Abstract
We examined malaria infection in asymptomatic blood donors from Mali, analyzing allelic diversity of Plasmodium falciparum (Pf) merozoite surface proteins (msp) -1 and -2 as well as the distribution of sulphadoxine-pyrimethamine (SP) resistance genes. A total of 140 genomic DNA samples were screened. Allele-specific nested polymerase chain reaction (PCR) analysis of Pfmsp-1 and Pfmsp-2 was performed, plus fragment analysis of the polymorphic regions to identify allelic diversity of the parasite population. We found parasite positivity due to Pf alone in 20.7% of these donors. Diverse allelic polymorphism of Pfmsp-1 and Pfmsp-2 was identified, with a high rate of multiplicity of infection (1.84 and 1.82 for Pfmsp-1 and Pfmsp-2, respectively). In addition, we found a high degree of SP resistance, with mutations at several dhfr and dhps codons. We conclude that there is an extensive diversity of Pfmsp-1 and Pfmsp-2 allelic types and SP drug resistance in Pf-infected donors from Mali.
Introduction
Malaria remains a disease of significant consequence in many parts of the world, especially among children in sub-Saharan Africa, despite advances in biomedical research.1–3 The most severe complications of the disease are primarily due to infection with Plasmodium falciparum (Pf). The disease is a significant contributor to reduced manpower and diminished economic output, and remains a scourge for non-immune travelers to endemic locations.4–7 Semi-immune individuals abound in endemic areas, their immune status being due to previous and multiple exposures. These individuals can be infected while being asymptomatic, and are responsible for transfusion-transmitted infections. Control programs are now focused on the design of new drugs, use of insecticide-treated bednets, vector control and aggressive vaccine design efforts. 8 Some of the vaccine design efforts are focused on blood stage subunit vaccines, of which merozoite surface proteins are principal components. Such efforts are hampered by Pf antigenic diversity of the merozoite surface proteins.9–12
Pf merozoite surface protein-1 (Pfmsp-1) is a major surface protein that plays a very significant role in erythrocyte invasion by merozoites. It is also a principal target in immune responses and is a candidate for blood stage subunit vaccine.13–15 Pfmsp-1 has 7 variable blocks separated either by conserved or semi-conserved regions. Block 2, a region near the N-terminal of the msp-1 gene, is the most polymorphic part and is under the strongest diversifying selection within natural populations. Multiple allelic types of msp-1 block 2 have been identified, including MAD20, K1 and RO33, ranging in size from 44 to 89 amino acids.11,12,16,17 Merozoite surface protein-2 (msp-2) is another subunit malaria vaccine candidate, 18 made up of highly polymorphic central repeats flanked by unique variable domains and conserved N- and C-terminal domains.19,20 Msp-2 has two allelic types, FC27 and 3D7. The extensive and varied antigenic diversity of msp-1 and msp-2 facilitates their usefulness as markers of malaria transmission dynamics and their ability to serve as effective surrogates of infection, infectivity and diversity in local populations.17,19,21,22
Drug-resistant parasites constitute another serious challenge hampering control efforts in endemic countries. These parasites interfere with prevention of malaria infection and pregnancy outcome among women in such locales, and could enhance the possibility of transfusion-transmitted infections, particularly among non-immune recipients.23,24 Sulphadoxine and pyrimethamine are widely available antimalarials, used for prophylaxis and treatment, to which parasites have developed resistance in recent years.25–33 Single nucleotide polymorphisms in the Pf genes for dihydropteroate synthase (dhps) and dihydrofolate reductase (dhfr) have been associated with resistance to sulphadoxine and pyrimethamine respectively, and such genetic markers serve as useful surrogates for monitoring emergence and dispersion of drug resistance.27,34–37 Evaluating the degree of drug resistance depends on identifying individual mutations in the genes dhfr (codons 50, 51, 59, 108 and 164) and dhps (codons 436, 437, 540, 581 and 613).38–40
Examining the degree of population diversity within malaria parasites in asymptomatic blood donors is essential for understanding the mechanism underlying disease pathology and the probability of transmission by transfusion. Additionally, unraveling the degree of drug resistance in such donors will assist in the establishment of a proper control strategy and the possible institution of a new therapeutic regimen. This study was designed to evaluate the genetic diversity of Pfmsp-1 and msp-2 in parasite isolates from blood donors in Bamako, Mali, and to assess the prevalence of mutations in specific genes involved in resistance to sulphadoxine and pyrimethamine.
Materials and Methods
Subjects, Sample Collection and Deoxyribonucleic Acid Extraction
A total of 140 asymptomatic blood donors were randomly selected from the Centre Nationale de Transfusion Sanguine (CNTS), Bamako, Mali and recruited for the study, after institutional review board approvals. All donors were male, which is representative of the global blood donor population. Following informed consent, blood was drawn into EDTA tubes, and genomic deoxyribonucleic acid (DNA) was extracted using the Puregene DNA extraction kit following manufacturer's instructions (Gentra Systems, Minneapolis, MN).
Multiplex polymerase chain reaction assay for single or mixed infections
We carried out a multiplex polymerase chain reaction (PCR) assay for differential diagnosis purposes and to determine if extracted genomic DNA samples had Plasmodium falciparum or Plasmodium vivax single or mixed infections. This assay utilizes the 18S rRNA gene, with primers, protocols and reaction setup following established methods. 41 Briefly, a universal forward primer, designed from a genus-specific region, and reverse primers specific to both the Pf and Pv gene sequences were utilized for PCR amplification. PCR was performed on an Eppendorf Mastercycler Gradient machine (Harlow Scientific, Arlington MA) in a total volume of 25 μL and amplified using the Lucigen EconoTaq Plus Green 2X Master Mix PCR system (Lucigen Corporation, Middleton WI). Final reaction mixtures contained 0.1 units/μL of EconoTaq DNA Polymerase, reaction buffer (pH 9.0), 400 μM dATP, 400 μM dGTP, 400 μM dTTP, 400 μM dCTP, 3 mM magnesium chloride (MgCl2), 1 μM forward and reverse primers and a proprietary mix of PCR enhancer/stabilizer and blue and yellow tracking dyes (Lucigen Corporation, Middleton WI). Five μL of PCR products were loaded onto a 2% ethidium-bromide stained agarose gel (SeaKem Agarose, Rockland ME) and band size was determined with the Doc-It LS Image Analysis Software (UVP Life Sciences, Upland CA), and a 1.5kb DNA ladder serving as marker (New England Biolabs, Boston MA).
Genetic diversity of msp-1 and msp-2
Primers for the polymorphic regions of interest (msp-1 block 2 and msp-2) were used for a two-step PCR amplification of genomic DNA, as previously published. 22 In brief, a primary amplification of the outer regions was followed by a secondary PCR using primary products as a template, with family-specific primer pairs. PCR conditions were altered to 25 μL final volume and amplified using the Lucigen EconoTaq Plus Green 2X Master Mix PCR system (Lucigen Corporation, Middleton WI). Final reaction mixtures contained 0.1 units/μL of EconoTaq DNA polymerase, reaction buffer (pH 9.0), 400 μM dATP, 400 μM dGTP, 400 μM dTTP, 400 μM dCTP, 3 mM MgCl2, 1 μM forward and reverse primers and a proprietary mix of PCR enhancer/stabilizer and blue and yellow tracking dyes (Lucigen Corporation, Middleton WI). For primary PCR reactions, 1 μL of genomic DNA template was used, and 2 μL of the primary PCR product served as the template for the nested PCR reactions. Nested PCR products were evaluated using 2% ethidium bromide stained agarose gel, visualized and photographed. The number of fragments obtained of appropriate size were compared for each sample, counted and combined across families for each gene.
Allelic distribution and multiplicity of infection
Prevalence of msp-1 (K1, R033 and MAD20) and msp-2 (3D7 and FC27) allelic types was evaluated as the presence of PCR products of the type in the total number of amplified bands for the corresponding locus. Individual alleles were identified by fragment length and by the corresponding allele-specific primers used. Size of PCR products was estimated with a 100 bp DNA ladder marker (New England Biolabs, Boston MA) and fragment size analysis carried out with the Doc-It LS Image Analysis Software (UVP Life Sciences, Upland CA). Allelic family fragment size allocation was done according to published methods. 42 The multiplicity of infection (MOI), or mean number of Pf genotypes per infected subject, was calculated by dividing the number of amplified PCR fragments reflecting parasite genotypes at any of the two msp loci.
Nested PCR assay of dhps and dhfr genes
Primary and nested PCR amplifications of genomic DNA samples for dhps and dhfr gene fragments were performed using standard methods and previously published primers. 43 Briefly, reaction mixtures in 25 μL volumes were amplified using the Lucigen EconoTaq Plus Green 2X Master Mix PCR system (Lucigen Corporation, Middleton WI). Final reaction mixtures contained 0.1 units/μL of EconoTaq DNA Polymerase, reaction buffer (pH 9.0), 400 μM dATP, 400 μM dGTP, 400 μM dTTP, 400 μM dCTP, 3 mM MgCl2, 1 μM forward and reverse primers and a proprietary mix of PCR enhancer/stabilizer and blue and yellow tracking dyes. Two primary PCR reactions (for dhfr and dhps genes) were carried out, as well as five secondary reactions utilizing PCR products from primary reactions as templates (codons 50/51/59 and 108/164 for dhfr and codons 436/437, 540/581 and 613 for dhps). Negative controls without a DNA template were included for all PCR reactions for confirmation purposes. Cycling parameters for both primary and nested PCRs are as described, 44 with amplified bands confirmed by running 10 μL of the nested PCR on 2% agarose gel.
Restriction fragment length polymorphism of drug resistance genes
A 5 μL nested PCR product for dhps and dhfr genes was incubated with mutation-specific restriction enzymes, following the manufacturer's instructions (New England Biolabs, Beverly MA), in a 15 uL final reaction. Restriction enzymes selection, incubation protocol and dhfr and dhps variant analysis followed previously published methods. 45 In brief, TSP5091, XmnI, AluI and DraI were selected for analysis of wild type and mutants of dhfr codons 51, 59, 108 and 164 respectively. As for dhps, AvaII, FokI and BstUI were selected for analysis of wild type and mutants of dhfr codons 437, 540 and 581 respectively. Dhfr and dhps variants were identified as previously described. 43
Results
Allelic diversity of Pfmsp-1 and Pfmsp-2
Frequency of Plasmodium falciparum merozoite surface proteins-1 (Pfmsp-1) and merozoite surface proteins-2 (Pfmsp-2) allelic forms.
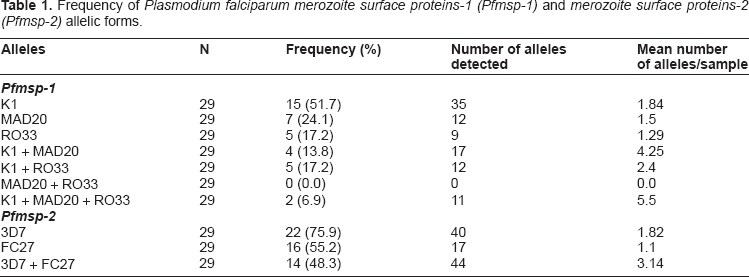
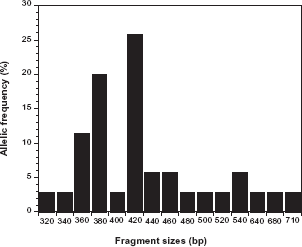
Allelic frequency distribution of Plasmodium falciparum merozoite surface protein-1 K1 alleles. K1 allelic type was the most common (55.6%), had fragment sizes ranging from 320–710 base pairs, with 420 bp (25.7%) the most common. Two (6.9%) samples analyzed had all three Pfmsp-1 allelic types.
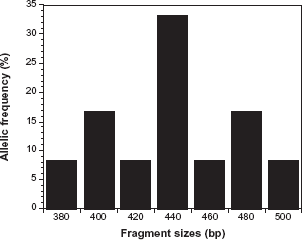
Allelic frequency distribution of Plasmodium falciparum merozoite surface protein-1 MAD20 alleles.
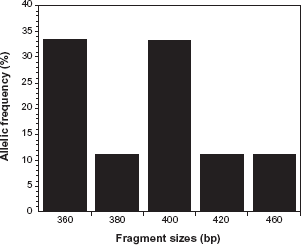
Allelic frequency distribution of Plasmodium falciparum merozoite surface protein-1 R033 alleles. RO33 allelic type was the least common (24.1%), with fragments ranging from 360–460 bp, with 360 bp (33.3%) the most common.
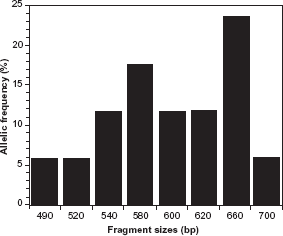
The Pfmsp-2 gene showed 20 total alleles for each family, with the 3D7 family being most prevalent (75.9% of overall alleles detected, compared to 55.2% of FC27). Additionally, a total of 57 allelic fragments, categorized into 17 (29.8%) and 40 (70.2%) fragments, assigned to the FC27 type and 3D7 type respectively, were recorded. Fourteen (48.3%) of all samples analyzed carried both of the Pfmsp-2 allelic types. 3D7 and FC27 allelic types had fragment sizes ranging from 490–780 base pairs, with 720 bp (22.5%) and 660 bp (23.5%) being the most common fragments for 3D7 and FC27 types, respectively (Figs. 4 and 5).
Allelic frequency distribution of Plasmodium falciparum merozoite surface protein-2 3D7 alleles. 3D7 allelic type was predominant (75.9%) and fragment sizes ranged from 520-780 bp, with 720 bp (22.5%) the most common. Allelic frequency distribution of Plasmodium falciparum merozoite surface protein-2 FC27 alleles. FC27 allelic type was less frequent (55.2%) and 660 bp (23.5%) most common fragment size. Fourteen (48.3%) of all samples analyzed carried both 3D7 and FC27 allelic types.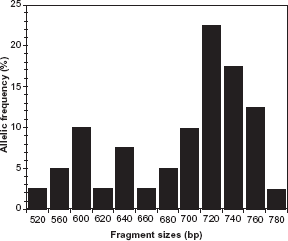
Restriction fragment length polymorphism (RFLP) digestion and analysis
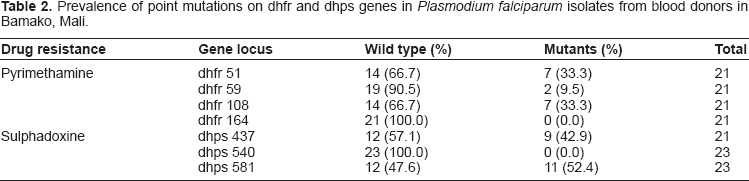
Molecular assays performed on the 29 malaria-positive samples from asymptomatic blood donors revealed that 21 (72.4%) and 23 (79.3%) were successfully amplified with specific Pf dhfr and Pf dhps primers respectively. Mutation analysis was carried out for dhfr isoleucine (Ile)-51, arganine (Arg)-59, asparagine (Asn)-108 and leucine (Leu)-164, with four restriction enzymes (AluI, XmnI, TSP509I and DraI), serving to detect mutations ascribed to antifolate resistance by the malaria parasite. Analysis of the dhps gene ascribed to sulphadoxine resistance for Glycine (Gly) -437, glutamate (Glu) -540 and Gly-581 was carried out with restriction enzymes FokI, AvaII and BstUI. The total prevalence of mutations found was 19.1% and 29.9% for dhfr and dhps genes respectively.
Pf dhfr polymorphism
Prevalence of point mutations on dhfr and dhps genes in Plasmodium falciparum isolates from blood donors in Bamako, Mali.
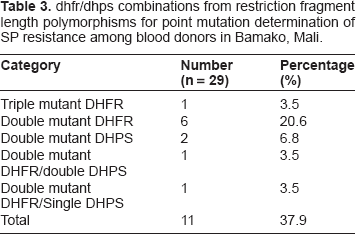
dhfr/dhps combinations from restriction fragment length polymorphisms for point mutation determination of SP resistance among blood donors in Bamako, Mali.
Pf dhps polymorphism
Analysis of dhps polymorphisms reveal PCR products of ~438 bp from nested PCR reactions. Evaluating the variants at codon 437 with the AvaII restriction enzyme reveal a wild type of 438 bp with mutations at this site shown by a cut fragment sized 404+ bp. This mutation indicates substitution of Gly for alanine (Ala). On the other hand, polymorphisms at codon 540 analyzed with the FokI enzyme produced fragments of 405 bp (wild type) or 320 and 85 bp (mutant). The wild type fragment indicates presence of lysine (Lys) while the mutant fragment reflects substitution of Glu for Lys. For codon 581, analysis with the restriction enzyme BstUI produced digests of sizes 105, 33 and 23 bp (wild type) and 138 and 23 bp (mutant). The wild type reflects presence of Ala while the mutant fragment reflects substitution of Gly for Ala (Table 2). Three dhps codons 437, 540 and 581 were evaluated for mutational analysis, as shown in Table 2, producing successful results at dhps codon 437 and 581, but not codon 540. For dhps, the frequency of mutations varied from a high of 47.8% (Gly-581) to a low of 42.9% (Gly-437). Additionally, some samples showed a mixed pattern with dhps double mutant Gly-437/Gly-581 (8.7%). Interestingly, we did not record any case of dhfr and dhps quintuple mutant (Table 3), carrying the dhfr triple mutant and dhps double mutant.
Discussion
Malaria remains a disease for which significant global efforts are focusing on integrated disease control. These efforts place an emphasis on vaccines that might be beneficial to non-immune travelers to endemic locales, and on reducing exposure to the parasite from transfused blood products.2,4,6 Our study of a group of blood donors from Bamako, Mali reveal malaria parasite positivity of 20.7%, and all infection due to Pf parasite. This report confirms previous studies, reinforces the significance of this species as a major disease agent in Mali, and the concomitant outcome arising from transfusing such infected blood or its products.
Deconvoluting the genetic diversity of the Pf parasite is critically important when designing a national treatment policy, vaccine immunogenicity studies and in terms of clarifying the local epidemiology of the parasite.10,12,42,46,47 In the present report, we examined allelic polymorphisms of Pfmsp-1 and Pfmsp-2 in 29 Pf-positive asymptomatic blood donors in Mali. We report extensive allelic diversity with a high rate of multiplicity of infection. This outcome is similar to reports from other endemic sub-Saharan African countries,42,46 the emphasis due to the fact these are blood donors.
Allele-specific analyses of Pfmsp-1 and Pfmsp-2 reveal different frequencies for both gene families, which confirms that the malaria parasite population in this group is highly diverse. Significantly, the number of Pfmsp-1 and Pfmsp-2 total alleles in the study group is higher than that found in studies involving patients with uncomplicated or severe malaria in sub-Saharan African or other endemic locations.48–50 This result clarifies the idea that there is a limited parasite population needed to cause symptomatic disease, which may not be the case in asymptomatic carriers. Collectively, these results suggest that highly diverse allelic variations of Pfmsp-1 and Pfmsp-2 exist in Mali blood donors, who are mostly mixed carriers.
Even though the allelic frequencies for both Pfmsp-1 and Pfmsp-2 alleles in the present study were considerably lower than that reported in some other patient studies,42,46,51 they are similar to those reported in a household survey analysis carried out in a malaria-endemic population from Haiti. 47 The population in our report are asymptomatic donors with probably very low parasitemia, not high enough to trigger clinical disease, but high enough to present a challenge to non-endemic travelers who might access these blood products. Additionally, these findings confirm that semi-immune status in residents of endemic countries is attributable to multiple exposures to the parasite over several years.
Molecular assays utilizing genetic markers permits determination of drug resistance in malaria. They are powerful tools for tracking drug resistance. The presence of dhfr and dhps mutations in Pf isolates obtained from asymptomatic donors reflects the effectiveness of treating malaria with sulphadoxine-pyrimethamine in Bamako, Mali. The prevalence of dhfr mutation in our study was as high as 33.3% in Ile-51 and Asn-108. Though this is lower than the 43% reported in Kolle, Mali, 52 it is similar to that reported in Ibadan, Nigeria. 44 The difference between our report and the other Mali group might be attributable to city conditions in Bamako compared to the rural Kolle area. In addition, the fact that our data compares with that reported for patients with symptomatic, uncomplicated malaria reveals the need for a revamping of the national malaria treatment policy, considering the use of sulphadoxine/pyrimethamine (SP) for chemoprophylaxis among pregnant women in West Africa. Prevalence of dhfr triple mutations in our study was 4.8%, lower than that from Tekete and colleagues. 52 This might be due to the fact that our patient population consists of asymptomatic donors, as opposed to malaria patients from other reports. Triple mutation frequency of dhps was as high as 47.8%, comparable to other reports. Interestingly, dhps double mutation was 8.7%, which also compares nicely to that found elsewhere. There is a rapid increase in SP resistance in Mali, which should be carefully analyzed, in case a change in national treatment policy is warranted.
In conclusion, we report that there is a high degree of SP resistance in our population, with mutations at several dhfr and dhps codons. Additionally, there are extensive polymorphisms with diverse allelic types of Pfmsp-1 and Pfmsp-2 genes among randomly selected Pf-infected donors, with a comparable degree of multiplicity of infection to patients with malaria infection. Naïve non-immune recipients of blood products from such donors have a very high degree of exposure to malaria parasite, and could receive transfusion-transmitted infection. Of serious significance to the local population is the possibility that patients not immune to a particular allelic subset may actually do worse if transfused with parasite-infected blood with those subsets. The high frequency of dhps and dhfr resistance genes calls into question the usefulness of SP as prophylactic or treatment agents in Bamako, Mali.
Funding Sources
This project was funded with the College of Science Faculty Education and Development Award and the Accelerated Research Funding Award, Rochester Institute of Technology (BNT). Patient recruitment and sample collection was carried out under an NIH/NIAID funding R01-AI-4236 (JMM).
Competing Interests
Author(s) disclose no potential conflicts of interest.
Author Contributions
Conceived and designed the experiments: BNT. Analyzed the data: BNT. Wrote the first draft of the manuscript: BNT. Carried out sample collection: JMM and DAD. Carried out the genotyping: BNT, CRP and SRC. Carried out the drug resistance studies: BNT and TJT. Contributed to the writing of the manuscript: SRC, TJT. Agree with manuscript results and conclusions: BNT, CRP, SRC, TJT, JMM, DAD. Jointly developed the structure and arguments for the paper: BNT, CRP, SRC, TJT. Made critical revisions and approved final version: BNT, JMM. All authors reviewed and approved of the final manuscript.
Footnotes
Acknowledgements
We are grateful to Mounirou Baby and Londyn Cullifer for technical assistance.
As a requirement of publication author(s) have provided to the publisher signed confirmation of compliance with legal and ethical obligations including but not limited to the following: authorship and contributorship, conflicts of interest, privacy and confidentiality and (where applicable) protection of human and animal research subjects. The authors have read and confirmed their agreement with the ICMJE authorship and conflict of interest criteria. The authors have also confirmed that this article is unique and not under consideration or published in any other publication, and that they have permission from rights holders to reproduce any copyrighted material. Any disclosures are made in this section. The external blind peer reviewers report no conflicts of interest.
