Abstract
Objectives
A growing number of studies have reported that genes involved in the repair of DNA double-strand breaks might be cancer-susceptibility genes. The x-ray cross-complementing group 4 gene (
Methods
We retrieved case–control studies that met the inclusion criteria from PubMed, Web of Science, Embase, and China National Knowledge Infrastructure databases. Associations between rs1805377 and cancer risk were evaluated by odds ratios (ORs) using a random effects model and 95% confidence intervals (CIs) as well as sensitivity and subgroup analyses.
Results
After inclusion criteria were met, the meta-analysis involved 24 studies that included 9,633 cancer patients and 10,544 healthy controls. No significant association was found between rs1805377 and the risk of cancer (pooled OR = 1.107; 95% CI = 0.955–1.284) in the dominant genetic model. Similarly, no significant association was observed in the subgroup analysis.
Conclusions
Through this meta-analysis, we found no association between the rs1805377 polymorphism and cancer occurrence. This may provide useful information for relevant future studies into the etiology of cancer.
Objectives
The occurrence of cancer is currently increasing because of an aging population, the prevalence of smoking, the lack of physical activity, and other lifestyle factors. 1 Cancer is a cellular abnormality initiated by uncontrolled growth caused by an accumulation of damage or mutations in genetically-mediated factors and environmental factors, resulting in cells evading the signal-mediated controls of cell growth and death. 2 Genetic factors have a greater effect on cancer initiation than environmental and lifestyle factors, 3 and a number of potential susceptibility genes and variations have been examined and identified to participate in cancer occurrence.
DNA damage repair involves known molecular pathways such as single-strand damage repair, double-strand break repair, and damage reversal.
4
Present evidence suggests that genes participating in the repair of DNA double-strand breaks might also be involved in modifying the risk of various cancers.
5
Among these, the x-ray cross-complementing group 4 gene (
Full-length
Meta-analysis is a useful method for identifying a common effect when considerable variation exists in study findings.
19
Another advantage is the increased sample size resulting from pooling relevant studies, which can, to some degree, decrease the occurrence of a false-positive or false-negative association generated by random error. Previous meta-analyses have investigated the association between
Methods
Identification of appropriate studies
A search of English (PubMed, Web of Science, and Embase) and Chinese language (China National Knowledge Infrastructure) databases was carried out to identify appropriate studies for inclusion in the meta-analysis using the following keywords:
Inclusion criteria were case–control studies involving cancer patients and reports of ATM allele and/or genotype frequencies. In the case of overlapping datasets, the most recent study was included. Exclusion criteria were the omission of healthy controls or duplication of previous data. With respect to studies lacking inclusion data, the authors were contacted by email to obtain missing information.
Data extraction
Data analysis
Data extraction from the publications was performed independently by two investigators, Xin-yuan Zhang and Xiao-han Wei. Extracted data included the first author surname, publication year, geographic region, genotyping method, sample size, and number of genotypes reported for both patients and controls. Data pertaining to patient ethnicity, control source, and cancer type were also extracted with a view to determining the contributions of underlying characteristics to the study findings.
Trial sequential analysis (TSA)
TSA was performed to evaluate whether the present meta-analysis had a sufficient sample size to generate firm pooled results about the effect of interventions. Evaluation criteria and calculation parameters were based on previous studies.23,24 TSA was conducted using TSA software (version 0.9.5.10; (http://www.ctu.dk/tsa/).
Statistical analysis
The chi-square goodness-of-fit test was used to calculate the Hardy–Weinberg equilibrium of control genotypes (significant at the 0.05 level), and odds ratios (ORs) and 95% confidence intervals (CIs) were employed to evaluate the strength of the association between rs1805377 and cancer. To calculate the pooled estimates of the ORs and 95% CIs among the studies, a random effects model was used to resolve inter-study heterogeneity. 25
For the measurement of pooled ORs, three genetic models (allele contrast, dominant, and recessive) were employed. As described in a previous study, 26 OR1 (AA vs. aa), OR2 (Aa vs. aa), and OR3 (AA vs. Aa) were compared, where A is the risk allele, from which the most appropriate genetic model was selected.27,28
A Q statistic was used to evaluate the degree of inter-study heterogeneity, with the absence of heterogeneity being defined as
A sensitivity analysis was used to evaluate whether the pooled effect size was potentially influenced by a single study. Each study was omitted from the meta-analysis in turn, then significant alterations to the pooled effect size were evaluated.
Funnel plots were generated for each study to evaluate publication bias. The standard error of log(OR) was plotted against log(OR); when the plot was asymmetrical, bias was determined. Accordingly, for the determination of the degree of asymmetry, an Egger test was performed;
Stata version 10.0 (Stata Corp., College Station, TX, USA) was used to perform all statistical calculations.
In silico analysis
To predict the potential association between rs1805377 and
Results
Online literature databases were used to identify relevant publications for inclusion in the meta-analysis. Twenty-four publications were included according to the established inclusion criteria.13,32–54 A flow diagram of this process is shown in Figure 1. Subjects involved in the studies are not overlapping. These 24 case–control studies collectively contained 9,633 cancer patients and 10,544 unaffected controls. Individuals with different genetic backgrounds and different types of cancer were included. The main characteristics of the included studies are listed in Table 1. Genotype and allele frequencies of rs1805377 and the Hardy–Weinberg equilibrium (HWE) in the controls are summarized in Table 2. Of the 24 studies, four publications deviated significantly from HWE.35,42,44,51

Flow diagram of literature screening.
Baseline characteristics of qualified studies in this meta-analysis.

Distribution of genotype and allele frequencies of the

Abbreviation:
Meta-analysis
Pooled ORs (with 95% CIs) in dominant, recessive, homozygous codominant, heterozygous codominant, and allele contrast genetic models were employed to evaluate the association of the rs1805377 polymorphism with cancer risk (Table 3 and Figure 2). The dominant model was selected to perform the pooled analysis according to the selection criteria of genetic models. The pooled results showed that there was no association between rs1805377 and the risk of cancer. The summary OR under a random effects model was 1.107 (95% CI = 0.955–1.284). Subsequent subgroup analysis also failed to detect any association of rs1805377 with cancer risk among East Asian and Caucasian patients (Table 4). Moreover, no association between rs1805377 and cancer was observed by subgroup analysis with respect to the control source (hospital or population). However, subgroup analysis according to cancer type revealed an association between rs1805377 and gastric antrum adenocarcinoma, but not other cancer types (Table 4).
Summarized ORs with 95% CIs for the association of the XRCC4 rs1805377 polymorphism with cancer.

Note: n, the number of studies; OR, odds ratio; CI, confidence interval;

Forest plot of the association between the
Stratified analysis of the association of the
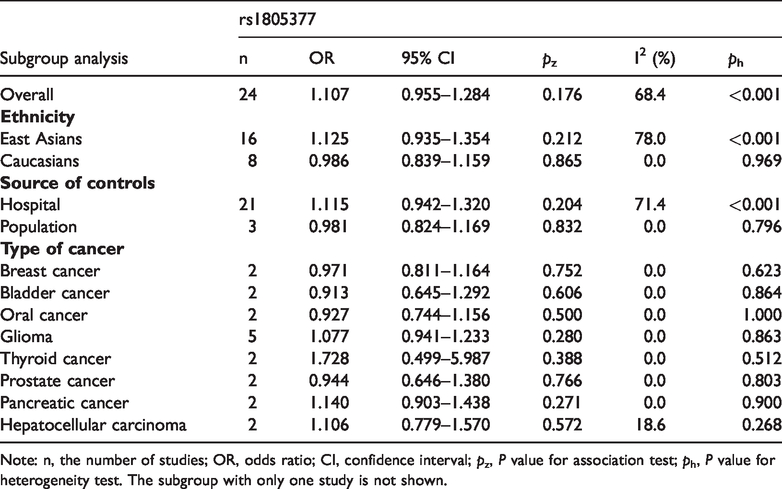
Note: n, the number of studies; OR, odds ratio; CI, confidence interval;
Sensitivity analysis
Sensitivity analysis was used to evaluate the extent to which each individual study contributed to the pooled results. Each study was eliminated from the analysis in turn, then pooled ORs were determined. No significant changes were detected between any of the analyses or the overall results; thus, we can be confident that the results of the meta-analysis display stability and reliability.
Assessment of publication bias
A funnel plot (Figure 3) was generated to assess publication bias, from which no significant effects were detected (Table 3).

Funnel plot analysis depicting publication bias in the association between the
TSA
In the overall analysis for dominant genetic model, the required sample size was 106,055 patients to reach the anticipated intervention effect (Figure 4). Results showed that the Z-curve did not cross the trail monitoring boundary, indicating that the present sample size was not sufficient and that further trials are required.

TSA for overall analysis under the dominant genetic model.
In silico analysis
eQTL analysis found that, compared with the A allele, the G allele of the rs1805377 locus leads to increased expression of

In silico analysis of
Discussion
The relationship between the
Previously, a putative association of rs1805377 with cancer occurrence was analyzed in three meta-analyses.20–22 While our meta-analysis overlaps somewhat with prior analyses, we included new analyses that have been conducted since these studies were published. Twenty-four studies were included to comprehensively investigate the role of rs1805377 in the occurrence of cancer. These consisted of patients with various types of cancer (breast cancer, bladder cancer, renal cell carcinoma, oral cancer, glioma, thyroid cancer, non-small-cell lung cancer, prostate cancer, gastric antrum adenocarcinoma, non-Hodgkin lymphoma, pancreatic cancer, hepatocellular carcinoma, and esophageal squamous cell carcinoma). With a view to evaluating the potential origins of heterogeneity and measuring stability, we performed subgroup analyses by ethnicity, control source, and cancer type. Therefore, to some extent, the final results of our meta-analysis are more accurate and comprehensive than previous meta-analyses.
There was considerable heterogeneity in our meta-analysis, which might reflect differences in genetic backgrounds. In subgroup analysis by ethnicity, we observed no significant heterogeneity in the East Asian subgroup, but strong heterogeneity in the Caucasian subgroup. This latter subgroup consisted of eight studies, including patients from the USA, Spain, Saudi Arabia, Germany, Portugal, and Australia. The observed heterogeneity may reflect the varied lifestyles and wide distribution of Caucasians, which can give rise to different cancer risks. 55
Our ability to conclusively define stable effects by subgroup, however, is limited by the relatively small sample size included in the subgroup analyses, particularly regarding cancers such as renal cell carcinoma, non-small cell lung cancer, gastric antrum adenocarcinoma, non-Hodgkin lymphoma, hepatocellular carcinoma, and esophageal squamous cell carcinoma. Our meta-analysis found an association between rs1805377 and the risk of gastric antrum adenocarcinoma, but this result should be interpreted with caution as only one study involving gastric antrum adenocarcinoma patients was included. Thus, we cannot conclude whether rs1805377 is associated with risk of cancer in these subgroups because of the limited sample size.
XRCC4 is required for non-homologous end joining, which is one of the major pathways for repairing DNA double-strand breaks. In its abnormal state it can lead to severe combined immunodeficiency,
9
but one reported patient with mutations in
Conclusion
The present study demonstrated no association between the XRCC4 rs1805377 polymorphism and cancer risk. Additional studies involving a wider range of ethnicities are now required to validate our subgroup analyses. Furthermore, environmental and epigenetic factors that contribute to cancer risk should also be studied.
