Abstract
A majority of extant life forms require O2 to survive and thrive. Oxidation is inevitably one of the most active cellular processes and one constant challenge that living organisms must face. Generation of oxidants including reactive oxygen species is a natural consequence of cellular metabolism of all biological systems during normal life cycle under different environments. These oxidants oxidize many biological macromolecules such as proteins and affect their functions. Oxidation of specific amino acids in proteins may cause damage to protein structure and impair function, or may also activate protein activities and promote cellular metabolism. As an example, the reversible oxidation of cysteine and methionine residues has a profound impact on protein function and cellular process. A recent study that examines the effect of Met oxidation on Ser phosphorylation in a mitochondrial enzyme, pyruvate dehydrogenase, provides another demonstration that protein oxidation is an important regulatory mechanism for organisms to deal with developmental and environmental challenges throughout life processes.
Over a period of 4,500 million years, our earth has evolved from an O2-free planet into an O2-rich world. A vast majority of life on earth are aerobic, thriving in the O2-rich environment (currently at about 21% of the atmosphere). The abundant presence of O2 also creates an unavoidable oxidative stress to life processes, due to oxidation of biological molecules. Oxidation, by strict definition, describes the process of the loss of electrons or an increase in oxidation number/state by a molecule, atom or ion. In a more biologically relevant sense, it can simply mean the interaction of biological compounds with molecular oxygen. Oxidants such as reactive oxygen species (ROS), examples of which include singlet oxygen ( 1 O2), hydrogen peroxide (H2O2), superoxide anion (O-2) and hydroxyl radical (HO•), are generated by cells during normal respiration and metabolism and in response to external environmental conditions. These reactive oxidants react in the cells with DNA, RNA, proteins, carbohydrates and lipids. 1 The effects of these oxidative reactions vary widely, ranging from no consequence to severe impairment of cellular functions and even death.
Protein oxidation is undoubtedly a frequently occurring process in all aerobic microbes, plants and animals. The oxidative damage to proteins/enzymes has received most attentions because of its implication in diseases and aging/senescence.1–5 However, protein oxidation could also serve as part of regulatory mechanisms along with phosphorylation, glycosylation, acetylation, ubiquitination and others, reflecting a strategy of organisms to deal with constantly changing developmental status and environmental conditions. Oxidation changes the chemical properties of amino acid side chains, leading to modifications of the protein structure and folding. In many cases, these changes result in protein impairment, but in others may instead cause activation and enhancement of protein function and metabolic processes. 6
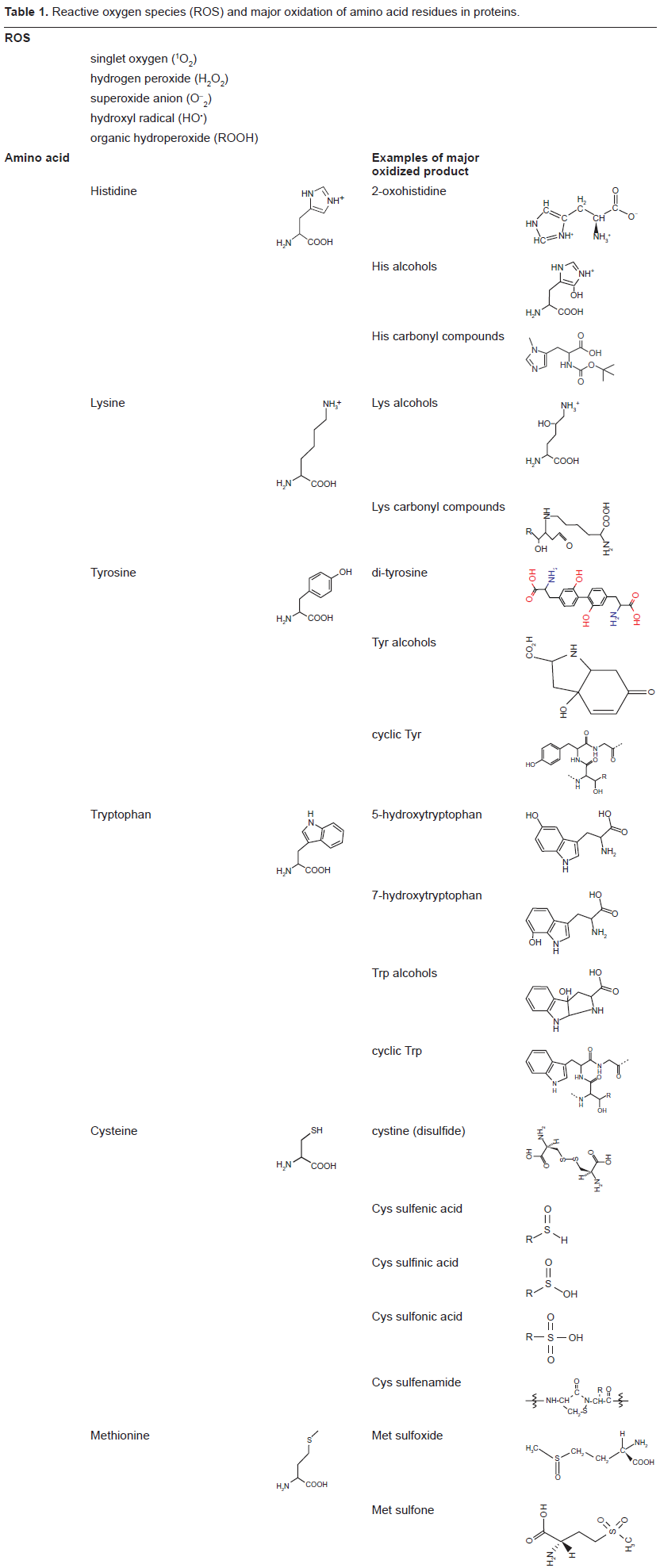
Oxidation of the side chains of some amino acids, notably His, Lys, Tyr and Trp, in proteins often is irreversible. Once oxidized, the affected proteins are permanently damaged and must be replaced. In contrast, the two sulfur-containing residues (Cys and Met) are very susceptible to oxidation but this reaction is reversible in biological systems (Table 1).1,2,7 Since these two amino acids, particularly Cys, often are part of protein conformation and activity, the status of oxidation/reduction of Cys and Met is intimately associated with the function of their residing proteins and other proteins, and may provide an effective mechanism for function regulation.
Reactive oxygen species (ROS) and major oxidation of amino acid residues in proteins.
Functional Regulation by Cys Oxidation
Oxidation of Cys residues in proteins and its role in functional regulation as a redox system have been extensively studied. ROS-induced oxidation of Cys to Cys sulfenic acid triggers a series of interactions with other Cys residues to form disulfide bonds, causing changes in protein activities. A variety of proteins employ Cys oxidation/reduction process, including transcription factors, protein kinases, protein phosphatases and molecular chaperones. For example, the OhrR is a peroxide sensor in bacteria that represses the gene expression of an inducible peroxiredoxin. When a specific Cys residue in OhrR is oxidized to Cys sulfenic acid, the OhrR still is able to bind DNA and suppress transcription. But further reactions with another Cys in Xanthomonas campestris or with other thiols in Bacillus subtilis leading to disulfide formation abrogate the DNA binding activity, consequently allowing peroxiredoxin transcription.8,9 For this protein, degrees of Cys oxidation determine its activity in transcription suppression and consequently the level of peroxiredoxin gene expression.
Formation of Cys sulfenic acid and subsequent disulfide bond following Cys oxidation also inactivate the ability of many protein tyrosine phosphatases to dephosphorylate their target kinase proteins, 10 therefore modulating cellular biochemical and physiological processes. A disulfide bond formed from the oxidized Cys in the catalytic subunit of cAMP-dependent protein kinase results in dephosphorylation of an essential Thr residue and the loss of enzyme activity. Reduction of the oxidized Cys restores both the kinase phosphorylation state and enzyme activity. 11 Likewise, reversible inactivation of MAP kinase phosphatase 3 by Cys oxidation 12 upregulates Jun N-terminal kinases that in turn activate other pathways (e.g. FoxO) in animals. The oxidation/reduction of critical Cys residues probably is the underlying molecular mechanism for ROS-activated signal transductions for development observed in many systems, as demonstrated in Drosophila. 13 Cys oxidation also leads to formation of disulfide bond between two subunits of cGMP–dependent protein kinase, which activates the kinase activity and enhances its affinity for its phosphorylating substrates. 14
Oxidation of a specific Cys residue to Cys sulfinic acid may cause structural changes and even functional conversions. The yeast peroxiredoxins (I and II) function as both peroxidases and molecular chaperones. Oxidation of Cys47 in the presence of the thioredoxin system induces a 10-fold increase in chaperone activity due to dimerization and a greatly reduced peroxidase activity. 15 Therefore, Cys oxidation acts as a functional switch for these proteins. In some cases, the oxidized Cys residues may not be converted to Cys sulfenic acid but still modulate the function. For example, the oxidized Cys cluster in Escherichia coli heat shock protein 33 (Hsp33) causes the release of Zn+2 and dimerization to become effective for preventing aggregation of unfolded proteins, whereas the reduced monomeric Hsp33 lacks the chaperone activity. 16 Again, the status of Cys oxidation plays an important regulatory role in protein functions.
Regulation by Met Oxidation
Like Cys, Met can also be easily oxidized by ROS such as H2O2 to Met sulfoxide [Met(O)]. Met(O) can be reduced back to Met by a group of specific enzymes called Met sulfoxide reductases (Msr) (see review by;17, 18 also7, 19, 20). Accumulating studies have demonstrated that the Msr system serves as an important protein repair and antioxidation mechanism in cells.17,21 Just as important, Met oxidation may have regulatory roles in cellular signal transduction, growth and developmental processes. For example, oxidation of Met to Met(O) in α1-antitrypsin results in a loss of proteinase inhibiting activity. 22 Reducing Met(O) to Met by human Msr can restore the activity of the oxidized α1-antitrypsin in vitro. 23 Since the cellular balance of proteinase/anti-proteinase is linked to the development of a disease condition called emphysema, regulation of proteinase inhibitor activity through Met oxidation/reduction is physiologically important. Likewise, oxidation of Met residues in the bacterial glutamine synthetase leads to a decrease in Mg+2-dependent, but not Mn+2-dependent, γ-glutamyl transferase activity, reflecting the physiological covalent regulation of this enzyme. 21 The stoichiometric status of the reversible oxidation of Met residues, i.e. the ratio of Met:Met(O) in this protein, provides a mechanism of fine-tuned regulation of enzyme activity, analogous to the regulatory mechanism of reversible phosphorylation/dephosphorylation.
The notion of Met oxidation as a functional regulator of proteins was explicitly proposed by Ciorba et al 24 in the study of a potassium channel. These researchers demonstrate that oxidation of a specific Met to Met(O) causes the potassium channel to recover rapidly from its inactivated state, whereas Msr-catalyzed reduction of the Met(O) to Met facilitates the maintenance of the inactivation. 24 Thus, Met oxidation and reduction plays an important regulatory role in the signal transduction. Recent studies show that in humans, oxidation of a single Met residue in apolipoprotein A-I impairs an enzyme activation critical for reverse cholesterol transport, 25 while Met(O) in calcineurin hinders the proper nuclear-localization of the nuclear factor of activated T cells and transcriptional activation in T cells, subsequently affecting cellular immune responses. 26 In chloroplasts of plants, the conserved Met residues of heat shock proteins must be in a reduced form in order to bind and protect other proteins from aggregation. 27 Furthermore, addition of an O atom to Met in many cases changes protein conformation, disrupts the protein native state, increases surface hydrophobicity to expose more hydrophobic residues and renders more susceptibility to proteolytic degradation by the proteasome, as first reported by Caldwell et al 28 for E. coli ribosomal protein L12 and subsequently demonstrated in many studies such as for staphylococcal nuclease, 29 calmodulin, 30 porcine lutropin 31 and thrombomodulin. 32 Therefore, cellular oxidation (and associated reduction) of protein residues affects the quantity, composition and activity of overall protein populations, thus regulates the biochemical processes.
As an effective regulatory mechanism, protein oxidation can also activate or enhance biological activity. For example, oxidation of Met-1 residue of ubiquitin to Met(O) gives rise to a 50% increase in reactivity, but further oxidation of Met(O) to sulfone decreases the activity. 33 Met oxidation activates calcium/calmodulin-dependent protein kinase II, leading to myocardial apoptosis, and the enzyme Msr can reverse its oxidation. 34 Similarly, Met oxidation activates the native human complement C5 protein so that it becomes able to bind other proteins. 35 Thus, the degree of protein oxidation offers the cell a means to regulate the abundance and activity of target proteins/enzymes, consequently the rate and direction of biochemical and physiological processes.
A recent study of the plant mitochondrial pyruvate dehydrogenase complex (PDC) by Miernyk et al 36 reveals a new mode of regulation by specific Met oxidation affecting protein phosphorylation. The mitochondrial PDC catalyzes the conversion of pyruvate to acetyl-CoA, leading to the Krebs cycle with concomitant energy generation or to lipid biosynthesis. PDC consists of three distinct components: pyruvate dehydrogenase (E1), dihydrolipoyl acetyltransferase (E2), and dihydrolipoyl dehydrogenase (E3). The PDC is regulated at several levels, one of which is reversible phosphorylation of E1. 37 Phosphorylation of the E1α subunit by pyruvate dehydrogenase kinase (PDK) switches off the PDC activity, whereas dephosphorylation by phospho-pyruvate dehydrogenase phosphatase (PDP) reactivates PDC. Plant mitochondrial E1α contains two Ser phosphorylation sites: Ser300 (site 1) and Ser306 (site 2). A conserved Met reside (Met301) is located next to Ser300. Miernyk et al 36 find that when this Met is oxidized to Met(O), the synthetic peptide containing site 1 Ser is no longer phosphorylated by PDK. Met oxidation can be detected in E1α isolated from intact pea mitochondria treated for H2O2 production. When the tobacco suspension cells are incubated with several oxidants known to induce ROS, the relative abundance of the phosphorylated PDC in these cells decreases in all treatments except for H2O2. These authors suggest that changes in physical properties due to Met oxidation and proximity of Met(O) to site 1 Ser cause the inhibition of E1α phosphorylation. To my knowledge, this is the first demonstration of Met oxidation affecting protein phosphorylation. Many phosphoserine-containing proteins such as 14-3-3 proteins possess immediately adjacent Met residues. Oxidation of these critically positioned Met residues could conceivably disturb or disrupt the binding ability and specificity of these proteins that mediate signal transduction. 38 Therefore, if the observations by Miernyk et al 36 prove to be a common feature in different systems, it will further support the notion that Met residue oxidation has an active regulatory role in protein/enzyme function.
Meanwhile, several questions can be raised. First, it would be interesting to see the enzymatic activities of PDC, particularly of E1, examined. One would expect to observe an increase in PDC activity in oxidant-treated cells when the E1α phosphorylation is inhibited by Met oxidation. Second, many environmental conditions such as high light intensity, low and high temperatures, long night and pathogen infections induce both ROS production and Met oxidation in plants.39–42 The notion that oxidative stress-induced ROS enhance PDC and respiration activities by inhibiting Ser phosphorylation by PDK (perceivably many other kinases as well) needs to reconcile with some of the known experimental observations. For example, plant mitochondrial PDC is normally off in the light 37 when ROS production should be actually more active. Finally, in contrast to chloroplasts and animal mitochondria where several Msr enzymes that convert Met(O) back to Met have been identified,7,20,43 plant mitochondria do not appear to have such an enzyme. Therefore, how plant mitochondria deal with Met oxidation and the associated effects on protein function needs future investigations. Perhaps, chloroplasts and mitochondria in plants handle Met oxidation differently, thus the occurrence and effect of Met(O) on protein modification and function may also differ in these organelles.
Responses to Oxidative Stress in Different Systems
As discussed above, the biochemistry of protein oxidation is likely to be similar, if not the same, among different organisms. However, the origin of and responses to oxidative stress must be different for microbes, plants and animals due to their distinct life cycle histories. I am not aware of the published comparative studies of the differences in protein oxidation between prokaryotes and eukaryotes. It is known that for bacteria nutritional conditions play an important role in protein synthesis, oxidation and degradation. 44 Other environmental factors such as pH, temperature and salt concentration influence the oxidation status in archaea. 7 Among eukaryotes, plants and animals display different responses to oxidative stress. For example, plants exposed to high light intensity, extreme temperatures or pathogen infections exhibit enhanced ROS production and elevated protein oxidation.40–42 In contrast, ROS biogenesis in animals (particularly mammals) seems to be less influenced by sudden environmental changes but is more closely associated with the endogenous developmental processes. Indeed, feeding fruit flies with ROS-inducing reagents like paraquat 45 or treating animal cell cultures with oxidant chemicals46–48 leads to observations of oxidative stress. But these kinds of lab-generated conditions probably do not truly reflect the physiology of these animals. Therefore, more comparative studies are needed to examine what and how environmental factors cause oxidative stress (ROS overproduction and macromolecule oxidation) in different systems.
It should be noted that protein oxidation by ROS is a non-enzymatic chemical process, therefore is non-specific. This is significantly different from other processes of protein modification and regulation such as phosphorylation, glycosylation, acetylation and ubiquitination where specific enzymes are dedicated to the process as part of gene expression regulation. The extent of protein oxidation probably depends on many factors such as ROS and target protein levels, abundance of vulnerable amino acid residues in the protein sequence and subcellular location of the protein relative to the source of ROS. For example, Met-free proteins (accounting for ca. 3.5% of all nuclear-encoded proteins in Arabidopsis thaliana; our unpublished analysis) would be expected to have less occurrence of oxidation, while proteins containing multiple Cys or Met residues are more likely to be the ROS targets. However, susceptibility of proteins to oxidation will ultimately be determined by the conformational positions of the side chains of these amino acids in the protein molecules. Those residues buried inside the protein will probably be more resistant to ROS attacks. One example is glutamine synthetase: 8 out of its 16 Met residues can be oxidized by H2O2, resulting in a loss of enzyme activity. All the 8 oxidizable Met are surface exposed whereas all the 8 unoxidized Met are buried (21). Thus, the structural positions of residues are a critical factor determining whether and how extensively a protein will be subjected to ROS oxidation and how the protein function will be affected. Regardless of the non-specific nature of oxidation, the biogenesis of ROS can be viewed as a biochemical and physiological response in cells to developmental and environmental conditions, therefore an important regulation signal for protein function. Now the advances in proteomics should allow a global view of protein oxidations (oxidative proteome) under different physiological conditions, analogous to phosphoproteomics 49 or glycomics. 50
In summary, oxidation of protein residues, particularly the reversible oxidation of Cys and Met, is a constantly occurring event during normal cellular activity and in response to stress. Through the oxidation and reduction of these residues, protein function may be up- or down-regulated or even altered due to protein structure changes and interactions with other residues, proteins or other cellular components. Therefore, protein oxidation, along with other protein modification processes, acts as an important regulatory mechanism that organisms utilize to modulate physiological and biochemical processes to survive and thrive in the richly oxic environment on earth.
Disclosure
The author declares no conflicts of interest.
Footnotes
Acknowledgements
The author thanks Herbert Weissbach for discussion and comments on the manuscript. This article is not aimed to provide a comprehensive review. Therefore, the author regrets omitting a large body of published research on this topic.
