EGFR T790M mutation cannot be overcome by first- and second-generation EGFR-TKI treatments. 11 Irreversibly covalently bound third-generation TKIs (osimertinib and almonertinib) overcome drug resistance caused by the EGFR-T790M mutation and have greater efficacy but fewer side effects. However, drug resistance caused by ocitinib treatment-induced EGFR C797S mutation has been reported. 12 Recently, epigenetic abnormalities have been found to be associated with EGFR-TKI resistance. Epigenetics refers to the reversible and heritable changes in gene expression without changes in the nucleotide sequence, including DNA and RNA methylation, noncoding RNA regulation, histone modification, and chromatin remodeling. Many studies have demonstrated that tumorigenesis and development are related to the epigenetic modification of RNA. RNA methylation accounts for more than 60% of all RNA modifications, and N6-methyladenosine (m6A) is the most abundant form of RNA post-transcriptional modification in eukaryotes. 13 The m6A methylation enzymes methyltransferase-like 3 (METTL3), alkB homolog 5 (ALKBH5), and alpha-ketoglutarate-dependent dioxygenase (FTO) promote NSCLC proliferation and metastasis via different pathways.14-16 Many m6A-modified genes were identified in afatinib-resistant cell lines, suggesting that m6A methylation may affect the sensitivity of NSCLC cell lines to afatinib by interfering with the cell cycle. 17 Thus, m6A modification is predicted to be involved in TKI resistance in NSCLC cells.
In this study, we aimed to explain the expression of m6A-related proteins in tumor tissues of patients with EGFR-mutant NSCLC using an online database and clinical specimens, explore the relationship between their expression and prognosis, and identify potential therapeutic targets for NSCLC.
Materials and Methods
Data Acquisition and Analysis
NSCLC data, including RNA sequencing profiles, clinical information, and gene variation profiles, were processed and downloaded using cBioPortal (https://www.cbioportal.org) from The Cancer Genome Atlas (TCGA; https://tcga-data.nci.nih.gov/tcga/). All data were analyzed using GraphPad Prism 10.0 (GraphPad Software Inc., USA) and Excel 2019 (Microsoft Inc., USA). The expression of m6A-related genes (the unit was RNA-seq by expectation maximization [RSEM]) and log2 values were obtained. m6A is the most prevalent, abundant, and highly conserved internal transcript modification, particularly in eukaryotic mRNAs. The m6A modification related regulatory genes include six “writers” (METTL3, METTL14, RBM15, RBM15B, WTAP, and ZC3H13), methyltransferases including two “erasers” (FTO and ALKBH5), demethylases including five “readers” (YTHDC1, YTHDC2, YTHDF1, YTHDF2, and YTHDF3), and m6A-binding proteins.18,19 The “writers,” “erasers” and “readers” play the functions of installation, removal, and recognition.
Patient and Clinical Sample Processing
PFS was set as the primary outcome; the estimated regression coefficient was 0.25, the incidence of PFS endpoint events at 60 months of follow-up was about 0.5, the standard deviation was 2, the test level α was set as bilateral 0.05, the test power 1-β was 0.8, and considering a 10% sample loss rate, the minimum sample size was set as 140 cases using the cox regression module in the sample size calculation software PASS 15. A total of 246 cases of NSCLC from 2016 to 2019 were randomly selected from Jiangmen Central Hospital for this retrospective study. Paraffin-embedded pathological specimens were sectioned at 4 μm thick and stained with hematoxylin and eosin (H&E) using standard procedures. Under the guidance of a pathologist, areas rich in tumor cells were scraped under a microscope (FTL-7063A; Cossim, Beijing, China) for DNA extraction (QIAamp DNA FFPE; Qiagen, Hilden, Germany). The specimens were analyzed by Hybribio (Guangzhou, China, https://www.hybribio.cn/) using Sanger sequencing to obtain EGFR mutation information for comparison with Cosmic (https://www.sanger.ac.uk/genetics/CGP/cosmic/). Detailed information is presented in the supplemental tables. A total of 103 patients (Supplemental Table 1) with wild-type EGFR (EGFR-wt) and 143 patients (Supplemental Table 2) with EGFR mutations (EGFR-mut) were collected. The
IHC and Analysis
IHC was performed to examine m6A-related protein expression in the NSCLC specimens. Briefly, the slides of paraffin-embedded sections at the 4 μm tick were dewaxed with xylene and hydrated with graded ethanol after heating at 65°C for 1 h. These were then antigen retrieved in Tris/EDTA (TE; pH 9.0) buffer for 10 min of microwave heating, blocked using hydrogen peroxide and goat serum, and incubated overnight at 4°C in a humidified box with the primary antibodies. The primary antibodies were diluted in antibody diluent (Abcam, Cambridge, MA, USA) with different concentrations of 1:400 ALKBH5 Rab mAb (ab195377, Abcam), 1:200 P53 Mou mAb (DO-7, ZSGB-BIO), 1:200 METTL3 Rab mAb (ab195352, Abcam), 1:100 RBM15 Mou mAb (66059-1-Ig, Proteintech), 1:100 METTL14 Mou mAb (MA5-24706, Thermo-Fisher), 1:200 WTAP Mou mAb (60188-1-Ig, Proteintech), 1:100 FTO Mou mAb (MA5-27142, Thermo-Fisher), 1:200 YTHDC1 Rab mAb (ab220159, Abcam), 1:400 YTHDC2 Rab poly (PA5-57920, Thermo-Fisher), 1:400 YTHDF1 Rab poly (17479-1-AP, Proteintech), 1:400 YTHDF2 Rab poly (24744-1-AP, Proteintech), and 1:200 YTHDF3 Rab poly (PA5-26983, Thermo-Fisher). After incubation, the slides were washed in tris-buffered saline or 0.05% Tween 20, incubated with biotin-conjugated secondary antibodies and biotin-conjugated streptavidin (Proteintech, Wuhan, China) for 30 min at 37°C, respectively, and stained by the 3,3′-diaminobenzidine (DAB) Enhanced Liquid Substrate System (Sigma-Aldrich, St. Louis, MO, USA). Images were collected at 40× objective magnification (numerical aperture, 0.75) using an M8 digital microscope (PreciPoint, Germany). The staining index (SI) provided by experienced investigators was averaged for further comparative evaluation of the m6A methylation regulatory factor expression. The proportion of tumor cells was scored as follows: 0 (no positive tumor cells), 1 (<10% positive tumor cells), 2 (10-35% positive tumor cells), 3 (36-70% positive tumor cells), and 4 (>70% positive tumor cells). The staining intensity was graded as follows: 0 (no staining), 1 (weak staining, light yellow), 2 (moderate staining, yellow-brown), 3 (strong staining, brown) and 4 (super strong staining, brown-black). The SI was calculated as the product of the staining intensity score and proportion of positive tumor cells. The expression of all m6A methylation regulatory factors in NSCLC was evaluated using the SI, with scores of 0, 1, 2, 3, 4, 6, 8, 9, 12 and 16. High and low expressions were sectionalized as follows: The SI ≤6 was divided into low-expression group, whereas the SI >6 was considered high-expression group.
Statistical Analysis
The mRNA expression of m6A methylation regulatory factor from TCGA is presented as mean ± standard deviation (SD) and analyzed using unpaired
Results
Expression of m6A-Related Regulatory Factors in EGFR, TP53, and KRAS Mutant NSCLC
First, a mutant spectrum of 13 m6A-related regulatory factors in pan-lung cancer was found infrequently with a 0.8-5% mutation rate, of which YTHDF3 was the first at 5% (Supplemental Figure 1). Therefore, we investigated the relationship between m6A regulatory factors and common mutations in TP53, EGFR, and KRAS in NSCLC. Driving mutations predominantly accounted for 67.2% of TP53 mutations, with 11.2% in EGFR-mutant and 19.7% in KRAS-mutant NSCLC (Supplemental Figure 2A-C). RBM15 and ALKBH5 were upregulated in TP53-mutant NSCLC (Supplemental Figure 2D), METTL3 was upregulated in EGFR (Supplemental Figure 2E), and METTL3 and RBM15 were downregulated in KRAS-mutant NSCLC (Supplemental Figure 2F). Thus, METTL3, ALKBH5, and RBM15 were used in subsequent experiments. Analysis of the three genes related to mutations in all 1017 lung cancer samples from TCGA showed that TP53-associated mutations accounted for 51.7% of the first mutations, 10.8% of KRAS, and 3.0% of EGFR (Figure 1A). The mRNA levels of the three genes in TP53-, EGFR-, and KRAS-mutant NSCLC showed that METLL3 and ALKBH5 were significantly upregulated in EGFR- and TP53-mutant NSCLC tissues (Figures 1B and C), respectively, whereas RBM15 did not change significantly (Figure 1D). The Expression of METTL3, ALKBH5 and RBM15 in EGFR, TP53 and KRAS Mutated Lung Cancer. (A) The Mutation Ratio in Lung Cancer. (B) mRNA Expression of METTL3. (C) mRNA Expression of ALKBH5. (D) mRNA Expression of RBM15. Each Bar Represents the Means ± Standard Deviation (SD) and 
Expression of m6A Modifications Involving Key Factors in NSCLC
“Writers”
The “writers” of m6A are mainly composed of METTL3, METTL14, and their cofactor WTAP and RBM15.
13
IHC staining of these four regulators revealed different levels of expression (Figure 2A). Compared to that in the EGFR-wt group, METTL3 expression was significantly upregulated in EGFR-mutant specimens. (Figure 2B), whereas METTL14, RBM15, and WTAP showed no significant differences (Figure 2C-E). The Expression of m6A “Writers” in EGFR Wild Type and EGFR Mutated NSLCL Tissues. (A) Typical Pictures of Immunohistochemical Staining of m6A “Writers” in EGFR-w.t. and EGFR-Mut. NSCLC Tissues. The Scale Bar in the Images Represents 100 μm and 20 μm Respectively. Images Were Collected at a 40× Objective Magnification (Numerical Aperture 0.75) Using an M8 Digital Microscope. (B) Immunohistochemical Staining Index of METTLE3 in EGFR-Mut. Tissues Compared With EGFR-w.t. (w.t., 
“Eraser”
Demethylation is caused mainly by FTO and ALKBH5. IHC staining showed that ALKBH5 was highly expressed in the EGFR-mut NSCLC tissues compared to the wild type (Figure 3A and B), whereas FTO showed no significant change (Figure 3A and C). The Expression of m6A “Erasers” in EGFR Wild Type and EGFR Mutated NSLCL Tissues. (A) Typical Pictures of Immunohistochemical Staining of m6A “Erasers” in EGFR-w.t. and EGFR-Mut. NSCLC Tissues. The Scale bar in the Images Represents 100 μm and 20 μm Respectively. Images Were Collected at a 40× Objective Magnification (Numerical Aperture 0.75) Using an M8 Digital Microscope. (B) Immunohistochemical Staining Index of ALKBH5 in EGFR-Mut. Tissues Compared With EGFR-w.t. (w.t., 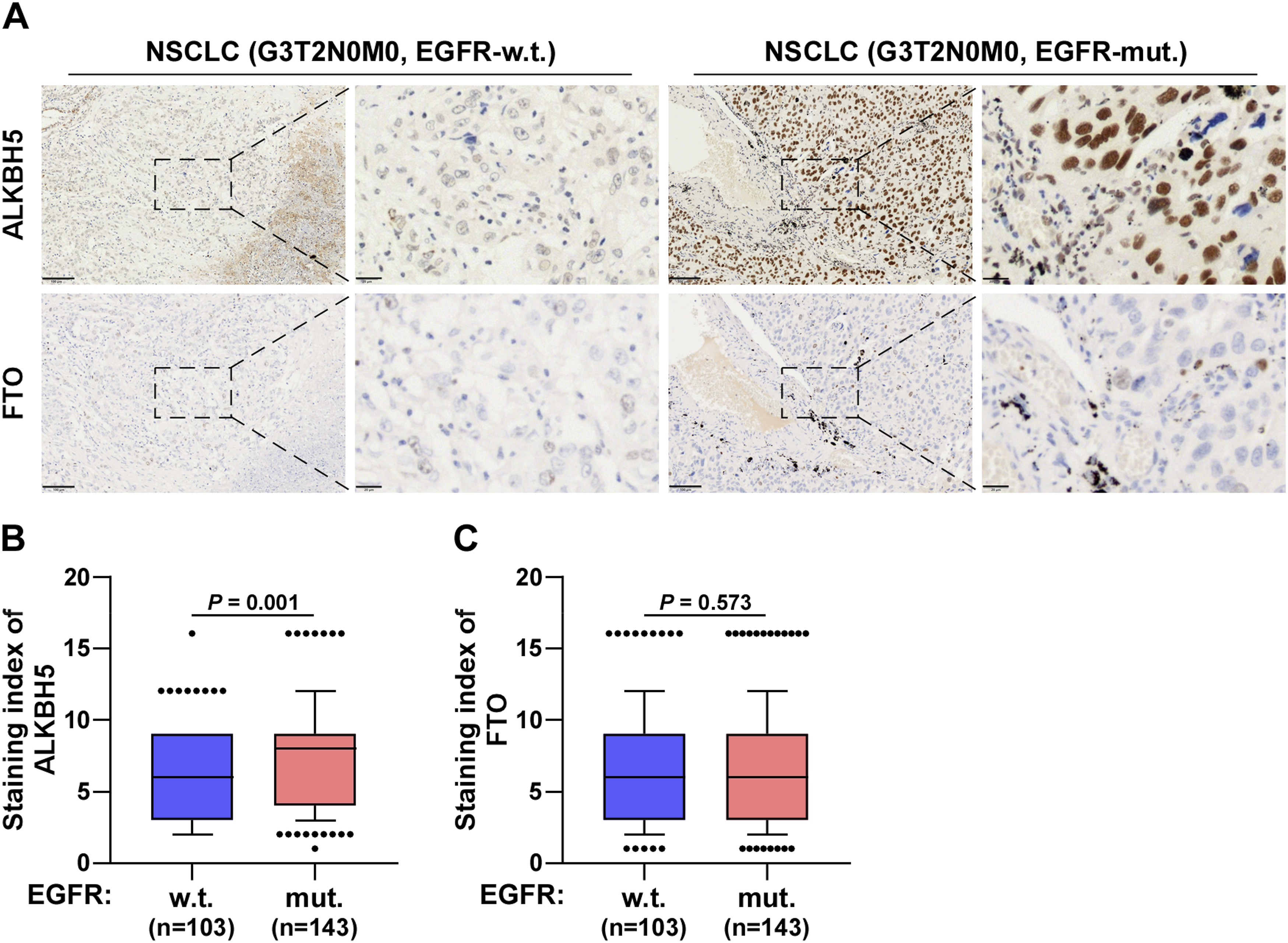
“Readers”
The “reader” in m6A can recognize modifications and combine with the “writers” and “erasers.” Different readers have different biological functions. The “readers” including YTHDC1, YTHDC2, YTHDF1, YTHDF2, and YTHDF3 were positively expressed in NSCLC tissues (Figure 4A). No significant differences were noted in the expression of these five genes between the wild-type and mutant groups (Figure 4B-F). The Expression of m6A “Readers” in EGFR Wild Type and EGFR Mutated NSLCL Tissues. (A) Typical Pictures of Immunohistochemical Staining of m6A “Readers” in EGFR-w.t. and EGFR-Mut. NSCLC Tissues. The Scale Bar in the Images Represents 100 μm and 20 μm Respectively. Images Were Collected at a 40× Objective Magnification (Numerical Aperture 0.75) Using an M8 Digital Microscope. (B) Immunohistochemical Staining Index of YTHDC1 in EGFR-Mut. Tissues Compared With EGFR-w.t. (w.t., n = 103; EGFR-Mut., n = 143). (C) Immunohistochemical Staining Index of YTHDC2 in EGFR-Mut. Tissues Compared With EGFR-w.t. (w.t., n = 103; EGFR-Mut., n = 143). (D) SI of YTHDF1 in EGFR-Mut. Tissues Compared With EGFR-w.t. (w.t., n = 103; EGFR-Mut., n = 143). (E) Immunohistochemical Staining Index of YTHDF2 in EGFR-Mut. Tissues Compared With EGFR-w.t. (w.t., n = 103; EGFR-Mut., n = 143). (F) Immunohistochemical Staining Index of YTHDF3 in EGFR-Mut. Tissues Compared With EGFR-w.t. (w.t., n = 103; EGFR-Mut., n = 143). Each Bar Represents the Median Values ± Quartile Values. 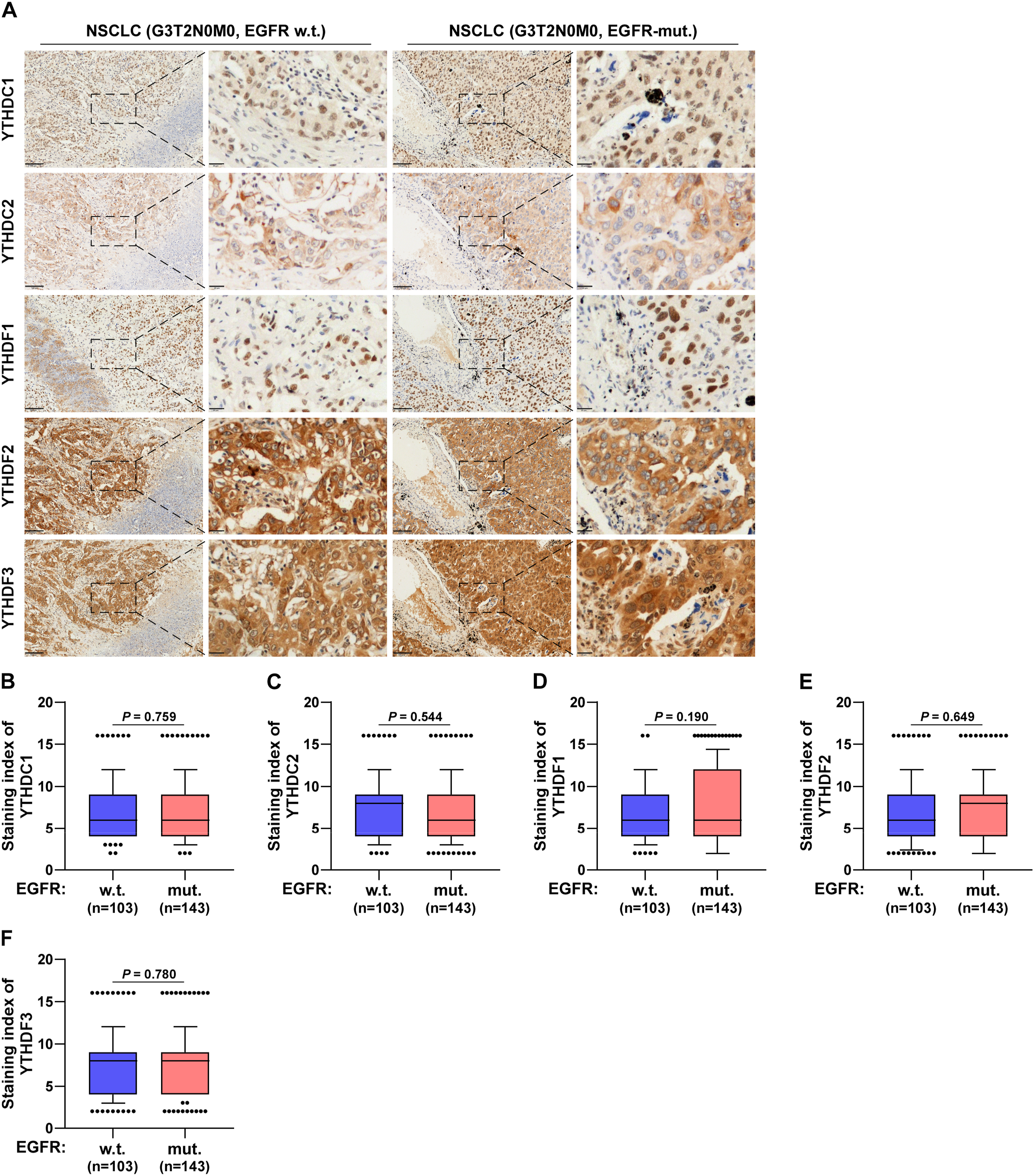
Thus, only METTL3 and ALKBH5 of the tested 11 m6A regulators showed different expression levels in wild-type and EGFR-mutant NSCLC tissues. These genes were significantly upregulated in the EGFR-mut group.
Relationships Between METTL3 and ALKBH5 Expression, Clinicopathologic Features, and Prognoses
METTL3 and ALKBH5 were upregulated in EGFR-mutant NSCLC. Among the 73 cases of the 19del, 57 cases of L858R, and 13 cases of other mutations, METTL3 showed no statistically significant differences between the three mutation types (Figure 5A), whereas ALKBH5 was significantly upregulated in the 19del and L858R mutations (Figure 5D). Compared with the G1 group, METTL3 and ALKBH5 were significantly upregulated in the G3 group, with no statistical difference in the G2 group (Figure 5B and F). In addition, METTL3 and ALKBH5 were significantly upregulated during stages II-IV (Figure 5C and E). These results suggest that the expression of METTL3 and ALKBH5 increased, followed by tumor development. The METTL3 low group (SI ≤6; n = 75) exhibited significantly longer median progression-free survival time of 55.2 months than the METTL3 high group (SI >6; n = 68) of 25.0 months (Figure 5G). The ALKBH5 low group (SI ≤6; n = 68) was found to have significantly longer median progression-free survival time of over 60.0 months than the ALKBH5 high group (SI >6; n = 75) of 24.1 months (Figure 5H). The Relationship Between METTL3 and ALKBH5 Expression, Clinicopathologic Feature and Prognosis. (A) Immunohistochemical Staining Index of METTL3 in EGFR-Mut. NSCLC Tissues Compared With Different Mutant Subtypes. (B) The Expression of METTL3 in Different Pathological Grades. (C) The Expression of METTL3 in Different Tumor Stages. (D) Immunohistochemical Staining Index of ALKBH5 in EGFR-Mut. NSCLC Tissues Compared With Different Mutant Subtypes. (E) The Expression of ALKBH5 in Different Pathological Grades. (F) The Expression of ALKBH5 in Different Tumor Stages. (G) Kaplan–Meier Analysis of Progression-free Survivals of EGFR-Mut. Patients. Patients Were Divided Into METTL3 Low Group (Blue, n = 75) and METTL3 High Group (Red, n = 68). (H) Kaplan–Meier Analysis of Progression-Free Survivals of EGFR-Mut. Patients. Patients Were Divided Into ALKBH5 Low Group (Blue, n = 68) and ALKBH5 High Group (Red, n = 75). Each Bar Represents the Median Values ± Quartile Values. 
High Expression of METTL3 and ALKNH5 is the Independent Risk Factor for EGFR-Mut NSCLC Patients
Univariate Cox regression analysis suggested that grade 3, stage II-IV, METTL3 high, and ALKBH5 high were related to PFS (Figure 6A). Multivariate Cox regression analysis indicated that METTL3 high and ALKBH5 levels were independent risk factors for PFS (Figure 6B). Compared to the 0-point group (blue; METTL3 and ALKBH5 low), when METTL3 high and ALKBH5 high coexisted (2-point; green), the PFS significantly decreased ( The High Expression of METTL3 and ALKNH5 is the Independent Risk Factor for EGFR-Mut NSCLC Patients. (A) Univariate COX Regression Analysis Indicates G3, Stage II-IV, METTL3 and ALKBH5 High are Significantly Correlated With PFS (n = 143). (B) Multivariate COX Regression Analysis Indicates METTL3 and ALKBH5 High are Significantly Correlated With PFS (n = 143). (C) Kaplan–Meier Analysis of Progression-Free Survivals of EGFR-Mut. NSCLC Patients. Patients Were Divided Into 0,1, and 2 Point Groups as METTL3 and ALKBH5 High Taking 1 Point Singly But 0 Point on the Contrary (0 Point, Blue; 1 Point, Red; 2 Points, Green; n = 143). 
Discussion
Lung cancer is the second most common cancer worldwide, and NSCLC is the most common lung cancer subtype with a high rate of metastasis and drug resistance. 21 In this study, METTL3 and ALKBH5 were upregulated in EGFR-mutant NSCLC, and their expression was related to the stage and pathological grade. High METTL3 and ALKBH5 expression levels may be related to shortened PFS and are predicted to be independent risk factors for EGFR-mutant NSCLC.
In recent years, compared with platinum-based chemotherapy, oncogene-targeted therapy has shown great efficacy in improving the quality of life and survival of patients. Thus, the mutation-related molecular mechanisms of oncogenes such as EGFR, TP53, and KRAS have attracted increasing attention. The modification of m6A methylation plays an important role in tumor progression-related molecules and pathways. The m6A modification process consists of the “writer” (methyltransferase), “eraser” (demethylase), and “reader” (m6A binding protein).18,19 The upregulation of the methyltransferase METTL3 and demethylase ALKBH5 promotes the development of lung cancer. METTL3 can participate in TGF-β-induced epithelial-mesenchymal transition (EMT) by regulating JUNB, a transcriptional EMT in lung cancer. 22 ALKBH5 promotes autophagy in NSCLC cells by targeting UBE2C to inhibit demethylation. 23 EGFR is the most common driver gene in NSCLC, and both METTL3 and ALKBH5 participate in its translation and expression. METTL3 promotes the proliferation of NSCLC cells by regulating EGFR mRNA translation,24,25 whereas ALKBH5 promotes tumor cell proliferation by destabilizing IGF2BP target genes. 26
Therapies targeting driver genes are on the rise. The Chinese Society of Clinical Oncology (CSCO) guidelines recommend EGFR site detection in lung cancer. EGFR-TKIs are widely used to treat NSCLC patients harboring EGFR mutations. However, drug resistance remains a challenge. The drug resistance mechanisms of EGFR-TKIs include both EGFR-dependent and independent mechanisms. The m6A modification has been reported to be involved in drug resistance. METTL3 can bind to MET to activate the PI3K/AKT signaling pathway, thereby reducing the sensitivity to gefitinib. 27 METTL3 promotes oxitinib resistance in lung adenocarcinoma by activating m6A modifications and enhancing YAP1 stability. 28 Only 0.8-5% mutation frequency of m6A-related regulatory genes was found in NSCLC; however, METTL3 and ALKNH5 were upregulated in EGFR-mut NSCLC tissues compared to EGFR-wt NSCLC tissues, indicating that the homeostatic cycle of m6A production and decomposition is frequently activated. Thus, we speculate that the rapid activation cycle is related to the development of resistance to EGR-TKIs in NSCLC. Recently, the targeting of related proteins in the m6A cycle has made promising progress. For example, the METTL3 inhibitor STM2457 has shown excellent results in preclinical studies of acute myeloid leukemia. 29 circ-FBXW7 represses the m6A-dependent translation of Wnt signaling to enhance TKI sensitivity in lung adenocarcinoma. 30 These results suggest that targeting METTL3, ALKBH5, and other mechanisms of the m6A cycle could be used as new therapeutics for reducing drug resistance in NSCLC.
In addition, survival prognosis and Cox regression analyses revealed that NSCLC patients with high METTL3 and ALKBH5 expression and EGFR mutations had shorter median progression-free survival periods. High METTL3 and ALKBH5 expression levels are independent risk factors for early progression. This suggests that m6A modification occurs frequently and rapidly in EGFR-mutant NSCLC, indicating that m6A modification and EGFR mutations may have a synergistic effect. Therefore, the detection of METTL3 and ALKBH5 may effectively evaluate the prognosis of NSCLC and predict the risk of early progression. High expression of m6A-related regulatory factors, such as METTL3, 31 METTL14, 32 WTAP, 33 ALKBH5, 34 and FTO 35 is associated with poor prognosis of NSCLC. On the contrary, the high expression of m6A “readers” YTHDC2,36,37 YTHDF1, and YTHDF2 38 are associated with favorable prognoses. In this study, IHC results showed that none of the other nine proteins detected significantly differed among the EGFR-wt and mutant group, except METTL3 and ALKBH5. Univariate and multivariate Cox regression analyses suggested that only METTL3 and ALKBH5 were associated with PFS in EGFR-mutant NSCLC patients. Thus, METTL3 and ALKBH5 may be more effective prognostic biomarkers than other m6A-related regulatory factors in EGFR-mutant NSCLC. Previous studies have shown that m6A regulators can be used as prognostic predictors in TP53‐mutant NSCLC patients. 34 M6A related regulators METTL3, IGF2BP2, HNRNPC, and HNRNPA2B1 may be effective prognostic and diagnostic factors for lung adenocarcinoma. 39 However, one of the two studies was purely based on bioinformatics research, and the other was a study on lung adenocarcinoma without genetic mutations. In our study, large numbers of immunohistochemical experiments on 11 m6A related genes were conducted to analyze the relationship between their expression and prognosis in patients with EGFR-mutant NSCLC, and further confirmed that high expression of METTL3 and ALKBH5 may be a risk factor for poor prognosis in these patients.
However, this study had some limitations. The m6A modification process involves multiple proteins, but only 13 were selected. The relationship between m6A-associated regulators and common driver gene mutations in lung cancer, except for EGFR mutations, has not yet been investigated. No cellular or animal experiments were conducted to verify these conclusions. Therefore, NSCLC specimens with mutations in other common driver genes, such as ALK and ROS1, will be collected for analysis, and further cellular and animal experiments will be planned to investigate the relationship between m6A-related regulators and EGFR-TKI drug resistance.
Conclusions
In this study, METTL3 and ALKBH5 were upregulated in EGFR-mutant NSCLC, and the PFS of patients with high METTL3 and ALKBH5 expression was shorter than that of patients with low METTL3 and ALKBH5 expression. METTL3 and ALKBH5 were predicted to be independent risk factors for EGFR-mutant NSCLC. These findings deepen our understanding of the clinical significance of m6A-associated regulators in NSCLC and help us to further explore the predictive biomarkers and new therapeutic targets.
Supplemental Material
Supplemental Material - M6A Methylation Regulators METTL3 and ALKBH5 are Risk Factors for EGFR-Mutant NSCLC
Supplemental Material for M6A Methylation Regulators METTL3 and ALKBH5 are Risk Factors for EGFR-Mutant NSCLC by Yaofeng Zhi, Silin Liu, Xuefei Chang, Wanxian Guan, Ronggang Li, Qiongru Liu, Jiaqing Chen, Jie Ling, Xulin Zhao, Aibin Liu, Jiarong Chen, Xin Zhang and Yanming Huang in Cancer Control
Footnotes
Acknowledgements
Thanks to the Department of Pathology of Jiangmen Central Hospital for the guidance and help in the interpretation of immunohistochemistry.
ORCID iDs
Ethical Statement
Author Contributions
All authors contributed to the study conception and design. Experimental studies, data analysis, statistical analysis and manuscript preparation were performed by Xin Zhang, Yaofeng Zhi, Silin Liu and Yanming Huang. Xuefei Chang, Wanxian Guan, Ronggang Li, Qiongru Liu, Jiaqing Chen, Jie Ling, Xulin Zhao, Jiarong Chen and Aibin Liu worked on the experimental studies, data analysis and statistical analysis. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Basic and Applied Basic Research Foundation of Guangdong Province (No. 2021A1515220180), National Natural Science Foundation of China (NO. 82273384, NO. 82203748 and NO. 82373057), Natural Science Foundation of Hunan Province (2022JJ30942) and Scientific Research Project of Hunan Health Commission (202203104934).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
