Abstract
The cryptic presence of a wide range of retroviruses with varying copy number holds biological significance for host reproduction and development. Most of the endogenous retroviruses with lost pathogenicity and replication ability still serve as transcriptional regulators of host cellular genes. These structural and functional features of proviruses present them as alternate promoters and enhancers for several host cellular genes involved in development and other biological processes. In addition, embryonic stem (ES) cells and induced pluripotent cells are known to effectively silence the expression of most of these proviruses through repressive epigenetic marks and proviral sequence heterochromatization, which is not a case for those of differentiated cells. In this review, we aim to dissect the underlying salient features of proviral silencing in embryonic stem cells and analyze the potential of these proviruses in cell fate determination.
Keywords
Introduction
The biology of retroviruses has been intensely studied and understood over the past decades. Most of the studies on retroviruses focus on their associated clinical complications and pathological manifestations. Distinctively, these viruses can pass their genetic information (RNA to DNA) in reverse to the usual direction for flow of biological information, ie, from DNA to RNA. 1 Most of the retroviruses have an RNA genome, which gets transcribed into a double-stranded DNA sequence within the host cellular machinery using reverse transcriptase (RT) and other polymerases (Fig. I). 1 Such a critical role of RT in infection and virus transmission makes these genomic entities crucial for grouping of viruses according to the similarity in the transcriptional binding and the mode of RT functioning. 2 Furthermore, these viral DNA sequences are integrated into the host genome using integrase (IN1) (Fig. 2). 3 These integrated viral genomic entities are referred to as proviruses 4 for their ability to be harbored within the host genome and passed onto subsequent generations in a vertical transmission pathway (Fig. 3). Proviruses can be endogenous retroviruses (ERVs), which have been integrated into the host genome in the past and passed through germ-line transmission, or can be exogenous retroviruses, which are recently exposed to the host and thus initiate the cascade of viral entry and integration with the host genome (Fig. 3). 5 Exogenous retroviruses are mostly infecting or pathogenic retroviruses that get processed and integrated into the host genome and ultimately silenced over generations to avoid any further pathogenic manifestations. 6 The flow and selection of these retroviruses within the host genome has been discussed in detail elsewhere.7–9
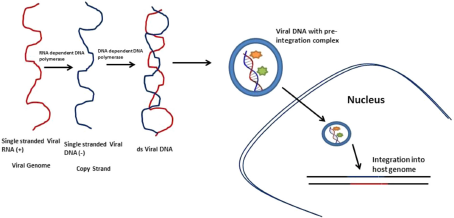
Viral RNA processing and integration into host genome. Retroviruses upon infection convert their single-stranded RNA genome into double-stranded DNA using DNA-dependent DNA polymerases via copying of its own RNA strand using ma-dependent DNA polymerase. In addition, the viral dsDNA along with the preintegration complex is integrated into host genome using integrase (IN1).
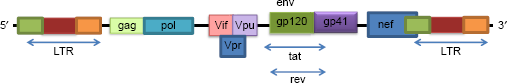
The genomic map of endogenous retroviruses. The map highlights the regulatory regions like 5‘-LTR and 3‘-LTR along with function gene products like envelope proteins, ie, env (gp-120/41), pol (reverse transcriptase), gag (group antigens), and rev (codes for rev protein that exports most of the unspliced and incompletely spliced mrNAs).
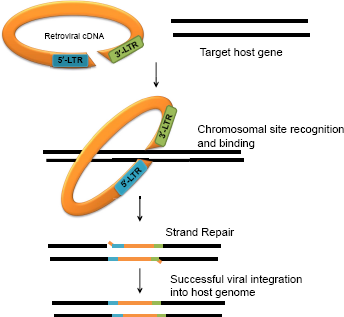
A schematic model of retroviral integration into host genome. The process of integration depends on the selection of specific host target site on chromosome and further downstream integration of viral genome. The functional product expressed from such a recombinant genomic element leads to modified protein product or gives rise to regulatory features like premature stop codons, mutation in coding sequences, or cryptic splice sites.
Thus, certain ERVs can replicate or act in trans to several genes in the chromosome (retrotransposon) 10 and insert germ-line mutations via such events of retrotransposition. 11 There are few other ERVs that are silenced and cannot replicate but add on to the host transcriptomic diversity by acting as alternate promoters and enhancers. 12 Thus, the ability to recombine, alter, and yield the downstream functional products such as proteins and also act as additional transcriptional regulators make these proviruses interesting. 13 With a wide range of functional interference and regulation exerted by these viral sequences, it is of relevance to understand the mechanism of how these viruses have evolved in the host genome for over a hundred millions of years. Mostly, retroviruses are known to infect the somatic cells and are thereby negatively selected and eliminated from the host. 14 The proviruses under study are those that have been vertically transmitted and selected by natural forces of selection to reside and coevolve with the host. However, these proviruses are silenced due to accumulating mutations leading to several premature stop codons, frame-shift mutations, absence of partial functional coding sequences like env or pol (Fig. 2), and sometimes absence of the whole functional coding sequence. 15 Even for the ERVs that are gradually silenced over several generations instead of those being silenced spontaneously, the number of replicative cycle is limited, and thus the chances of retaining pathogenicity get limited. These silenced proviruses have often been referred to as domesticated, 16 tamed, 17 or co-opted viruses. 18
ERVs are classified as class I, class II, and class III ERVs based on the relative geological time scale of integration of these ERVs into the host genome. 19 This makes class III ERVs the most recent ERVs and the ones with relatively minimal mutations or modifications. 7 In contrast, class I and class II ERVs were integrated much earlier than class III ERVs and harbor several silencing mutations. 20 A bias for specific host has been noted among these proviral elements. Most of these proviruses are known to be present in basal vertebrates, although few of them have also been reported in invertebrates, insects, and other lower animals. 21 Approximately 400 different families of pro-viruses are known to be harboring within the human genome. These proviruses constitute about 10% of mouse genome and 58% of human genome. 22 Such a high prevalence of viral genomic fragments substantiates for the fact that these proviruses hold certain beneficial contribution to host fitness and survivability. 23
In embryonic cells and induced pluripotent cells (iPSCs), the ERVs are known to be under transcriptional and post-transcriptional silencing modifications such as regulation of transcription factor binding and epigenetic repressive marks such as methylation, acetylation, and sumoylation of histones as well as the proviral DNA sequences (Fig. 4). 24 These mechanisms of proviral silencing are known to be unique to stem cells. 25 Thus, proviral silencing in embryonic cells and embryonic carcinoma cells has been an important regulation of ERVs and most often studied using mouse model system. 26 The advantage of proviral silencing in stem cells has been used for retroviral and lentiviral vector-based transgene delivery. Such an inherent silencing mechanism of proviruses may facilitate cellular reprogramming, of which very little is known till date. 27 In addition, silenced proviruses have been found to be effective in transgene therapy by overcoming the interference of the viral transgenes. 27 However, owing to a dynamic and stochastic pattern of provirus silencing, there are reports wherein transgenes limit the efficiency of cellular reprogramming and inhibit the differentiation of reprogrammed cells to specific lineages. 28 Furthermore, viral transgenes can interfere with the cellular transcriptome and proteome by modulating the expression of noncoding RNAs, which are known to influence several protein-coding mRNAs. As a result, these wide-scale interferences of viral transgenes with host genetic machinery lead to pathogenic or oncogenic transformations of host cells. 28 Therefore, understanding proviral silencing in stem cells is essential to identify the provirus silencing machinery and shed light on how the absence of this process regulates lineage-specific development and cell fate determination.
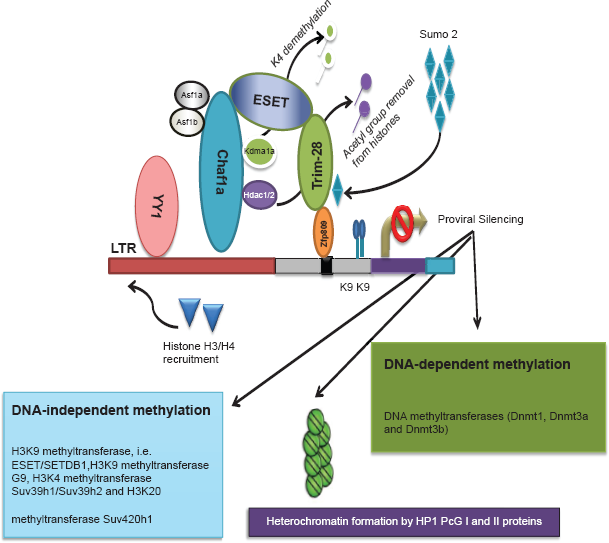
Proviral silencing mechanism in ES cells. The silencing machinery of provirus in embryonic cells involves DNA/histone methylation-, deacetylation-, and sumoylation-based epigenetic modifications in a Trim-28-mediated pathway for Pro-specific PBS and in a YY1-dependent pathway for alternate PBS.
Salient Features of Provirus Expression
The proviral genetic elements within the host genome and their expression pattern are summarized as follows:
Viral genetic entities are integrated within the host genome at gene fragile sites or cell-specific sites of integration. These integrated genetic elements influence the expression of their neighboring genes on the same chromosome. 29
Although these elementary regulatory sequences are retained in proviruses, yet in most cases, there is a loss of functional nucleocaspids or packaging signal, which further ceases viral replicability and horizontal transmission. 30
Domesticated proviruses with lost replicative ability can be reactivated by functional protein product of spontaneous exogenous retrovirus infection and, thus, may exhibit transient replication ability. 31
The viral sequences regulate the expression of several genes through their flanking regulatory regions, ie, long-terminal repeat (LTR) region containing insulators, enhancers, and promoters along with other regions such as primer binding sites (PBS), packaging signals (Ψ), and the polyurine tract signals. 32
Overall, these viral transgenes have served as transposable elements and also as fixed transgenes in the host genetic network. These genetic elements add more diversity and alter the host genetic map leading to an active genetic rewiring of host genome, 32 often termed as “Restless genome.” These transgenes are known to be highly sensitive to external stimulus and thus form an important part in understanding the systemic biology. 32 For example, the activation of ERVs by spontaneous infection or the activation of exogenous viruses possesses another dimension of viral transgene-mediated regulation of host genes and systemic responses. 31 Even viruses with defective genetic entities or those that are silenced can be reactivated by using the polymerases of active virus infection. 32 In addition, viral genetic fragments devoid of replicative ability or replication-deficient ERVs are known to be duplicated by mere errors during host genomic replication, 32 especially human ERVs K (HERVK), also known as the most recently acquired retroviruses with multiple copies of open reading frames, resulting in HERVK-encoded viral protein. However, recent work by Grow et al 33 has identified that the presence of DNA hypo-methylation at LTR regions followed by transactivation by Oct-4 binding in the HERVK genetic inserts leads to reactivation of these transgenes in human embryos and germline cells. These widespread mechanisms of viral integration, replication, or duplication of retroviruses have presented these “proviruses of interest” for their role in evolution, fitness, and adaptability. However, much interest has been focused recently on understanding the unique expression pattern of these proviruses in stem cells in comparison to that of the differentiated cells.
Endogenous Retrovirus Family and Diversity
The conventional classification of the viruses using the International Community of Taxonomy of Virus (ICTV) relies on factors such as the presence of oncogenes, viral core, and accessory gene elements and the range of their host.19,33 Unfortunately, most of the ERVs (largely present as fragments within the host genome) cannot be mapped to their infectious exogenous virus counterparts and, thus, do not find a place in the conventional ICTV classification. These ERVs also vary among themselves depending on their primer binding sites as these are important sites for the start of reverse transcription.19,33 Different viruses use different tRNA primers (tRNAPro/tRNAGlu, etc.) specific to their PBS sequences. 33 Copy number of ERVs also plays a crucial role in the silencing of these ERVs and their expression in differentiated cells. 34 ERVs with high copy number are known to be gradually silenced than those of ERVs with low copy numbers, which are silenced spontaneously. 34 Thus, the expression of different families of ERVs not only varies according to their PBS or other regulatory sequences but also varies according to their classes (ie, class I, class II, or class III) and their relative copy number in the host genome. Interestingly, it is also known that the proviral integration within the host genome is skewed for host preference, and thus, the classification of viruses is also based on the range of host that they can infect. 35 Different classes of ERVs mostly infect vertebrates, and it has been known that such a host preference is not a result of competitive exclusion of the hosts. Thus, the selection of host, copy number, and frequency of proviral expression holds mutualistic benefits in the host-virus relations. Table 1 summarizes different classes of ERVs as per the ICTV system of virus classification.
Classification of ERVs as per the International Community of Taxonomy of Virus (ICTV).
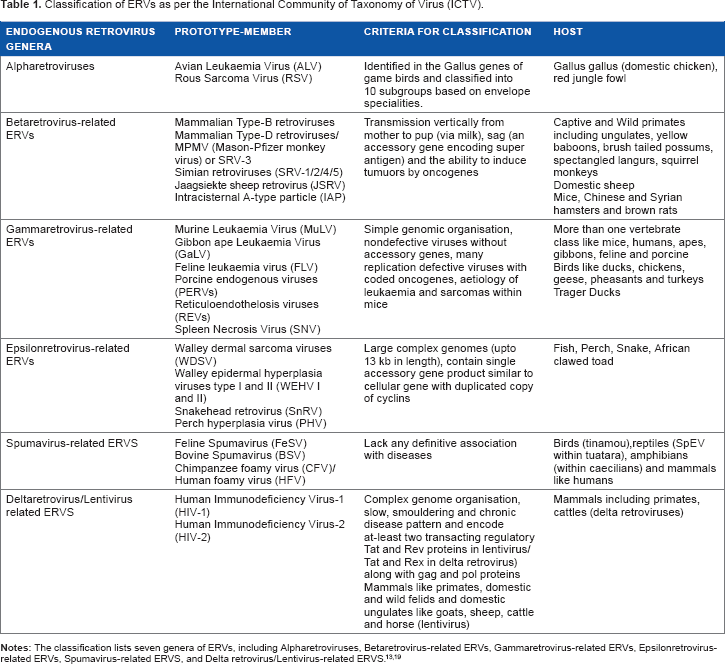
Class I ERVs in humans are known as HERVs and contain the largest families of ERVs in human (around 19 families) with copy number varying from one to several hundreds. 12 However, most of the class I ERVs have been widely defective due to different mutations and silencing mechanism and, thus, are partially or fully intact. Class II HERVs (with minimum 10 independent families) is also referred as HERVK superfamily. 12 In comparison to class I and class III HERVs, this is the least populated class of ERVs. In contrast, class II ERVs are the most populated ERVs in mouse justifying the host-specific bias of integration of proviruses. Class III HERVs are the second most populated human ERVs consisting of four families of ERVs with a copy number of 200-500. 12
Proviral Silencing Machinery in Embryonic Stem Cells
The silencing of ERVs is heralded by the binding of a zinc finger DNA-binding protein, ie, Zfp809, to the proviral DNA sequence at the primer binding site. 25 The binding of Zfp809 is dependent on the recognition of specific PBS sequence, which is tRNAPro in most cases. Subsequently, Zfp809 recruits tripartite motif-containing 28 (Trim-28), also known as KAP-1 or Tifb-1. 25 Trim-28 is the universal regulator of proviral silencing via recruitment of several chromatin remodeling factors to the proviral DNA sequence and epigenetic repressive marks on the proviral DNA or histones (Fig. 4). 36 The most common repressive marks include DNA methylation by DNA methyltransferases (Dnmt1, Dnmt3a, and Dnmt3b) as well as histone methylation by H3K9 methyltransferase, ie, ESET/SETDB1, 24 H3K9 methyltransferase G9, 37 H3K4 methyltransferase Suv39h1/Suv39h2, 38 and H3K20 methyltransferase Suv420h1 (Fig. 4). 15 It has been well understood that the proviral gene silencing occurs via both a DNA methylation-dependent and -independent manner (histone methylation). For most cases of histone methylation, ESET-mediated H3K9me3 is known to be crucial and indispensable for proviral silencing. 24 G9-mediated H3K9me2 is known to be the other arm for H3K9 methylation, which acts in response to be spontaneous and transient activation of retroviruses upon exogenous virus infections. However, G9-mediated methylation is dispensable for prolonged proviral silencing. 37 Apart from methylation, it has been recently shown that events of deacetylation and sumoylation are crucial in silencing the ERVs in embryonic stem (ES) cells. 39 Yang et al 39 through a genome wide screen of mouse ES cells have shown the role of histone chaperons Chaf1a and Chaf1b and sumoylation factors like Sumo2, Ubei 2 and few others in Trim-28-mediated proviral silencing (Fig. 4).
ES cells, which otherwise exhibit a large open chromatin for expression of a wide range of genes required for development, can still harbor local heterochromatized structure spanning the ERV sequences leading to a silencing of these ERVs in undifferentiated or stem cell populations. ERV silencing is also achieved by local heterochromatization of the proviral genetic fragments. 40 This process of heterochromatinformation is guided by HP1 (heterochromatin-associated protein 1) 40 and PcG I and II proteins (polycomb group I and II proteins). 41 These proteins lead to the H3K27me3 enrichment of class I and class II ERVs and KDM1A (lysine-specific demethylase-1A) enrichment of class III ERVs. 42
Although the Trim-28-dependent silencing machinery depends on upstream recognition of specific PBS (tRNAPro). Most of the ERVs are also known to use alternate PBS for the binding of primers for initiation of reverse transcription. However, all such ERVs are also effectively silenced in embryonic cells by Trim-28-independent ERV silencing. 42 Schlesinger et al have shown that another zinc finger DNA-binding protein Ying Yang 1 (YY1) can repress proviral silencing for such ERVs using alternate PBS recognition, and even Rex1 (a YY1 family protein) is known to be important for proviral silencing. YY1 is known to bind to the regulatory region within the LTR of ERVs (Fig. 4). 42 In addition, Schlesinger et al have shown that in ES cells, YY1 can partially interact with Trim-28 and mediate proviral silencing, and such an interaction is lost in differentiated cells. 42
Interestingly, a 3'D4Z4 insulator sequence along with HS4 insulators (in the LTR region) protects the ERVs from silencing and repressive epigenetic marks, leading to a variable and persistent expression of ERVs for a limited number of cell cycles. 43 3'D4Z4 does not target integration sites on specific chromosomal location but confers protection from proviral silencing. 43 Therefore, a critical understanding of the proviral silencing machinery aims to highlight the role of the LTR regions flanking the ERV genetic elements in cell fate determination and regulation of cellular development in comparison to their undifferentiated counterparts.
Pattern of Proviral Silencing
The retroviral silencing machinery in ES cells is composed of different subunits with several factors regulated in a cell-specific manner. The role of PBS in proviral expression has been studied by Taichi et al 44 and Baghi et al 45 using Moloney murine leukemia virus in mouse. 46 It is known that the retroviral silencing is dependent primarily on the recognition of primer binding site (PBS) of the ERVs. 46 Therefore, viruses using similar primer for recognition of PBS are classified together as a group of related viruses and, thus, are expected to be under a similar regulatory mechanism for expression or silencing in different cell types. The proviral silencing machinery in ES cells inhibits the pro-viral gene expression by exerting transcriptional repression through insulator sequences of LTR region, inhibition of transcription factor recruitment and binding to proviral promoters/enhancers, as well as induction of local hetero-chromatization of gene harboring the proviral sequence. 47 In addition to this transcriptional inhibition, several posttranscriptional modifications such as methylation, deacetylation, sumoylation, and characteristic histone marks regulate the pattern of proviral silencing.
The pattern of proviral silencing involves a mixed population of ES cells, which include cells with spontaneously silenced ERVs, cells with moderate expression of ERVs, and cells with high expression of ERVs (rare population also known as escapees) (Fig. 5). 48 For any population of embryonic cells reactivated by exogenous retrovirus infection, the silencing of all ERVs may not be all at the same time but may be gradually silenced over a few generation of cell passages (in vitro), and thus, in between this process, there is always a chance to recover all three types of ES cells with a spectrum of ERV expression, ie, from a silenced or almost negligible ERV expression with moderate-to-high ERV (rare) expressing cells. 48 In addition, the repressive DNA/histone marks seen in ES cells leading to proviral silencing is seen across all the subpopulation of cells such as those with completely silenced ERVs to those with moderate and high ERV expres-sions.48 Therefore, the recruitment of the methyltransferases, acetyl transferases, or other sumoylation factors is spontaneous upon exogenous retrovirus infection, although the silencing of ERVs is attained in a more gradual manner. Thus, when the ERVs were integrated for the first time into the host genome, the silencing of the ERVs was rather gradual than the anticipated spontaneous and global silencing of ERVs. This hints for a possible selection of gradual silencing machinery in response to the beneficial or detrimental effects of the ERV expression.
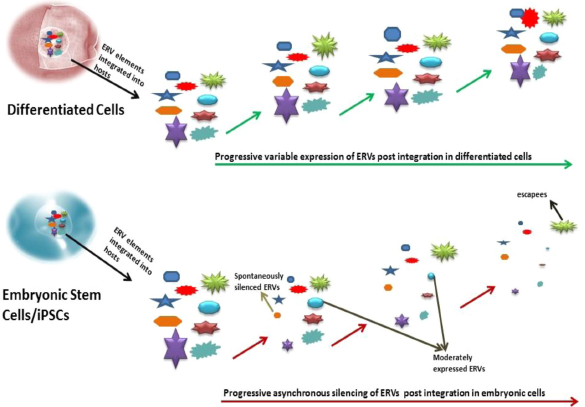
Asynchronous provirus silencing in embryonic cells in comparison to that of differentiated cells. The ERVs are known to be silenced in a local, asynchronous, and gradual process in embryonic cells depending on the site of integration, the class of ERVs, and the copy number of these ERVs. The population of ES cells includes cells with spontaneously silenced ERVs, cells with moderate expression of ERVs, and cells with high expression of ERVs (escapees). In comparison, differentiated cells exhibit a variable ERV expression guided by the nearby genes and copy number.
In addition, it has been appreciated that the silencing pattern of ERVs is greatly influenced by the copy number of the ERVs. 49 Viruses integrated into the host genome with high copy number may also lead to a gradual silencing effect on ERVs, although the histone/DNA epigenetic marks are enriched spontaneously. 49
Thus, the mode of retroviral silencing is more local, stochastic, and asynchronous than that of a global and synchronous retroviral silencing (Fig. 5). 48 The herald of retroviral silencing machinery may not be understood as block and silence but must be viewed as block and progressing toward silence. Therefore, the silencing of ERVs aims at attaining an effective cell type and virus copy number-dependent silencing pattern, which is largely follows a local, stochastic, and nonsynchronous model. Understanding the mechanism of transgene silencing can facilitate targeted gene therapy or virus-based cellular repro-gramming. Interestingly, both the types of provirus silencing dependent and independent on the cell type are known to use a broad strategy of provirus silencing machinery. As discussed earlier, this broad choice of silencing machinery involves using both PBS-binding tRNA primers and PBS-independent silencing machinery.
Provirus in Cell Fate Specification and Determination
Embryonic cells are known to be under wide range of epi-genetic regulatory network facilitating the differentiation of these cells into different cell types. 50 Often such cells are under bivalent histone modifications of both activating nature and repressing nature, leading to selective regulation of transcription. This ensures a stage-specific transcriptional network rewiring and the onset of differentiation from undifferentiated progenitor cells. 50
The relevance of proviruses in regulating the nearby host cellular genes is essential to understand that silenced provi-ruses in stem cells can downregulate the expression of the nearby genes and, reciprocally in differentiated cells, these proviral elements can regulate the host cellular genes sitting close to them in order to specify and determine the cell fate. 51 For example, the use of 5-aza‘-2 ‘-deoxycytidine (blocking DNA methyltransferase) suppressed the methylation of pro-viruses and blocked the expression of the nearby host cellular genes. In addition, the importance of the chromosomal site of integration for proviral silencing has been shown in several other reports.52–54 A change in ERV expression can modulate the epigenetic drift in stem cells and, thus, play a major role in cell lineage specification and fate determination. Thus, it is reasonable to predict the role of proviruses in cell fate determination via host cellular genes neighboring these viral elements.
The ability of the ERVs to act as retrotransposons can regulate the expression of host cellular genes. Retrotransposons have been widely appreciated for their role as controlling elements as they can generate variable expression of host genes arising from accumulating mutations, alternations in coding sequences of host genes, and activation of several proto-oncogenes. 55 For example, retrotransposon-based activity of ERVs has been widely studied for targeting salivary amylases. 56
ERVs can also act as alternate enhancers and promoters for host cellular genes. ERV sequences acting as promoters can induce tissue-specific gene expression as well as code for specific regulatory elements like (long noncoding RNAs). 37 Such an ERV-based regulation of host cellular genes is known to be self-regulatory and acts in trans to other host cellular genes. For example, the “solo” LTR of mouse ERVL is known to act as an alternate promoter of nearby genes. 57 Mouse-sex limited protein (Slp) gene is known to harbor ERVs close to its transcription start site controlling the expression of genes important for placental development and pregnancy. 58 Human ERV-H can regulate cell fate by interacting with Oct-4 and acting as promoter enhancers via LTR7 for nearby genes to maintain pluripotency. 58 In mouse and human genomes, ERV1 is known to carry transcription factor-binding site for pluripotency-associated genes such as Oct-4 and Nanog, exerting regulation of host cell pluripotency. 58 Statistically, about 30% of 5‘-capped human mRNA transcripts and 25% of the coding genes (at their 3‘-UTR) harbor repetitive ERV elements. 58 Such an ERV-mediated regulation of host cellular genes usually occurs in smaller magnitude, although it affects a wide range of host genes. Given that several ERVs are present in high copy numbers, each ERV can act as alternate promoter and enhancer for several host cellular genes.
Several ERVs and ERV-associated gene regulators are known to be crucial for host development and cell fate decision. Approximately 25% of mouse ERVL copies are known to be activated during embryogenesis and is known to affect cellular genes (307 protein-coding genes), leading to the expression of around 625 chimeric transcripts. 58 Such an activated MERVL expression during gastrulation is also associated with epigenetic marks like H3K4 me3 and downregulation of KDM1A, which otherwise demethylates histones. 59 Although most of the ERVs are silenced in embryonic cells, certain ERVs are selectively upregulated for their role in host cell development. LINE-1 (long interspersed nuclear elements) forms the group of rare ERVs, which are upregulated in ES cells in comparison to differentiated cells. 60 In addition it has been shown that ERVs are found in approximately one-third of p53 protein-binding sites and, thus, pose their relevance for cellular development. 61
Gene therapies using retroviral or lentiviral vector system are known to induce host genotoxicity due to insertional mutagenesis. 62 However, the impact and relative scale of genotoxicity is different for undifferentiated and somatic cells. 63 In a study, Zheng et al 64 have compared the retrovirus-mediated insertional mutagenesis in T-cell-committed lymphoid cell lines and in iPSCs. The results indicate that retroviral silencing in iPSCs results in downregulation of major host cellular genes and, therefore, fail to commit to specific lineage, whereas T-cell lymphoid cell lines show upregulated expression of host cellular genes neighboring upregulated ERVs. 64 In another study, Ramos-Mejía et al have shown that the escapees or ERVs that escape retroviral silencing can prevent differentiation of iPSCs into cord blood CD34+ progenitor cells. 65 This indicates that retroviral transgene expression can potentially limit cell differentiation or lineage specification.
Reactivated ERVs can also determine cell fate. Reactivation of ERVs is dependent upon spontaneous exogenous retrovirus infection, recombination with exogenous retroviruses during infection and alteration in their genetic regulatory elements induced by external factors. For example, infection by avian leukemia virus in provirus-harboring cells leads to recombination leading to newer pathogenic variants, modulation of host immune response upon actual infection, as well as destruction of specific target cells upon viral infection. 66 There are reports of neoplastic transformations in certain cells due to insertional activation of oncogenes mediated by ERVs. 67 Interestingly, most of the ERVs implicated for pathogenicity or deleterious functionality upon reactivation belong to the most recent class of retroviruses integrated into the host genome.
Conclusion
Retroviral genome poses a diverse aspect of evolutionary biology in case of complex organisms. These retroviruses are well known for their fast evolving capacity and their coevolution pattern with respect to their vertebrate hosts mainly humans. The presence of ERVs pertains to a more complex biological question than what is being currently investigated through a binary simplification of the case, ie, whether the proviruses are active or silenced. The large time scale of integration of these retroviruses into the host genome and the effective silencing of these ERVs during vertical transmission suggest for an intertwined host-provirus relationship. The mutual benefit of such a co-opted relationship gets substantiated by the fact that these ERVs have a wider role in diversifying the transcriptional network of the host. The fitness of host, especially humans, largely depends on the ability to maintain different cellular populations, including stem cells, undifferentiated progenitor cells, and terminally differentiated cells. This presents a situation of stage-specific and cell origin-dependent mechanism of cell fate determination that requires an undulating transcriptional landscape enriched with a wide variety of epi-genetic modifications (both activating nature and repressing nature). The potency of proviruses to rewire the transcriptional regulatory network through the LTR region to act as alternate promoters and enhancers and regulate the expression of noncoding RNAs can be viewed as pivotal in pluripotency maintenance, lineage specification, and cell fate commitment. However, such a hypothesis does not apply to all the provi-ruses; there is diversity among different ERVs belonging to different classes.
This review aims to understand (a) the evolutionary diversity of retroviruses, (b) identification and characterization of their potential to act as transcriptional regulatory elements, (c) their skewed host preference for vertebrates, and (d) the long time scale of coevolution with the complex organisms. The intricate relationship between the host cellular genes and residing proviruses has been critically analyzed in this study with special focus on their role in host cell fate determination and development. An effective proviral regulation can render improved gene therapy, control random gene expression associated with diseases, and develop a fine tuning of proviral gene expression for using these as markers of cell fate (ie, pluripotency and differentiation).
Author Contributions
Conceived and designed the experiments: SS. Analyzed the data: SS. Wrote the first draft of the manuscript: SS. Made critical revisions and approved final version: SS. Author reviewed and approved of the final manuscript.
