Abstract
It has been widely reported that periodontitis may lead to bone tissue and teeth loss and result in failure of prosthodontics or implants. Interleukin-1 (IL-1) is a potent proinflammatory cytokine that plays an essential role during the pathogenesis of periodontitis. However, the gene polymorphisms of IL-1α, IL-1β and IL-1RN and the relationship between these protein expressions in healthy people and patients with chronic periodontitis (CP) in China have not been fully elucidated. We investigated the gene polymorphisms and protein expression of IL-1α, IL-1β and IL-1RN in healthy subjects and CP patients, and our data suggest that these gene polymorphisms are associated with CP. The frequency of the C/C genotype of IL-1α was 55% in CP patients, while in the control group it was 20% (p<0.0001). The C/C genotype of IL-1β was also higher in CP patients (51%) than in controls (21%) (p<0.0001). For the 2/2 genotype of IL-1RN, CP patients showed a 30% frequency, while in controls this was 15% (p<0.0001). Protein levels evaluated by enzyme-linked immunosorbent assay demonstrated a significant difference in secretion between patients and controls for IL-1α and IL-1β. These results indicate that genotype and protein production of IL-1α, IL-1β and IL-1RN are associated with CP in a Chinese population, and might be putative risk indicators for chronic periodontitis.
Introduction
Periodontitis is a chronic inflammatory disease initiated by gram-negative microorganisms in the plaque biofilms and mediated by host responses (1). The progression and outcome of this disease is highly dependent on the individual. The results of several twin studies suggest that genetic factors may be responsible for the clinical variance in periodontitis (2, 3). Current genetic research in periodontitis has focused on gene polymorphisms of cytokines, chemokines, receptors and enzymes that play essential roles in the innate, inflammatory and immunological response to this disease. These include the interleukin (IL)-1 family, IL-4, IL-6, IL-10, tumor necrosis factor, transforming growth factor-β1, and others (4).
Potent proinflammatory agents, such as the IL-1 family including IL-1α, IL-1β and IL-1 receptor antagonist (IL-1Ra), have been reported to initiate and regulate the pathogenesis of periodontitis (5). IL-1α and IL-1β are agonists that bind to a specific receptor, which can result in extracellular matrix degradation and alveolar bone resorption. IL-1Ra is an antagonist that regulates the activities of IL-1α and IL-1β by competitively inhibiting binding to the same receptor (6, 7).
IL-1α possesses a wide spectrum of biological functions and plays critical roles in the regulation of immune responses. Studies show that overexpression of IL-1α may cause periodontal disease (8). IL-1β, the predominant form of IL-1, stimulates a wide range of cell types to produce protease and mediators in different periodontal diseases. IL-1Ra is a 22-kDa protein encoded by the IL-1RN gene, which binds competitively to the same IL-1 receptor without triggering intracellular signaling. It is also important for normal physiology and pathological states by acting as an antagonist. Studies have shown that the balance between IL-1β and IL-1Ra is closely related to the severity of periodontitis (9, 10).
Because of its essential biological function during the development of periodontitis, the IL-1 gene cluster appears to be the most investigated gene related to periodontal disease. The IL-1 gene cluster is located on human chromosome 2q13 (11). IL-1 gene polymorphisms were demonstrated to be associated with clinical variance in periodontal disease, such as severity, tooth loss (12), and bleeding on probing (BOP) (13). Several studies show that IL-1 gene polymorphisms could be an important risk factor for periodontitis in different populations (14, 15). Owing to multiple factors and different criteria for patient selection, the identification of genetic risk factors for periodontitis is complex. First reported by Kornman et al (16), the periodontitis-associated genotype of IL-1 contributes to periodontal disease severity in Caucasians. This result was confirmed by Galbraith et al and Gore et al (17, 18). However, studies in Chinese, Greek and Thai populations failed to find a relationship between IL-1 genotype and chronic periodontitis (CP) susceptibility or severity (19-21).
The aim of this study was to investigate single nucleotide polymorphisms (SNPs) in the IL-1α and IL-1β genes at positions −889 (C/T), −511 (C/T) and an 86-bp variable number of tandem repeats (VNTR) polymorphism in intron 2 of IL-1RN in CP patients and healthy controls. The protein production of IL-1α and IL-1β was also investigated.
Materials and Methods
Subject enrollment
A total of 491 Chinese subjects including 399 healthy individuals and 92 CP patients were recruited for this case-control study. Written informed consent was obtained from all enrolled subjects and clinical examinations were performed at the West China Hospital of Stomatology, Sichuan University (SCU). All individuals were confirmed to be free from systemic diseases such as diabetes, hypertension and cardiovascular disease. Subjects with a past or current smoking habit, swelling of lymph nodes, temporomandibular joint disease, and soft tissue abnormalities were excluded from the study. According to the inclusion criteria, only subjects with a definite diagnosis of CP and clinically and radiographically proven periodontitis-free subjects were included.
Intraoral examination of periodontal conditions was performed in 399 healthy subjects. Clinical parameters including physical examination results, medical and dental history, supragingival calculus, gingival recession, BOP, probing depth (PD), clinical attachment loss (CAL), tooth mobility and plaque were collected as well. Periodontal disease was diagnosed based on the 1999 International Classification of Periodontal Disease and Conditions (22). The study subjects were classified into 2 groups: a healthy group consisting of 399 subjects without any signs of periodontal disease in the absence of gingival recession, CAL and sites with PD >3 mm; a CP group consisting of 92 patients who presented with PD >5 mm and a CAL >4 mm degree of gingival recession and tooth mobility. Blood samples of patients and control subjects were taken at SCU. The study protocol was approved by the SCU ethics committee.
Sample preparation
Blood samples were collected in lithium heparin tubes (Vacuette, Austria) and centrifuged for 10 minutes at 1500 rpm. Plasma was removed for enzyme-linked immunosorbent assay (ELISA) analysis. The remaining cells were transferred to a 50 mL centrifuge tube along with red blood cell lysis buffer, up to a final volume of 45 mL. The mixture in the tube was inverted several times then centrifuged for 10 minutes at 1500 rpm. The supernatant was discarded and the remaining cells were washed with 0.9% PBS to be used for DNA extraction.
Extraction of DNA and PCR
Genomic DNA was extracted from all samples using a commercially available Genomic DNA Mini Kit (Qiagen, USA), following the manufacturer's instructions. The extracted DNA was labeled and stored at −80°C until use.
Polymorphism in the IL-1α gene at position C -889T
A PCR Master Mix kit was used in the study (Promega, USA). All procedures were carried out in a sterile and stable environment to prevent external contamination. The promoter region containing the polymorphic site of IL-1α was amplified using the following primers: 5′-AAG CTT GTT CTA CCA CCT GAA CTA GGC-3′ and 5′-TTA CAT ATG AGC CTT CCA TG-3′ (16, 23). Twenty units of nuclease-free water, 25 units of master mix, 0.5 units of each primer and 3 units of the extracted DNA sample were mixed to undergo thermal cycling. The thermal cycler (MJ, Bio-Rad, USA) was programmed at 94°C for 5 minutes of denaturation, 30 cycles of 94°C for 30 seconds, 56°C hybridization for 30 seconds and 72°C for 1 minute and a final extension for 5 minutes at 72°C. Products from the thermal cycling were labeled accordingly and stored at −80°C until use. Restriction fragment length polymorphism was performed with 3 units of NcoI at 37°C overnight to detect allele C (16 bp + 83 bp) and allele T (99 bp).
Polymorphism in the IL-1β gene at position C -511T
IL-1β was amplified using the primers 5′-TGG CAT TGA TCT GGT TCA TC-3′ and 5′-GTT TAG GAA TCT TCC CAC TT-3′ (14, 24). Amplification reactions were performed with 500 ng genomic DNA in a total volume of 50 μL, containing 10 mM Tris–HCl (pH 8.3), 50 μM KCl, 1 μM of each primer, 200 μM each of dATP, dCTP, dGTP and dTTP, 4 mM MgCl2, and 2.5 U Taq DNA polymerase. The cycling consisted of 2 cycles of 2 minutes at 95°C, 1 minute at 53°C and 1 minute at 74°C, followed by 38 cycles of 1 minute at 95°C, 1 minute at 53°C and 1 minute at 74°C. Products from the thermal cycling were labeled accordingly and stored at −80°C until use. The products were digested with 3 units of AvaI at 37°C overnight to detect allele C (114 bp + 190 bp) and allele T (304 bp).
Polymorphism in the IL-1RN (intron 2) gene – VNTR
IL-1RN was amplified using the primers 5′-CTC AGC AAC ACT CCT AT-3′ and 5′-TCC TGG TCT GCA GGT AA-3′ (25). Amplification reactions were performed with 500 ng genomic DNA in a total volume of 50 μL, containing 10 mM Tris-HCl (pH 8.3), 50 mM KCl, 1 μM of each primer, 200 μM of dATP, dCTP, dGTP and dTTP, 1.5 mM MgCl2, and 2.5 U Taq DNA polymerase. The reaction was incubated for 1 minute at 96°C, followed by 35 cycles of 1 minute at 94°C, 1 minute at 60°C and 2 minutes at 70°C, and an additional 5 minutes at 70°C and 5 minutes at 55°C. In the second intron of the IL-1RN gene, 2 alleles were defined by different numbers of an 86-bp segment repeat. Genotypes were determined by comparing the size of the bands with a molecular weight ladder, with separation into allele 1 (4 repeats −410 bp) and allele 2 (2 repeats −240 bp).
Electrophoresis and visualization of digest product
Five microliters of amplicons and 1 mL of Ready-Load 1 kb DNA ladder (Invitrogen, Spain) were loaded into 3% agarose gel (Invitrogen, Spain) containing 0.5 mg/mL of ethidium bromide. The gel underwent electrophoresis at 100V, 100 mA for 30 minutes, and it was visualized using a Dolphin-DOC ultraviolet illuminator (Wealtec, South Africa).
ELISA assay
Plasma samples were used to perform ELISA analysis according to the manufacturer's instructions of the IL-1α and IL-1β ELISA kits (Diaclone, France). The procedures were as follows: 50/100 μL of the standard group solutions and plasma of each subject were pipetted into a 96-well plate. The plate was incubated for 2 to 3 hours at 350 rpm and washed with washing buffer 3 times. The wells were then dried and 200 mL of substrate tetramethylbenizidine was added into each well for 20 minutes in the dark at room temperature. The plates were read at 450 nm wavelength using a Sunrise universal microplate reader (Tecan, Austria). The levels of IL-1α and IL-1β in the samples were obtained by comparison with the standard curve generated from standards supplied by the manufacturer.
Statistical analysis
The differences in genotype and allele frequencies and carriage rates between the CP and control groups were examined by the chi-squared test (χ 2 ). For risk analysis of alleles and genotypes, the odds ratio (OR) with 95% confidence intervals (95% CI) was calculated. The soluble protein levels of IL-1α and IL-1β were evaluated with an independent t-test, and a p value of <0.05 was considered statistically significant. Statistical analysis was performed by SPSS 18.0 for Windows (SPSS Inc., USA).
Results
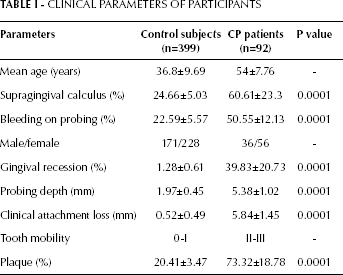
The clinical parameters of CP patients and healthy controls are summarized in Table I. Healthy subjects and CP patients had a mean age (± standard deviation [SD]) of 36.8±9.69 and 54±7.76 years, respectively. The percentage of supragingival calculus was significantly higher in the CP group than the control group (60.61% vs 24.66%, p=0.0001). There were also significant differences in BOP, gingival recession, PD, CAL and plaque between the CP group and the control group (p<0.0001).
CLINICAL PARAMETERS OF PARTICIPANTS
Distribution of IL-1α and IL1-β genotypes and allele frequencies
The distribution of IL-1α polymorphisms among the CP and control groups is depicted in Table II. For IL-1α −899T, homozygous C alleles were represented by 2 DNA bands with sizes of 76 and 153 bp; homozygous T alleles were represented by 3 DNA bands with sizes of 76, 29 and 124 bp. For heterozygotes of IL-1α, −899T displayed a combination of both C and T alleles (16, 153, 29 and 124 bp). There was a significant difference between the CP and control groups in the distribution of IL-1α genotypes (χ 2 84.42, p<0.0001). In the C/C genotype, the CP group presented a frequency of 55.4% compared to 20.0% in the control group. The OR for carriage of the IL-1α allele (T/T and C/T genotypes combined compared with the C/C genotype) was 4.89 (95% CI, 3.23-7.40, χ 2 62.48, p<0.0001) in the CP group (Tab. II). There was a significant difference between groups in allele frequencies of IL-1α (χ 2 7.80, p=0.005; OR1.51, 95% CI, 1.13-2.02). The distribution frequency of homozygous C alleles of IL-1α was higher in the CP group (64.7%) than in the control group (55.3%).
GENOTYPE AND ALLELE FREQUENCY OF IL-1α AND IL-1β IN ALL STUDY PARTICIPANTS
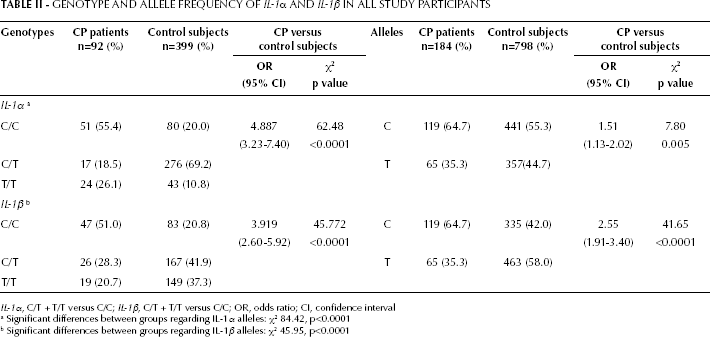
IL-1α, C/T + T/T versus C/C; IL-1β, C/T + T/T versus C/C; OR, odds ratio; CI, confidence interval
Significant differences between groups regarding IL-1α alleles: χ2 84.42, p<0.0001
Significant differences between groups regarding IL-1β alleles: χ2 45.95, p<0.0001
The polymorphism distribution of IL-1β in the CP and control groups is also shown in Table II. For IL-1β −511T, homozygous C alleles were represented by 2 DNA bands with sizes of 190 and 114 bp, while the homozygous T allele was represented by 3 DNA bands with a size of 304 bp. For heterozygotes of IL-1β −511T displayed a combination of both C and T alleles (190, 114, and 304 bp). There was a significant difference between groups in the distribution of IL-1β genotypes (χ 2 45.95, p<0.0001). In the C/C genotype, the CP group presented a frequency of 51.0%, compared to 20.8% in the control group. The OR for carriage of the IL-1β allele (T/T and C/T genotypes combined compared with the C/C genotype) was 3.92 (95% CI, 2.60-5.92, χ 2 45.77, p<0.0001) in the CP group. The distribution frequency of homozygous C alleles of IL-1β was higher in the CP group (64.7%) than in the control group (42.0%). The OR was 2.55 (95% CI, 1.91-3.40, χ 2 41.65, p<0.0001) between the CP and control groups (Tab. II).
Distribution of IL-1RN genotypes, allele frequency and IL-1RN2 allele positivity
With regard to the distribution of IL-1RN genotypes, the difference between the CP and control groups was significant (χ 2 15.23, p<0.0001). The IL-1RN2 genotype frequency was 29.5% in the CP group and 14.5% in the control group. The OR for carriage of the IL-1RN allele (RN1/2 and RN1/1 genotypes combined compared with the RN2/2 genotype) was 2.44 (95% CI, 1.54-3.85, χ 2 15.20, p<0.0001). The frequency of IL-1RN2-positive genotypes of the IL-1RN gene was also significantly higher in the CP group than the control group (χ 2 12.69, p<0.0001; OR 2.19, 95% CI, 1.41-3.39). The distribution frequency of RN2 alleles of IL-1RN2 was higher in the CP group (31.5%) than in the control group (16.5%) (χ 2 27.66, p<0.0001; OR 2.30, 95% CI, 1.68-3.15) (Tab. III).
GENOTYPE DISTRIBUTIONS AND ALLELE FREQUENCIES IN IL-1RN2-POSITIVE AND IL-RN2-NEGATIVE STUDY PARTICIPANTS
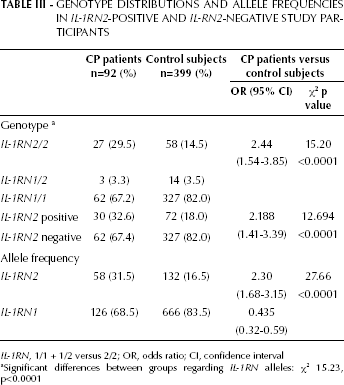
IL-1RN, 1/1 + 1/2 versus 2/2; OR, odds ratio; CI, confidence interval
Significant differences between groups regarding IL-1RN alleles: χ2 15.23, p<0.0001
Clinical parameters (mean ± SD) of IL-1RN2-positive and IL-1RN2-negative CP subgroups
We also investigated the relationship between the IL-1RN2 allele and different clinical parameters to find out whether IL-1RN2 positivity and negativity were associated with clinical variance. The nonparametric Mann-Whitney U test showed no significant differences in clinical parameters between subjects showing IL-1RN2 positivity and IL-1RN2 negativity within the CP group (p>0.05) (Tab. IV).
CLINICAL PARAMETERS (MEAN ± SD) OF THE CP GROUP DIVIDED INTO IL-1RN2-POSITIVE AND IL-1RN2-NEGATIVE SUBJECTS
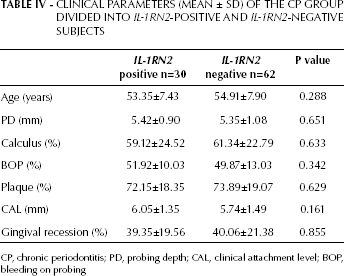
CP, chronic periodontitis; PD, probing depth; CAL, clinical attachment level; BOP, bleeding on probing
Protein production of IL-1α and IL-1β
The serum level of IL-1α was significantly higher in the CP group than in the control group (60.42 pg/mL ± 11.01 pg/mL vs 23.95 pg/mL ± 4.55 pg/mL, p=0.0001) (Fig. 1A). The serum level of IL-1β was also higher in the CP group than in the control group (79.56 pg/mL ± 17.28 pg/mL vs 19.51 pg/mL ± 3.74 pg/mL, p=0.0001) (Fig. 2A). There was no significant difference between the serum expression amongst genotypes of each biomarker in the CP group (Fig. 1B and 2B).
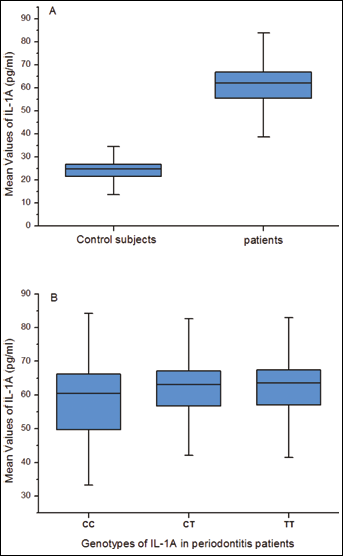
Serum levels of IL-1α in patients with periodontitis and control subjects. (A) Comparison of the serum level of IL-1α between patients and controls. (B) Serum genotype distribution of IL-1α in patients with periodontitis.
Discussion
The quality and quantity of alveolar bone are critical factors that affect the final outcomes of prosthodontics and implants. Patients with periodontitis have an increased failure rate of dental implants (26). In order to prevent the bone loss caused by periodontitis at an early stage, the present study aimed to find out whether gene polymorphisms are risk indicators of CP.
Periodontitis is a multifactorial chronic inflammatory disease, and genetic factors may play a critical role in host responses to this disease (27, 28). More than 20 candidate genes have been investigated and the polymorphisms of these genes were also found to be associated with both aggressive and chronic periodontitis (4). The IL-1 gene cluster might be the most studied among these genes.
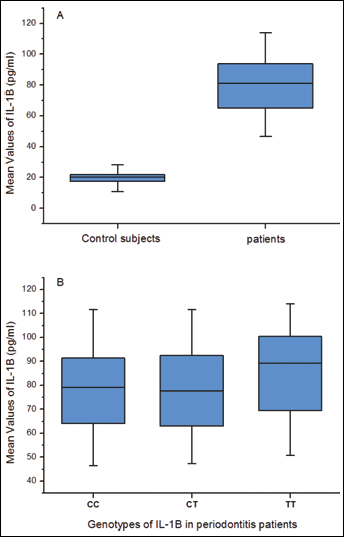
Serum levels of IL-1β in patients with periodontitis and control subjects. (A) Comparison of the serum level of IL-1β between patients and controls. (B) Serum genotype distribution of IL-1β in patients with periodontitis.
In our study, SNPs in the IL-1α −889, IL-1β −511 and IL-1RN (intron 2) genes, and IL-1α and IL-1β protein production in patients with CP and healthy people were analyzed. Susceptibility to CP was also investigated. This study has demonstrated a significant difference between the CP group and the healthy control group in association with IL-1α −889 polymorphic sites in a Chinese population. The C/C homozygous genotype was approximately 55% (51/92) in the CP group, much higher than in the control group (20%; 80/399) (p<0.0001). The C allele frequency was higher than the T allele frequency by 30% (Tab. III). The SNPs of IL-1α were, as previously reported, associated with periodontitis (16, 17, 29, 30). Another study, however, failed to find positive connections to periodontitis, but showed that the level of IL-1α was highly elevated in the gingival crevicular fluid of patients with periodontitis (31).
In the present study, ELISA results showed that protein production of IL-1α in CP patients was significantly higher than in the healthy group (Fig. 1A). It might indicate that genetic variations may increase the levels of protein production. However, the different genotypes seem not to be associated with protein production in the CP group (Fig. 1B). The absence of genotypic variance might indicate that there is no genetic predisposition to the production of IL-1s among CP patients.
As for the IL-1β −511 polymorphism, a significant difference was also observed in the genotype and allele frequencies between the CP and control groups. The C/C genotype was approximately 51% (47/92) in the CP group, much higher than in the control group, where it accounted for 21% (83/399) (p<0.0001). Our results were supported by other studies, which demonstrated a positive association with the severity of PD (16, 32). A meta-analysis also demonstrated a weak positive relation between IL-1β and PD (33). Other studies involving different population samples demonstrated no association between IL-1β polymorphism and PD (14, 24).
The protein production of IL-1β was found to be higher in the CP group than in the control group (Fig. 2A). Interestingly, the T/T genotype was associated with the highest protein production compared to the C/C and C/T genotypes in the CP group (Fig. 2B). The results of IL-1β expression of previous studies were controversial. Pociot et al reported that increased IL-1β secretion was correlated with a genetic polymorphism at IL-1β +3954 in periodontitis patients (34). However, Mark et al demonstrated no association of this genotype with monocyte IL-1β expression in heterozygous individuals with mild to moderate adult periodontitis (35). The possible reason for the differences might be the different criteria of subject selection.
Regarding IL-1RN (intron 2) polymorphism, the genotype 2/2 was found to be associated with susceptibility to CP in a Chinese population. The carriage rate of allele 2 was approximately 33% in CP patients, similar to what was observed for Brazilians (37%) according to Trevilatto's research results (14). While in the healthy control group allele 1 of the IL-1RN genotype occurred much more frequently (82%), this result was also similar to Berdeli's finding (13). The frequency and carriage rate of allele 2 was found to be increased not only in periodontitis but also in other chronic inflammatory diseases such as rheumatoid arthritis, inflammatory bowel disease, systemic lupus erythematosus, and nephropathy (4, 36, 37). However, a recent study in a Chinese population demonstrated a similar carriage rate of the IL-1RN2 allele between the periodontitis group and healthy cases (38).
The association of clinical parameters and the IL-1RN2 allele between the CP and the control group was also evaluated (Tab. IV). However, we failed to find any positive connection between clinical variations and the IL-1RN2 allele. Berdeli showed that the percentage of BOP was significantly higher in IL-1RN2-positive subjects in the CP group (13). Studies also demonstrated that IL-1RN2 might regulate the balance between IL-1Ra and IL-1β by enhancing the release of IL-Ra and decreasing the secretion of IL-1β (39, 40). By contrast, other studies showed different results regarding the association of IL-1RN2 and secretion of IL-1Ra and IL-1β (41, 42).
A vast number of papers have demonstrated the role of genetic variations and environmental factors in diseases. The susceptibility of individuals to periodontitis is also based on this gene-environment interaction. This might explain the different proportions of this disease in different population groups. Thus, identifying the candidate genes that contribute to the pathogenesis of periodontitis can be significantly important. SNPs are found to be the most common source of genetic variations. The SNPs in candidate genes related to periodontitis should be identified for more accurate diagnosis as well as improving prognosis and developing novel therapies.
In conclusion, the findings of the present study support the association of IL-1α, IL-1β and IL-1RN gene polymorphisms and the secretion of proteins with CP in a Chinese population. The results suggest that individuals with C/C allele of IL-1α, IL-1β and IL-1RN2 may increase the risk of developing CP. It is also worthwhile to explore the association between the extent of disease with the serum levels of IL-1, which might help with diagnosis. Further investigation into the use of SNPs of these genes as a risk indicator of this disease is warranted.
