Abstract
This short note highlights a procedure to distinguish the residues having similar masses, oxidized methionine and phenylalanine containing peptides using MALDI TOF/TOF. The isotope intensities give a preliminary recognition of peptides containing oxidized methionine. In the peptides with partial oxidation of methionine a mass difference of 16 Da can be observed in the mass finger print of the peptide. Neutral loss of methane sulphenate (CH3 SOH) in the MS/MS spectra is the most abundant ion in the peptide containing oxidized methionine, whereas this fragment ion is not produced from phenylalanine containing peptide. The mass spectra of methionine, oxidized methionine and phenylalanine containing peptides were examined from the proteins of
Introduction
In Proteomics, proteins are identified using mass spectra of the trypsin-digested peptides called peptide mass figure print (PMF), and subsequently obtaining the sequence of these peptides using tandem mass spectra. Identification of peptide sequences by mass spectra involves the use of several algorithms such as MASCOT and SEQUEST. These programs also give an estimate of the false positive rates of identifying the peptide sequences. Proper identification of peptide sequence is essential for the identification of proteins, particularly when the genome sequence of the species under study is not available.
The origin and control of some of the oxidation reactions of methionine, cysteine and tryptophan residues in proteins are known. 11 Partial oxidation of methionine is a common phenomenon during some of the processes used in proteomics. The protein spots on the SDS-gel most often contain molecular species having oxidized methionine and methionine. Both of these molecular species appear at the same position and therefore can't be distinguished. The tryptic peptides containing methionine and the peptides containing oxidized methionine show two different signals in the PMF with a mass difference of 16 Da. Identification of oxidized methionine using FAB—MS in peptides was reported earlier. 12 However, the oxidized methionine and phenylalanine have similar mass of 166.0532 and 166.0862 respectively. Hence it is difficult to distinguish the peptides containing these residues. Therefore it is necessary to develop a methodology to identify oxidized methionine containing peptides. Mass spectrometry is extensively used to identify the various oxidation reactions in proteins and peptides. 13 Several factors of mass spectra are useful. The distribution of the isotope peaks of the parent ion serves as signatures of sulfur-containing amino acids. The y-series and b-series of ions can be used for determination of the sequence of peptides. In addition to these, neutral loss of methane sulfenate in these spectra confirms oxidation of methionine (Fig. 1).
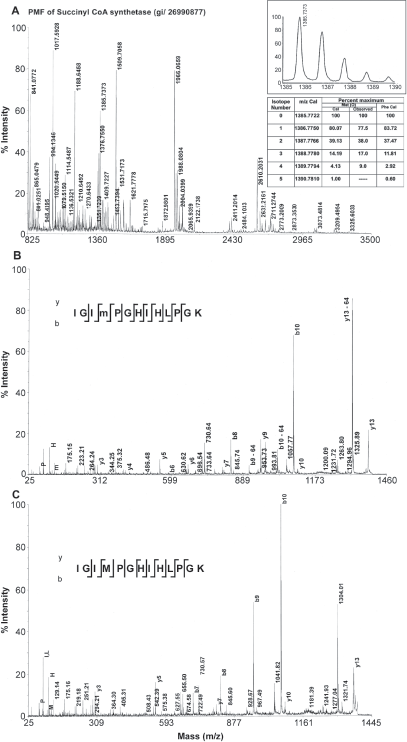
PMF of a protein from
Our laboratory is involved in identification of proteins from cold-tolerant bacterium
Materials and Methods
The isotope abundances for various peptides can be calculated using the web site http://prospector.ucsf.edu. MS-Isotope utility program was used from this web site to calculate the isotope abundances of peptides. The experimental intensities of the isotope peaks and mass values can be obtained from the spectra. The genome sequence of
Results
Phenylalanine and oxidized methionine have same molecular weight, but they differ in the isotope distribution. The natural abundance of sulfur isotopes is 32S-95.02%, 33S-0.75%, 34S-4.21%, 36S-0.02%. 16 The isotope distribution of methionine containing peptides in general showed a higher intensity of isotopic peaks due to increased natural abundance of 34S from second isotope peak (Fig. 1A inset). This is suggestive of a preliminary indication to distinguish between oxidized methionine containing peptide and phenylalanine containing peptide. A decrease in relative intensity of the first isotope peak and an increase in the relative intensity of third isotope of a tryptic peptide obtained from succinyl CoA synthetase alpha sub unit were provided as an example (Fig. 1A). The mass and the intensities of different isotopes were calculated by replacing oxidized methionine with phenylalanine. The oxidized methionine residues could be differentiated from phenylalanine residues by the higher intensity of the second isotope (34S-4.21%) associated with the former. Table 1 shows some tryptic peptides containing methionine, oxidized methionine and phenylalanine peptides from a few identified proteins using database search. The peptide mass finger print of a protein succinyl CoA synthetase that had produced peptides with partial oxidation of methionine was shown in Figure 1. Both oxidized methionine-containing peptide and methionine-containing peptide appeared in the mass spectrum of the tryptic peptide with a mass difference of 16 Da (Fig. 1 A, Table 1). Loss of 64 Da from precursor ion has been found to be the most abundant ion in the MS/MS spectrum (Fig. 1B). The peaks help to distinguish oxidized methionine form phenylalanine are shown in the Table 1. Manual verification of the MS/MS spectra clearly identified the oxidized methionine ruling out the possibility of the residue being a phenylalanine.
Analysis of some tryptic peptides form the proteins of
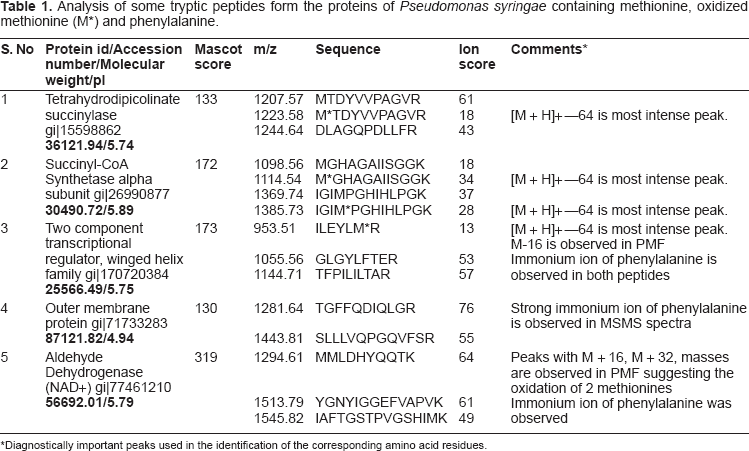
Diagnostically important peaks used in the identification of the corresponding amino acid residues.
Discussion
Tryptic digests of several peptides obtained from the proteins of an Antarctic bacterium
In conclusion, using methionine oxidation as a chemical marker, the sequence of methionine containing peptides has been determined. This has helped in the identification of peptide sequences containing these residues from an Antarctic bacterium. Manual verifications the MS/MS spectra are essential for correct identification.
Disclosure
The author reports no conflicts of interest.
Footnotes
Acknowledgements
I acknowledge the financial support from the department of Biotechnology, New Delhi through a research grant (BT/PR7383/BRB/10/474/2006). I sincerely thank Dr. M. K. Chattopadhyay for his help in the preparation of this short note.
