Abstract
This note follows up on an observation concerning a possible frameshift in the DNA sequence of exon 2 of rabbit interleukin 4 (IL4) made in a comparative study of the rabbit Th2 cytokine region sequences from two different rabbits. One was from the tuberculosis-susceptible Thorbecke strain, whose whole genome was sequenced at Broad Institute (OryCun2.0), and the other a normal NZW rabbit studied as part of the ENCODE project. If present, a frameshift could have resulted in exon skipping and production of IL4δ2 protein. We resequenced DNAs of the Thorbecke OryCun2.0 donor rabbit, another rabbit of the same strain, and a third rabbit of the inbred B/J strain in the region in question. All three had sequences identical to the normal NZW in the ENCODE ENm002 assembly in that region and hence no frameshift. The sequence information was submitted to GenBank and assigned accession numbers JQ687218 and JX073284.
Introduction
Alternative splicing of RNA encoding rabbit interleukin 4 (IL4) was reported by Perkins and coworkers more that twenty years ago. 1 Although the functional significance of protein products produced from alternative splicing was not known, recent studies have suggested that forms such as those lacking the second exon (IL4δ2) may have unique functional properties.2,3 In the course of a study comparing genomic DNA sequences of two different rabbits, 4 we found that the Thorbecke strain animal chosen for whole genome sequencing of the rabbit (OryCun2.0) had a possible frameshift in exon 2 of IL4 compared to a normal NZW rabbit DNA sequenced by the ENCODE project (ENm002). Although the Broad Institute's sequence coverage averaged 6.51x in the OryCun2.0 assembly, they had only two traces that may have been used in the assembly of the region in question. We raised the possibility that if a frameshift was indeed present, forced exon skipping and production of IL4δ2 could have affected some immunological responses of rabbits of this strain. 4 They were already known to be more susceptible to Mycobacterium tuberculosis infection than outbred NZW from the same supplier. 5 To follow up on this observation, we obtained and sequenced DNA from the OryCun2.0 rabbit and another rabbit of the same strain. DNA from another inbred rabbit of the B/J strain was also sequenced as a control. Our sequencing clearly showed that there was no frameshift. It was the borderline quality of the Broad's sequence of only two traces for the region of interest that led to the apparent frameshift included in the OryCun2.0 assembly.
Methods
Primers
Primers were synthesized by Integrated DNA Technologies (WWW.IDTDNA.COM). The external PCR primer pair Forward FP and Reverse RP were selected using NCBI primer BLAST and internal primer pair Forward FPN and Reverse BPN were selected using MacVector 12.5 (Table 1). The product lengths were expected to be 476 base pairs (bp) for external primer pair and 319 bp with the internal nested primers.
Primers used for PCR, nested PCR, and sequencing reactions.

DNA
Frozen samples of approximately 5 µg of DNA from the OryCun2.0 donor rabbit labeled 272.0 and from a second rabbit of the same strain labeled 299.0 were a kind gift from Jessica Alfoldi and Jeremy Johnson, of the Broad Institute. DNA from another inbred rabbit strain (B/J) was used as a control. Dr. Akira Seto formerly of the Shiga University of Medical Science Shiga, Japan shipped the B/J DNA to NIH after he retired. It had been stored frozen as an alcohol precipitated pellet since 1995.
PCR
Great care was taken to avoid contamination at all steps of the project. Gloves were changed frequently and DNA samples were diluted in a room distant from where PCR reactions were set up. No rabbit DNA had been previously used in either room. The HotStarTaq PLUS DNA Polymerase kit (QIAGEN #203601) was used following the manufacturer's directions with two different MgCl2 concentrations; a set of duplicates with 1.5 mM MgCl2 in the final reaction volume, and an additional set of duplicate PCR reactions with a final concentration of 3.5 mM MgCl2. For the PCR, target DNA samples 272.0 (OryCun2.0 donor), 299.0 (second Thorbecke rabbit) and B/J were each diluted to 5 ng per µl so that 5 µl would contain 25 ng. PCR reactions were programmed into an MJ Research Inc. PTC-100 Thermocycler with hot bonnet. First the DNA samples were denatured at 95 °C for 10 min followed by cooling to room temperature. A separate tip for each tube was then used for additions of 45 µl of a master mix containing 0.5 µM primers, dNTP mix (200 µM of each dNTP), and HOTSTARTaq polymerase 2.5 units per reaction. The first round of PCR consisted of 5 min at 95 °C to activate polymerase, then touchdown steps at 94 °C 1 min, 68 °C 30 sec, 72 °C 30 sec, decreasing by 2 °C over the next 4 rounds. A 1 min 30 sec extension at the fifth 58 °C round was followed by 29 rounds with 94 °C 1 min, 56 °C 30 sec, 72 °C 30 sec, followed by a final 5 min extension at 72 °C. A second set of nested PCR reactions used 2.5 µl of the first round reaction products added to 47.5 µl of the master mixes with the internal primers (Table 1). After the 95 °C step to activate the polymerase, the touchdown PCR protocol was similar to the first round except that it started at 66 °C decreased by 2 °C down to 58 °C and after the 1 min 30 sec 72 °C step, extension continued at 56 °C for 30 rounds and a 5 min extension step at 72 °C. Gel electrophoresis was used to determine that products of the expected sizes were obtained. The Qiagen MinElute PCR Purification kit was used to purify first round products according to the Manufacturer's directions because sufficient products for sequencing were obtained from them.
Sequencing
Tubes containing purified first round products (15 ng) premixed with either the forward (FPN) or reverse (BPN) internal primers (Table 1) were sent for sequencing by standard Sanger DNA sequencing at GENWIZ (www.genwiz.com) following the instructions they provided. The 272.0 and 299.0 products were from reactions in 1.5 mM MgCl2 and the B/J product was from reactions in 3.5 mM MgCl2.
Results
Figure 1A shows a portion of the sequence of IL4 in the Oryctolagus cuniculus ENCODE region ENm002 genomic scaffold (gi|217273035|gb|DP001045.1|). The encoded amino acid sequence between positions 830654-830788 (IL4 exon 1) is shown below the codons. Similarly the encoded amino acids of IL4 exon 2 are shown below codon positions 830964-831011. The external primers are shown in red and underlined. The internal nested and sequencing primers are highlighted in yellow and the region where a fifth C was found by Broad within exon 2 in the OryCun2.0 sequence is highlighted in blue. Figure 1B shows alignment of OryCun2.0 with the corresponding ENCODE region between positions 830551 and 831300. There are three locations where an extra base was reported in the OryCun2.0 sequence compared to ENm002. These three extra bases were not found in any of our sequences. Figure 1C shows alignment of our sequences of DNA sample 272.0 from the OryCun2.0 donor compared to ENm002. The second Thorbecke inbred rabbit's DNA sample 299.0 gave the identical result. The B/J rabbit also had identical sequence for the regions in question but an extra G was found in duplicate PCR products on both strands after intronic position 153 of the 465 bp sequence shown in Figure 1C. The sequences were submitted to GenBank and given accession numbers JQ687218 (Thorbecke rabbits) and JX073284 (B/J rabbit).

Sequences of the region of interest in rabbit interleukin 4 (IL4). (
Figure 2 shows the PCR products from the first round of amplification that were obtained and used for sequencing. Products obtained in duplicate from the OryCun2.0 donor DNA 272, from the second rabbit 299 of the Thorbecke strain and from the DNA of the B/J strain are shown with the MassRuler DNA ladder (Fermentas). They were of the predicted sizes. Panel A shows products obtained with amplifications containing 1.5 mM MgCl2 from DNA of rabbits 272 and 299 were purified for sequencing. Panel B shows products obtained with amplifications containing 3.5 mM MgCl2 from DNA of the B/J strain were purified for sequencing.
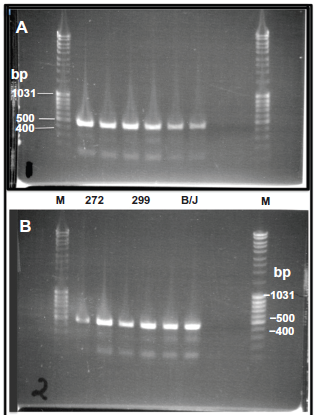
PCR products. PCR products obtained from the first round PCR that were used for sequencing. Lanes marked M show the MassRuler DNA ladder mix loaded at 10 µl/well (100 ng of 1031 bp and 500 bp bands and 40 ng of 400 bp band). (
Figure 3 shows sequencing chromatograms using the forward primer FPN where positions 192–195 are the four Cs and using the reverse primer BPN where the complementary four Gs are at positions 80–83.
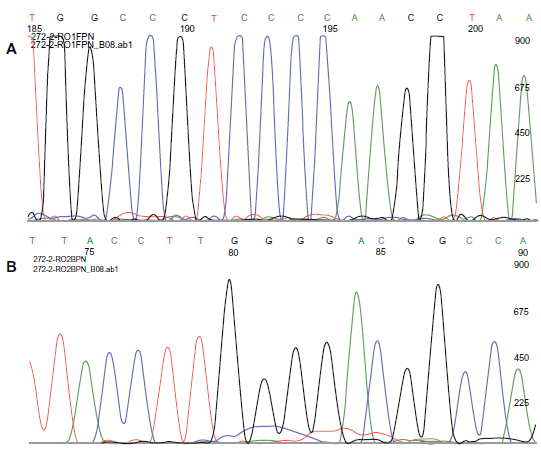
Sequencing chromatograms of a sample of OryCun2.0 272.0 donor DNA When forward primer FPN was used, positions 192–195 correspond to four Cs. When the reverse primer BPN was used, the complementary four Gs are found at positions 80–83.
Concluding Remarks
The trace archives of the Broad Institute's sequence of Oryctolagus cuniculus were reported to have 7.5x coverage. However, the OryCun2.0 assembly reflects 6.51x coverage because it only utilized traces that passed their quality control cut off. In spite of the high overall quality of the Orycun2.0 assembly, only two traces of questionable quality corresponding to the region of interest in exon 2 of rabbit IL4 were in the trace archive. The sequence in question that showed the frameshift in exon 2 of IL4 and was included in the OryCun2.0 assembly was 569 bases into the sequence of gnl|ti|2047213760. Our sequencing showed that the sequence that could have led to alternatively spliced IL4δ2 was incorrect. No frameshift is present and thus the IL-4 DNA sequence of the Thorbecke strain is identical to the normal NZW rabbit sequenced for ENCODE.
Authors Contributions
Conceived and designed the experiments: RGM, MGM. Analysed the data: RGM, MGM. Wrote the first draft of the manuscript: RGM. Contributed to the writing of the manuscript: RGM, MGM. Agree with manuscript results and conclusions: RGM, MGM. Jointly developed the structure and arguments for the paper: RGM, MGM. Made critical revisions and approved final version: RGM, MGM. All authors reviewed and approved of the final manuscript.
Funding
This research was supported by the Intramural Research Program of the National Institutes of Health, NIAID.
Disclosures and Ethics
As a requirement of publication authors have provided to the publisher signed confirmation of compliance with legal and ethical obligations including but not limited to the following: authorship and contributorship, conflicts of interest, privacy and confidentiality and (where applicable) protection of human and animal research subjects. The authors have read and confirmed their agreement with the ICMJE authorship and conflict of interest criteria. The authors have also confirmed that this article is unique and not under consideration or published in any other publication, and that they have permission from rights holders to reproduce any copyrighted material. Any disclosures are made in this section. The external blind peer reviewers report no conflicts of interest.
Footnotes
Author(s) disclose no potential conflicts of interest.
Acknowledgements
We appreciate the gifts of Thorbecke strain DNA from Jessical Alfoldi and Jeremy Johnson of the Broad Institute and B/J DNA from Akira Seto, Shiga, Japan. Thanks to David Margulies and William Paul for making laboratory space, equipment and supplies available and to Jinfang Zhu and Kannan Nataranjan for helpful advice. We also appreciate comments on the manuscript from David Margulies, Kannan Nataranjan, Jinfang Zhu and Alejandro Schäffer.
