Abstract
Over the years, the term “rhizobia” has come to be used for all the bacteria that are capable of nodulation and nitrogen fixation in association with legumes but the taxonomy of rhizobia has changed considerably over the last 30 year. Recently, several non-rhizobial species belonging to alpha and beta subgroup of Proteobacteria have been identified as nitrogen-fixing legume symbionts. Here we provide an overview of the history of the rhizobia and the widespread phylogenetic diversity of nitrogen-fixing legume symbionts.
Introduction
Members of the Leguminosae, comprising about 19,000 species form the largest plant family on earth and play an important ecological role. Leguminous plants are very diverse in morphology, habitat, and ecology, ranging from Arctic annuals to tropical trees and their importance as food crops worldwide is large. Some 25% of the world's major crop production is derived from legumes, and more than one-third of humanity's nutritional nitrogen requirement comes from legumes. Besides, legumes are very important both ecologically and agriculturally because they are responsible for a substantial part of the global flux of nitrogen from atmospheric N2 to fixed forms such as ammonia, nitrate, and organic nitrogen.
They probably evolved approximately 60 million years ago (Ma), early in the Tertiary period and their evolutionary success can be largely attributed to their ability to form symbiosis with specific bacteria known as rhizobia. The symbiotic relationship between nitrogen-fixing rhizobia and legume plants has been studied for over 100 years as a classic example of mutualistic associations. Rhizobia or sometimes Rhizobium is the general name given to a phylogenetically diverse group of soil bacteria that form nitrogenfixing symbioses with leguminous plants. These bacteria are environmental heterotrophs with great metabolic plasticity that may survive in many different environments, sometimes unassociated with plant hosts but they have significant interest because of their ability to form symbiotic associations with legumes and are able to elicit symbiotic association on the species of Leguminosae, forming nodules in roots or stems (Fig. 1).

Typical effective nodules formed by Rhizobium in Phaseolus.
Symbiosis between rhizobia and legumes is the result of millions of years of co-evolution being a process complex and exquisitely regulated that plays an important role because it offers the ability to convert atmospheric molecular nitrogen into forms useable by the plant, a process called biological nitrogen fixation. Whatever the true figure, this symbiosis represents the most important nitrogen input to many ecosystems and is among the most important ecological interactions to humans and ecosystems. Currently, the subject of biological nitrogen fixation is of great practical importance because the use of nitrogenous fertilizers has resulted in unacceptable levels of water pollution increasing concentrations of toxic nitrates in drinking water supplies and the eutrophication of lakes and rivers. Over the years, the term “rhizobia” has come to be used for all the bacteria that are capable of nodulation and nitrogen fixation in association with legumes but the rhizobial species described so far are very diverse and do not form an evolutionary homologous clade. In this article, we expose the widespread phylogenetic diversity of nitrogen-fixing legume symbionts and their taxonomic history.
From the Rhizobia Discovery until Bergey's Manual from 1974
The microorganisms able to establish nitrogen fixing symbiosis with legumes were discovered in XIX century, being Frank who firstly named them Rhizobium leguminosarum. 1 Since this date all bacteria able to nodulate legumes are called rhizobia and therefore they were involved in the history of the Bacteriology since the beginning of this science. Frank was contemporary of Ferdinand Cohn who published a bacterial code system for which is considered the father of the bacterial taxonomy, based at that time on the shape and general appearance of bacteria. 2 After, Orla-Jensen proposed the use of physiological characteristics as basic criterion in bacterial taxonomy. 3 In addition to these, other criteria that include microbe interactions with higher organisms were used in some groups such as rhizobia although their taxonomic value was considered of doubtful since the beginning of Bacteriology. 4
In the beginning of XX century, an American microbiologist, Bergey, initiated the publication named Bergey's Manual 5 in which the bacteria were classified on basis of their phenotypic characteristics. In the case of rhizobia their classification as Gram negative, aerobic, non-sporulated and motile rods do not allow their differentiation from other soil bacteria with the same phenotypic characteristics. Therefore their ability to induce nodules in legumes was from this date the basic criteria for differentiation of rhizobia and their taxonomy reached lower development than that of other soil related bacteria whose taxonomy was much more developed just due to their inability to form plant nodules.
The Bergey's Manual played a fundamental role in rhizobial taxonomy since it recorded the history of the species of rhizobia from the beginning of the Bacteriology until 1980 in which they were officially validated. 6 In the Bergey's Manual of Determinative Bacteriology edited in 1974 six species were included in the genus Rhizobium. 7 Only few phenotypic characteristics of the species of this genus were included and the authors of this chapter indicated that the data about plant infection are essential for species classification. 7 These characteristics were also essential to differentiate Rhizobium from genus Agrobacterium the second genus included in the family Rhizobiaceae. Whereas Rhizobium contained species nodulating legumes, Agrobacterium included species able to induce different pathogenic symptoms in plants.
The species of genus Rhizobium were divided into two groups containing the fast and slow growing species on YMA medium, respectively. 8 The fast growing species were R. leguminosarum nodulating Pisum, R. trifolii nodulating Trifolium, R. phaseoli nodulating Phaseolus and R. meliloti nodulating Medicago (Fig. 2). The slow growing bacteria included R. lupini and R. japonicum nodulating Lupinus and Glycine, respectively. In contrast with the genus Rhizobium, the species of the genus Agrobacterium were differentiated on the basis of several phenotypic characteristics since two species of this genus, A. tumefaciens (Fig. 3) and A. rubi produced the same symptoms in plants, tumours. From the other two species of this genus A. rhizogenes produces hairy roots and A. radiobacter is not pathogenic for plants.
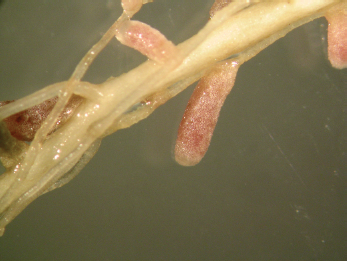
Typical effective nodules formed by Ensifer meliloti (formerly Sinorhizobium meliloti, formely Rhizobium meliloti) in Medicago sativa.

Tumours formed by A. tumefaciens in Nicotiana tabaccum.
The only molecular datum included in the 1974 edition of Bergey's Manual was the G + C content of the bacterial species although in the 60's decade techniques as DNA-DNA hybridization for species differentiation was reported in several works after the discovery of DNA renaturation by Marmur.9–11
Taxonomy of Rhizobia between 1974 and 2000
In 70's decade it was discovered that the symbiosis and pathogenicity genes are harboured in plasmids, in many cases autoconjugative, in both Agrobacterium 12 and Rhizobium. 13 The symbiotic genes included those involved in nodulation (nod) and nitrogen fixation (nif) and the pathogenicity genes those involved in tumour or hairy roots (vir). The nod genes are responsible for the synthesis of the nod factors (lipochitin-oligosaccharides) that are receptors for the plant favonoid signal.14–18 The nodD is a regulatory gene of the operon nodABC19–21 whose genes are determinants of the host range.22–27 The nifH gene codifies a subunit of the enzyme nitrogenase involved in nitrogen fixation and is carried by rhizobia but also by free-living nitrogen fixers.28–30
Although, the genes that are easily transferred among species are not useful in taxonomy, when the first edition of Bergey's Manual of Systematic Bacteriology was published in 1984, the symbiotic and pathogenic criteria not only were not eliminated from the rhizobial taxonomy but used to reclassify some species. 31 This was the case of the species R. phaseoli and R. trifolii that were reclassified into R. leguminosarum and from this moment were considered by rhizobiologists as biovars from R. leguminosarum. Then, the genus Rhizobium contained only three species, the two old species R. leguminosarum and R. meliloti, and a new species named R. loti proposed by Jarvis et al 32 to include rhizobia nodulating legumes other than those nodulated by R. leguminosarum and R. meliloti. 31
It is necessary to take into account that in 1980 was published the valid list of bacterial species by Skerman et al 6 that included R. leguminosarum, R. phaseoli, R. trifolii and R. meliloti. From this date all species must be described or validated in the International Journal of Systematic Bacteriology (IJSB) being not valid the proposal performed outside this Journal. As the proposal of Jordan in the Bergey's Manual was not validated in IJSB, the species R. phaseoli and R. trifolii were maintained in the list of the valid species. The species Rhizobium lupini disappeared from the Manual but as it was not reclassified in any species and neither was officially rejected it currently remains as valid species.
In the Bergey's Manual from 1984 two new genera were added to family Rhizobiaceae, the genus Bradyrhizobium (Fig. 4) with a single species, B. japonicum, that was originated by the segregation of the slow growing species from genus Rhizobium 33 and the genus Phyllobacterium isolated from plant leaves nodules with two species P. myrsinacearum and P. rubiacearum. 34 Whereas the species B. japonicum was described in the IJSB, those of genus Phyllobacterium did not, but were validated in the same year of the Bergey's publication. 35

Typical effective nodules formed by Bradyrhizobium in Lupinus albus.
In 1988 a new genus named Azorhizobium able to nodulate stems of Sesbania rostrata in Africa was added to family Rhizobiaceae 36 and the species R. fredii, a fast growing species nodulating soyabean, officially described in 1984 37 was reclassified into a new genus named Sinorhizobium on basis of phenotypic characteristics 38 and 16S rRNA gene sequences. 39
From 1984 ahead deep changes in the taxonomy of prokaryotes had been occurred from the Woese's proposal to classify them on basis of their 16S rRNA gene sequences 40 (Fig. 9). The analysis of this gene allowed the phylogenetic classification of bacteria with independence of their phenotypic or symbiotic characteristics. Nevertheless, during many years the symbiotic characteristics have a dominant role in the rhizobial taxonomy and the 16S rRNA gene was not included in the minimal standards for rhizobial species description until 1991. 41 According to them the cultural and morphological characteristics, the 16S rRNA gene analysis, the hybridization of DNA-DNA or rRNA-DNA, the RFLP and MLEE analysis and the symbiotic characteristics are recommended to describe new species of rhizobia.
From 1984 until 1994, year in which was published the ninth edition of the Bergey's Manual of Determinative Bacteriology, only Bradyrhizobium elkanii, a new slow growing species nodulating soyabean, was described outside IJSB 42 although was later validated in this journal. 43 Since this date a clear dichotomy was produced between Bergey's Manual and IJSB because the species published in IJSB were not completely recorded in the Manual. In this way in the edition of 1994 were included S. xinjiangensis, 38 whose name was later corrected as S. xinjiangense, a species nodulating soyabean, and R. galegae, a species nodulating Galega. 44 However, Agrobacterium vitis inducing tumours in Vitis vinifera 45 and two species of genus Rhizobium, R. huakuii and R. tropici, nodulating Astragalus and Phaseolus, respectively were not recorded.46,47
Recent Changes in the Taxonomy of Rhizobia
From Woese's proposal, in the 90's decade and mainly from year 2000 significant changes were produced in the taxonomy of all bacteria since the sequencing of 16S rRNA gene was mandatory for bacterial species description. From this moment, although the nodulation of legumes was not excluded from rhizobial taxonomy, it became a secondary criterion in species description. This was a revolutionary change in rhizobia because the identification of these bacteria was possible with independence of the isolation source.
In the second edition of the Bergey's Manual of Systematic Bacteriology published in 2005 the bacteria were arranged in different phylogenetic groups obtained after the analysis of 16S rRNA gene. The rhizobia were included in the class alpha Proteobacteria and distributed in several families within the new order Rhizobiales belonging to alpha-Proteobacteria. 48 This new order was not validated in IJSEM because it is not rightful 49 since includes the genus Hyphomicrobium and this genus was previously included in a valid order named Hyphomicrobiales. 6 In this way the Manual recovered the different taxonomic changes proposed in the taxonomy of this group of bacteria since the publication of the previous Manual of Systematic Bacteriology from year 1984. In this Manual the former family Rhizobiaceae includes the former genera Rhizobium, Sinorhizobium, Agrobacterium and the new genus Allorhizobium isolated from Neptunia nodules in Senegal, 50 together with a genus that does not was isolated from legumes nodules named Ensifer. The genus Azorhizobium was included in the family Hyphomicrobiaceae and the genus Bradyrhizobium in the family Bradyrhizobiaceae. This family is not valid because includes the genus Nitrobacter included in a previously validated family named Nitrobacteraceae. 6 A new genus proposed in 1997 to include the species of intermediate growth named Mesorhizobium 51 (Fig. 5) was included in the family Phyllobacteriaceae that also included the former genus Phyllobacterium.

Typical effective nodules formed by Mesorhizobium in Cicer arietinum.
In spite of the efforts of Bergey's Manual for the adaptation of its contents to the current taxonomy of rhizobia, we can affirm that the dichotomy between Manual of Bergey's and IJSEM is until more marked in the 2005 edition than in previous ones. In spite of the increasing of the number of rhizobial species recorded in the Bergey's Manual some species described several years before its publication are missing such as R. yanglingense, 52 R. sullae 53 and R. indigoferae 54 within genus Rhizobium. In the case of genus Sinorhizobium, S. arboris and S. kostiense, 55 S. kummerowiae 54 and S. morelense 56 were not included. In the genus Agrobacterium it lacks A. larrymoreii. 57 Finally, in the case of genus Bradyrhizobium, B. yuanmingense 58 was not included. It is also surprisingly that in the Manual was not recorded the proposal to include the genera Agrobacterium and Allorhizobium into genus Rhizobium since the author responsible of the complete chapter from order Rhizobiales was one of the coauthors of the reclassification of Agrobacterium and Allorhizobium into genus Rhizobium.
Although the current classification of bacteria is based in the 16S rRNA gene that has allowed the description of non-nodulating species of rhizobia, it has limitations to differentiate among close species59,60 and for this purpose several metabolic genes (housekeeping) have been proposed in several groups of bacteria. 61 In rhizobia the two first genes analyzed were recA and atpD 62 and currently they have been sequenced in many rhizobial species showing their usefulness in differentiation of species whose 16S rRNA genes are nearly identical.59,60 In the last years new schemes for identification and phylogenetic analysis of bacteria named MLSA (Multilocus sequence analysis) and MLST (Multilocus sequence typing) based in the analysis of several housekeeping genes have been applied phylogenetic analyses of concrete groups of rhizobia as Ensifer63–65 and Bradyrhizobium.66,67 For instance, these studies indicate that housekeeping genes sequencing is superior to DNA–DNA hybridization for the assessment of genetic relatedness between Ensifer species and support the suggestion that Ensifer xinjiangensis is a later heterotypic synonym of Ensifer fredii. 64 The ad hoc committee for re-evaluation of the species definition suggested that ‘species should be identifiable by readily available methods (phenotypic, genomic)’ and that one promising approach towards this goal is the determination of a minimum of housekeeping genes 68 and Zeigler 69 suggested that analysis of less than five suitable housekeeping genes might be sufficient for a reliable classification. For this reason, in the last descriptions of new rhizobial species the analysis of at least two housekeeping genes have been included commonly to know the closest related species before to perform DNA-DNA hybridization experiments.
Besides the housekeeping genes also named “core” genes, some “auxiliary” or “accessory” genes involved in the legume symbiosis are commonly included in species description of rhizobia and in some MLST analysis.70,71 The symbiotic genes from rhizobia are codified in plasmids in fast and some middle growing species and in symbiotic islands in the middle and slow growing ones.72–85 From these genes, the most studied in rhizobia are nodD, nodA, nodC and nifH.25,70,86–90 Nevertheless, the symbiotic genes are not useful in taxonomy because due to their ability to be transferred in nature91,92 from plasmids to islands, 93 from bacteria to plants 95 and among bacteria. 96 Therefore the analysis of symbiotic genes is overall useful to identify non-rhizobial species able to nodulate legumes (Fig. 10 and Fig. 11) and to carry out biogeography studies of legume endosymbionts. 97 Particularly the nodulation genes are useful to define biovars within rhizobial species.26,98–100
Currently there are species from classic rhizobial genera in which the symbiotic genes nodD and nifH were not found as occurs in Bradyrhizobium betae isolated from roots of Beta vulgaris (Fig. 6) affected by tumour-like deformations. 101 Also several species have been not isolated from nodules although they were able to nodulate Medicago sativa as Rhizobium daejeonense isolated from a cyanide-treatment biorreactor 102 and R. cellulosilyticum isolated from a pulverized decaying wood of Populus alba. 103 Other species have been isolated from plant rhizosphere such as and R. alamii 104 and plant root inner tissues as R. oryzae. 105

Tumour-like formations in Beta vulgaris from which was isolated Bradyrhizobium betae.
The complete list of valid species of rhizobia is recorded in the List of Prokaryotic Names with Standing in Nomenclature that is performed and revised by Dr. Euzeby (http://www.bacterio.cict.fr). According to this list currently the genus Rhizobium contains 33 species and includes to the former genera Agrobacterium and Allorhizobium. The genus Sinorhizobium, currently named Ensifer, contains 12 species, the genus Mesorhizobium, 19 species, the genus Bradyrhizobium, 8 species, the genus Phyllobacterium, 7 species and the genus Azorhizobium, 2 species (http://www.bacterio.cict.fr). This high number of species able to nodulate legumes (Table 1) was unexpected only two decades ago and the previsions are that it will be increased in next times since currently there are eight new species in press in IJSEM.
Traditionally considered rhizobia species and nodulating non-rhizobia species classification.
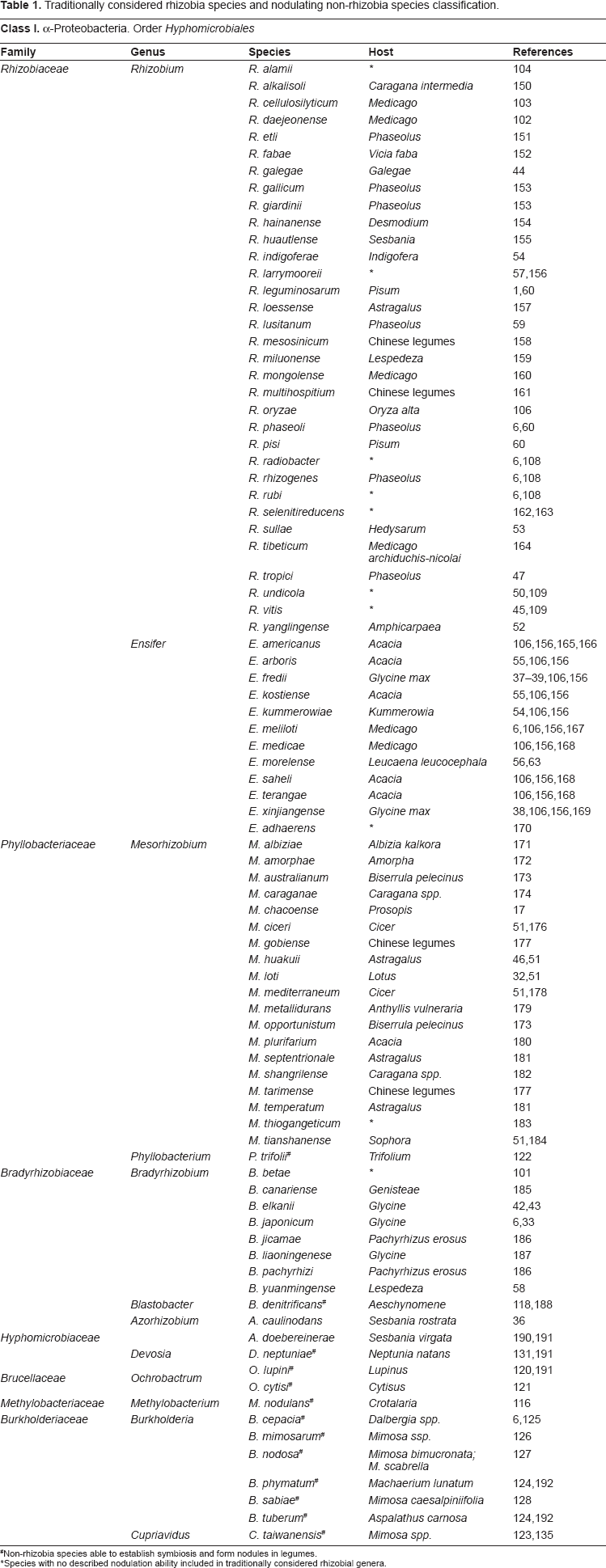
Non-rhizobia species able to establish symbiosis and form nodules in legumes.
Species with no described nodulation ability included in traditionally considered rhizobial genera.
Some Problems in Rhizobial Taxonomy
Along the rhizobial history some problems have been produced in the taxonomy of these microorganisms because rhizobiologists do not take into account the existence of general rules for species nomenclature. They do not consider that the species reclassifications carried out in the Bergey's Manual are not official and that while they are validated in the official journal of bacterial systematics the old names remain valid. This occurred in the case of R. phaseoli and R. trifolii, which were considered not valid from 1984 because Jordan reclassified these two species into R. leguminosarum in his chapter of Bergey's Manual. 31 However, as this reclassification was never validated, all these species remain valid until 2008 in which our investigation group revised the taxonomic status of these species. 60 Our results showed that the valid species are R. leguminosarum and R. phaseoli, whereas R. trifolii is a later synonym of R. leguminosarum. Moreover, a problem with the type strains of R. leguminosarum deposited in different culture collections was detected and a new species named R. pisi was proposed to include one of these two strains deposited in DSMZ and NCIMB collections from Germany and United Kingdom, respectively.
Nomenclatural problems also affected to the order Rhizobiales and family Rhizobiaceae that as was already mentioned are illegitimate names because it was not considered the existence of previous valid names for these taxa. The same type of problems had occurred in the case of genus Sinorhizobium whose name has been recently changed to Ensifer. 106 This problem was originated because a species named Ensifer adhaerens described before than genus Sinorhizobium was not considered at the time of its description. Considering the rules of the Bacteriological Code, the name Ensifer has priority over Sinorhizobium since it was described before and thus it is clear that the proposal by Willems et al 107 to reclassify E. adhaerens into genus Sinorhizobium is not acceptable as was pointed out in the decision of the Judicial Commission of the International Committee on Systematics of Prokaryotes whose decision has been very recently published. 108
The use of virulence or symbiotic genes in taxonomy has also caused many problems such as that detected in the former genus Agrobacterium when Sawada et al 109 showed the total identity between the 16S rRNA gene sequences of the type strains from A. radiobacter and A. tumefaciens. Thus they proposed to reject the name A. tumefaciens since the name A. radiobacter has priority according to the nomenclature rules. Therefore the species A. radiobacter contains strains tumorigenic and non-tumorigenic strains depending on the presence of the plasmid pTi containing the virulence genes.109,110 A second error affected to the former species A. rhizogenes (Fig. 7) that contains strains able to induce hairy-roots in plants. 110 From the works of Willems and Collins 111 and Yanagi and Yamasato 112 it was found that this species phylogenetically belong to the genus Rhizobium and concretely it is close to R. tropici. This initial error was crucial in the later decision for reclassification of genus Agrobacterium into genus Rhizobium mainly based in the 16S rRNA gene sequences. 108

Atypical ineffective nodules formed by Rhizobium rhizogenes in Phaseolus vulgaris.
It is typical among the rhizobia researchers do not easily accept the official taxonomic changes in the systematics of these bacteria. They were reluctant to accept the change of the name Rhizobium meliloti to Sinorhizobium meliloti, also the proposal to reject the name Agrobacterium tumefaciens has not been until completely accepted and this name is currently used by some researchers. Also the reclassification of Agrobacterium and Allorhizobium into genus Rhizobium is subject of controversy and several letters to the editor have been published in IJSEM.113,114 However, since this reclassification was accepted in this journal must be accepted. Finally, in the last meeting of the International Committee on Systematic Bacteriology Subcommittee on the Taxonomy of Agrobacterium and Rhizobium the members decided to do not accept the name Ensifer and proposed to use the name Sinorhizobium.
The Non-Rhizobia Nodulating Legumes
In the past eight years, several bacteria capable of forming nodules and fixing nitrogen in legume roots have been documented and grouped within alpha and beta Proteobacteria, which include Methylobacterium nodulans,115,116 Burkholderia sp., 117 Blastobacter denitrificans, 118 Devosia neptuniae, 119 Ochrobactrum lupini 120 and O. cytisi, 121 Phyllobacterium trifolii, 122 Ralstonia taiwanensis (renamed as Cupriavidus taiwanensis), 123 Burkholderia tuberum, B. phymatum, 124 B. cepacia, 125 B. mimosarum, 126 B. nodosa 127 and B. sabiae. 128
As has been previously mentioned a common error in the rhizobial taxonomy was the use of symptoms in plants as a criterium for classification and identification of isolates. This lead to other error that was to consider the nodulation of legumes as an exclusive ability of rhizobia and thus the strains isolated from nodules that do not present the typical colonies on YMA plates are discarded. For instance, the colonies with this typical aspect are named Rhizobium even without further identification. This situation dramatically changed from 2001 ahead when the 16S rRNA gene sequencing was applied to the identification of the isolates. In this year was published in Nature the nodulation of legumes by Burkholderia, a genus that belong to beta-Proteobacteria. 117 This genus contains diverse species with different physiological and ecological properties, is a common soil habitant bacterium, often associated with plant roots, but also has been isolated from animals and humans. Moulin et al 117 isolated a strain of Burkholderia from nodules of the South African legume, Aspalathus carnosa. This isolate is also able to form nodules in the roots of Macroptilium atropurpureum, a tropical legume, confirming to have the ability of symbiotic nitrogen fixation. The Burkholderia strain carried nodulation (nodABC) genes phylogenetically related to those found in legume symbionts of the class alpha Proteobacteria (“classic” rhizobia) supporting the hypothesis of lateral gene transfer in the rhizosphere, crossing the boundary between class alpha Proteobacteria and beta Proteobacteria.
In the same year the nodulation of Crotalaria by a strain of genus Methylobacterium from alpha Proteobacteria 115 later named M. nodulans 116 was reported. Phylogenic analysis based on 16S rRNA of nodulating strains of Methylobacterium shows their close relationship to other species of Methylobacterium 115 whereas it was confirmed that this species harbors the common nodulation nodABC genes and nifH gene encoding structural nitrogenase enzyme.115,116 Phylogenic analysis based on nifH gene shows that it is related to Gluconacetobacter diazotrophicus, a sugarcane nitrogen fixing endophyte. 129
A typical case of bacteria classified into the genus Rhizobium by its morphology in YMA medium was that of Rhizobium neptuniae isolated from Neptunia natans nodules. 130 When we analyzed the LMW RNA profiles of these bacteria we proved that they are not typical of any rhizobial genera 119 and the analysis of the 16S rRNA gene sequence confirmed that they belong to a new species of genus Devosia, named Devosia neptuniae. 131 The strains of this species carry nodD and nifH genes closely related to those of R. tropici CIAT899T indicating that they were transferred to D. neptuniae from R. tropici an American species nodulating Leucaena. 47 Later this hypothesis was supported by the finding that R. tropici nodulated Neptunia in America. 132
In the year 2002, besides that of Neptunia by Devosia, the nodulation of Aeschynomene indica by Blastobacer denitrificans was reported. 118 Aschynomene indica is most frequently nodulated by Bradyrhizobium japonicum and Bradyrhizobium elkanii. 133 Nevertheless, Blastobacter denitrificans is able to form effective nodules in the roots of A. indica and reduce the atmospheric nitrogen 118 which was again confirmed by presence of nifHDK genes by southern hybridization. Although the reclassification of Blastobacter denitrificans into genus Bradyrhizobium has been proposed 134 as this proposal has been not until validated in IJSEM it cannot be considered as official.
In 2003 the nodulation of Mimosa by Ralstonia taiwanensis or Wautersia taiwanensis was published. 123 This species was isolated from nodules of Mimosa pudica and Mimosa diplotricha. In this case the strain was erroneously classified as Ralstonia and it has been later named Cupriavidus taiwanensis. 135 At present, Cupriavidus is a beta Proteobacteria, belonging to family Burkholderiaceae of order Burkholderiales. Several species of this genus were isolated from soil and human clinical specimens 135 but C. taiwanensis was the only species able to form effective nodules and fixes atmospheric nitrogen in legumes. C. taiwanensis carries ten nodulation genes nodBCIJHASUQ and one regulatory gene nodD on pRalta. 136 Next to nod genes, C. taiwanensis carries 19 genes, presumably arranged in five operons and covering 25 kb that are involved in nitrogenase synthesis and functioning. These operons are nifA, encoding a sigma 54-dependent regulator, nifENfdxBnifQ and a modified nifX, nifVWfixABCX, nifBfdxNnifZfixU and the nitrogenase structural genes nifHDK. 136
In year 2004 strains from genus Ochrobactrum from family Brucellaceae were found in nodules of Acacia mangium but no information on their symbiotic genes was reported. 137 Later a new species of this genus carrying symbiotic genes close to those of rhizobia was reported as endosymbiont of Lupinus honoratum nodules. 120 It harbors megaplasmids of 1500, 200 and 150 kb and the nodulation (nod) and nitrogen fixation (nif) genes were detected in all the sym plasmids using nifH and nodD probes. Besides, in 2007 a second new species of Ochrobactrum named O. cytisi was isolated from Cytisus scoparius nodules in Spain. 121 O. cytisi strains contained nodD and nifH genes on megaplasmids that were related phylogenetically to those of rhizobial strains nodulating Phaseolus, Leucaena, Trifolium and Lupinus.
In the same year 2005 a new species named Phyllobacterium trifolii was isolated from Trifolium pratense nodules. 122 Phyllobacterium species was originally proposed to accommodate bacteria isolated from leaf nodules of Rubiaceae and Myrsinaceae tropical plants 34 but the plasmid profile revealed that P. trifolii harbors symbiotic plasmids in which nodulation and nitrogen fixation genes are located and experiments with this species revealed that it forms atypical nodules in the roots of Trifolium repens and Lupinus albus 122 (Fig. 8).

Atypical ineffective nodules formed by Phyllobacterium trifolii in Lupinus.
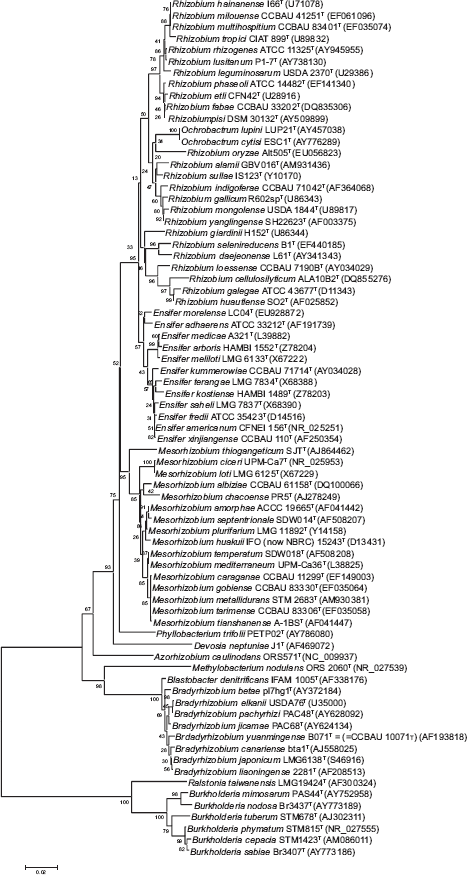
Neighbour-joining phylogram showing relatedness of partial 16S rDNA gene sequences of type strains from bacteria able to nodulate legumes. Bootstrap confidence values were estimated from 1000 replications of each sequence. Bar, 2 substitutions per 100 nucleotide positions.
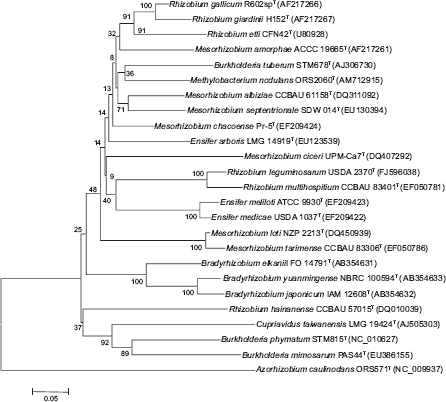
Neighbour-joining phylogram showing relatedness of partial nodC gene sequences of representative type strains from bacteria able to nodulate legumes. Bootstrap confidence values were estimated from 1000 replications of each sequence. Bar, 5 substitution per 100 nucleotide positions.
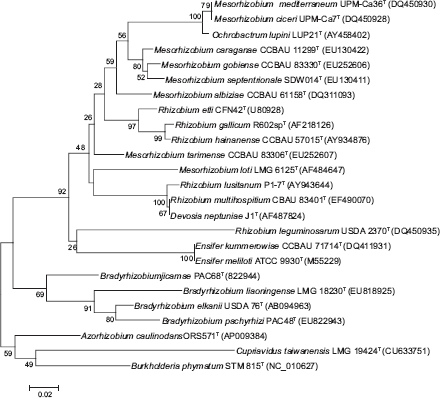
Neighbour-joining phylogram showing relatedness of partial nifH gene sequences of representative type strains from bacteria able to nodulate legumes. Bootstrap confidence values were estimated from 1000 replications of each sequence. Bar, 2 substitutions per 100 nucleotide positions.
On the other hand in the same year we demonstrated the nodulation of Phaseolus vulgaris by two pathogenic strains of the former species Agrobacterium rhizogenes (currently Rhizobium radiobacter) and the presence in these strains. The nodD and nifH genes found in these strains were close to those nodulating Phaseolus vulgaris and together with the symbiotic plasmids the presence of tumourigenic or hairy-root inducing plasmids was shown. 138
From 2006 to 2008 several new species of Burkholderia have been isolated from Mimosa nodules, B. mimosarum, 126 B. nodosa 127 and B. sabiae. 128 B. mimosarum was isolated from root nodules of M. pigra and M. scabrella from Taiwan, Brazil and Venezuela and several isolates showed effective nodulation in Mimosa species. The presence of nif and nod genes in the genomes has been demonstrated.139,140 B. nodosa was isolated from root nodules on Mimosa bimucronata and Mimosa scabrella from Brazil and produced N2-fixing nodules on Mimosa pudica, M. diplotricha and M. pigra. Also the nodulation of Mimosa by B. phymatum have been reported. 141 Although all the most reports are from mimosoid legumes, there is evidence that papilionoid legumes such as Cyclopia that is nodulate by B. tuberum 142 and Dalbergia louveli 125 that is reported for the first time that a strain belonging to the Burkholderia cepacia complex are able to induce efficient nodules on these legume plants. Even very recently it has been shown that some strains of Burkholderia are more competitive than R. tropici for the nodulation of Mimosa. 143 Also, preliminary evidence suggests that nodulating strains of both Burkholderia may have obtained nod genes from local rhizobia. 140
The fact that the closest known relatives of these species in the 16S rRNA tree are nonsymbiotic taxa and presence of very similar and phylogenetically related nodABC genes in nodulating alpha and beta Proteobacteria strongly implies that the genes required for legume nodulation are thought to have been acquired subsequently by lateral transfer from undefined sources, thus converting soil saprophytes into symbionts. 143 This hypothesis has been supported by many works125,144,145 but it is not clear whether a single transfer event was responsible for the spread of nodulation genes.
Finally, several studies reported the presence of gamma Proteobacteria in legume nodules but until now symbiotic genes have been not detected in these strains.146–148 Nevertheless as Benhizia et al 146 say they could nodulate legumes since recently it has been reported the legume symbiosis may exist in absence of nod genes as occurs in the case of photosynthetic bradyrhizobia. 149
Anyway, all these findings showed that the legume symbiosis are until poorly understood and the needed of further studies on the legume endosymbionts in different ecosystems and geographical locations mainly of those remaining unexplored from a symbiotic point of view. The set of genes available, data from several genes as housekeeping genes and the availability of new set of tools can provide useful insights to unravel relationships in these groups. Today, with modern molecular techniques this is a realistic goal. Inclusion of such molecular information in future phylogenetic analyses could lead to a reassessment of the position of these species and help to resolve the taxonomic problems.
Publish with Libertas Academica and every scientist working in your field can read your article
“I would like to say that this is the most author-friendly editing process I have experienced in over 150 publications. Thank you most sincerely.”
“The communication between your staff and me has been terrific. Whenever progress is made with the manuscript, I receive notice. Quite honestly, I've never had such complete communication with a journal.”
“LA is different, and hopefully represents a kind of scientific publication machinery that removes the hurdles from free flow of scientific thought.”
Available to your entire community free of charge
Fairly and quickly peer reviewed
Yours! You retain copyright
Footnotes
The authors report no conflicts of interest.
