Abstract
CD14 is a multifunctional receptor expressed on many cell types and has been shown to mediate immune response resulting in the activation of an inflammatory cascade, with polymorphism of its promoter (rs2569190) found to be associated with susceptibility to several diseases. In malaria infection, the CD14 gene demonstrated a pathogenic profile in regulating experimental cerebral malaria, with reports of elevated levels of soluble CD14 in serum of patients but no definitive conclusion. We present a detailed analysis of genetic diversity of CD14 promoter gene (snp −159 C/T; rs2519190) polymorphism between a malaria-infected group and uninfected controls and its association with clinical parameters of disease. Genomic DNA samples obtained from 106
Introduction
Inflammatory reactions, secondary to pathogenic stimuli, are important for the initiation of necessary immune response for combating the external stimuli when effectively and efficiently deployed. Infection with
In published reports, genetic variations leading to dysfunctional monocyte functions have been implicated in disease susceptibility and outcome, probably clarifying how human populations have adapted to major infectious diseases through the ages.
14
Genetic variants of the CD14 gene (a C-to-T transition, snp −159 C/T) have been shown to be associated with susceptibility to multiple diseases and conditions including allergic rhinitis,
15
functional dyspepsia,
16
inflammatory bowel disease,
17
gram-negative bacteria, and respiratory infections, including brucellosis.18–21 Its significance in allergic reactions and outcome in patients with asthma have also been documented.21,22–24 In human, malaria infections particularly, elevated levels of soluble CD14 have been found in patients with acute infections,
25
concluding that soluble CD14 levels may play a critical role in disease severity. A murine infection model has been the clearest to delineate a specific pathogenic role for CD14 during experimental cerebral malaria infection, showing a protective outcome in
Although differing outcomes have been reported for the critical role of TLRs alongside CD14, its co-receptor, in mediating disease susceptibility or outcome in malaria infection,26–29 it is abundantly clear that this promoter gene potentially has a significant role in disease outcome. Would CD14 promoter gene (rs2519190) single-nucleotide polymorphism (SNP) be a mediating factor in who develops severe or acute disease in sub-Saharan Africa? Will the hereditary variants regulate initiation of immune response thereby affecting (positively or negatively) phagocytosis and the inflammatory cascade following infection as well as clinical parameters of disease? Dissecting the essentiality of CD14 promoter gene polymorphism in acute
Our objective is to investigate any association between CD14-snp159 C/T (rs2569190) gene polymorphism and susceptibility to acute malaria infection or other measures of disease severity, such as parasitemia, fever, and anemia. To do this, we conducted a detailed analysis of genetic diversity of CD14 promoter gene polymorphism in an infected group versus uninfected controls, recruited from sub-Saharan Africa. Genomic DNA samples obtained from 106
Materials and Methods
Subjects and demographics
A total of 106 patients (age range: 6 months-5 years), who presented with presumptive, acute malaria infection (initial diagnosis positive for
PCR confirmation for malaria infection
To confirm microscopy results, PCR assay targeting the 18S ribosomal RNA gene of
Genotyping for CD14 promoter gene (rs2569190) polymorphism
To assay for the SNP of the CD14 promoter gene (rs2569190), we used a previously published primer and protocol, 33 with some amendments. We amplified 1 μL of genomic DNA as template, with conditions optimized to 25 μL of final volume, as described. 1 About 5 μL of PCR products was examined on a 2% ethidium bromide–stained agarose gel; positive amplification yielded products of 561 base pairs (bp) and size estimated with a GeneRuler 100bp Plus DNA ladder (Thermo Fisher Scientific Inc, Springfield Township, NJ, USA).
RFLP assay
To determine genotypic and allelic frequencies of the polymorphic variants, we set up a RFLP assay using the FastDigest
Statistical analysis
Tests for deviation from Hardy-Weinberg equilibrium (HWE) were performed, with SNPs rejected based on the recommended threshold of
Results
We examined the genetic diversity of the CD14 gene promoter (−159 C/T; rs1800829) SNP that has been demonstrated to exhibit significant relevance within and between disease groups and evaluated its association with disease outcome in
Genotypic and allelic frequencies of CD14 (snp −159 C/T; rs2519190) promoter gene polymorphisms between patients with
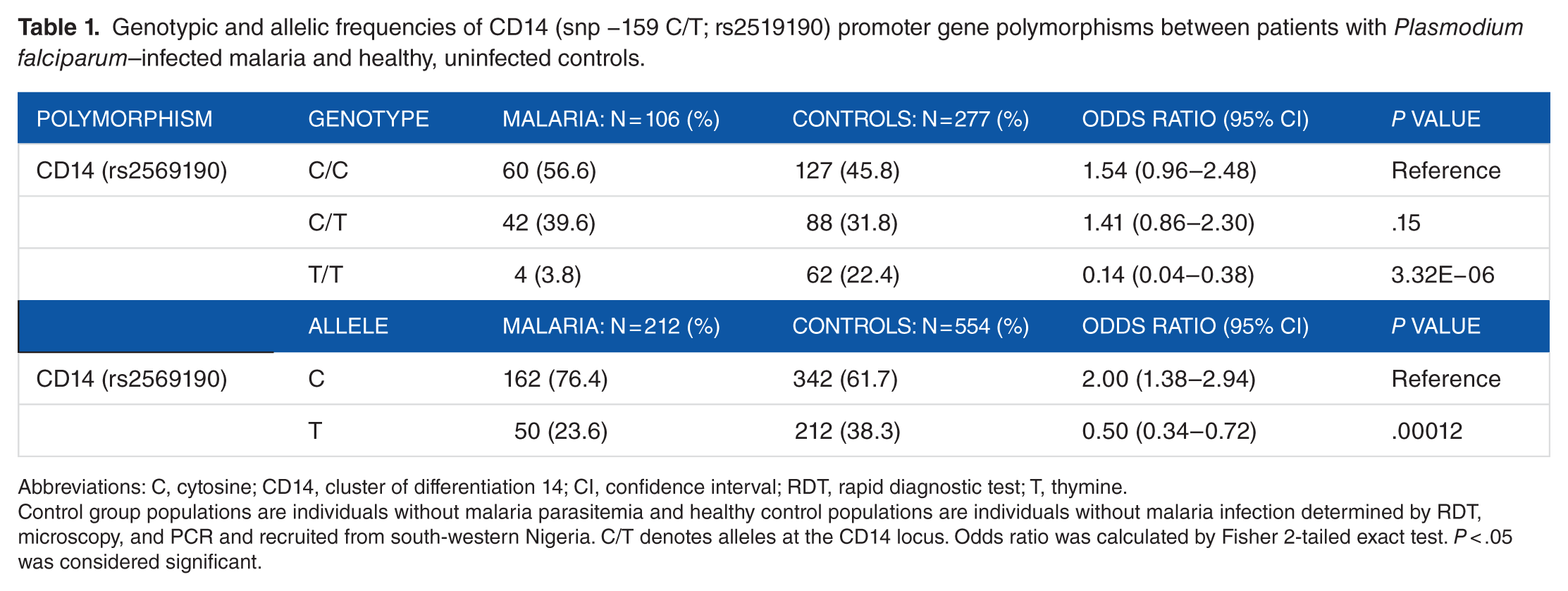
Abbreviations: C, cytosine; CD14, cluster of differentiation 14; CI, confidence interval; RDT, rapid diagnostic test; T, thymine.
Control group populations are individuals without malaria parasitemia and healthy control populations are individuals without malaria infection determined by RDT, microscopy, and PCR and recruited from south-western Nigeria. C/T denotes alleles at the CD14 locus. Odds ratio was calculated by Fisher 2-tailed exact test.
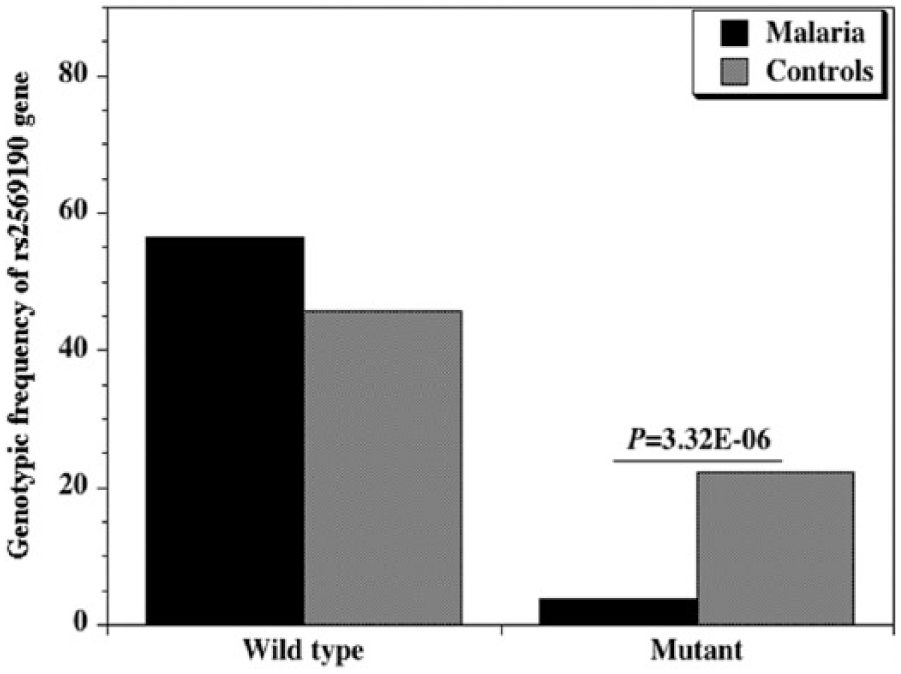
Genotypic frequency of CD14 gene promoter polymorphism (snp −159 C/T; rs2569190) between
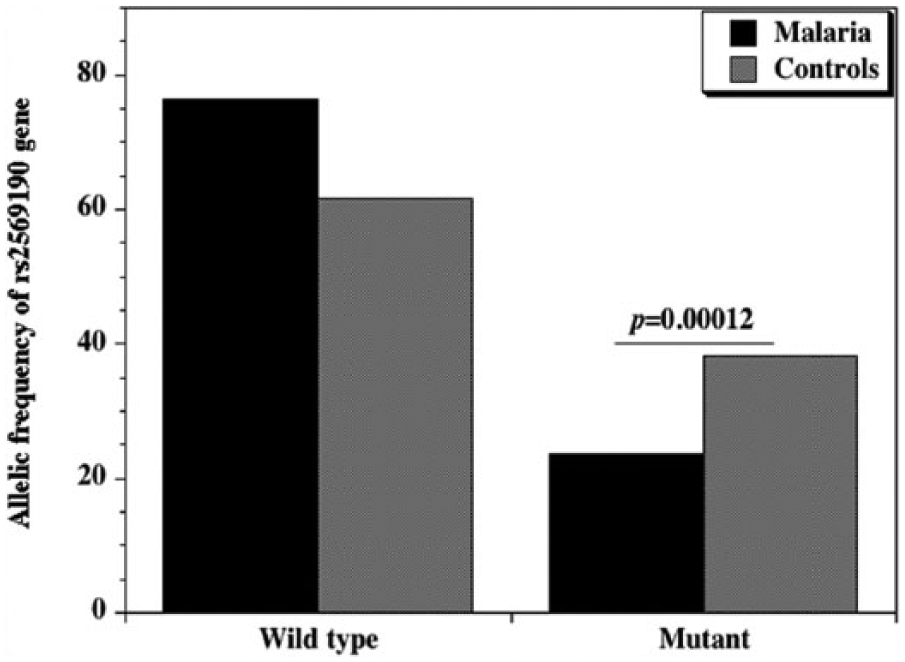
Allelic frequency of CD14 gene promoter polymorphism (snp −159 C/T; rs2569190) between
To determine whether the CD14 gene promoter genetic diversity results correlate with markers of clinical disease or severity, we evaluated parasite load (parasites/μL of blood), fever (°C), anemia (PCV), and age (months) among malaria-infected patients. We found no statistically significant difference or any association for age (mean age: 31.9 versus 38.7 months;
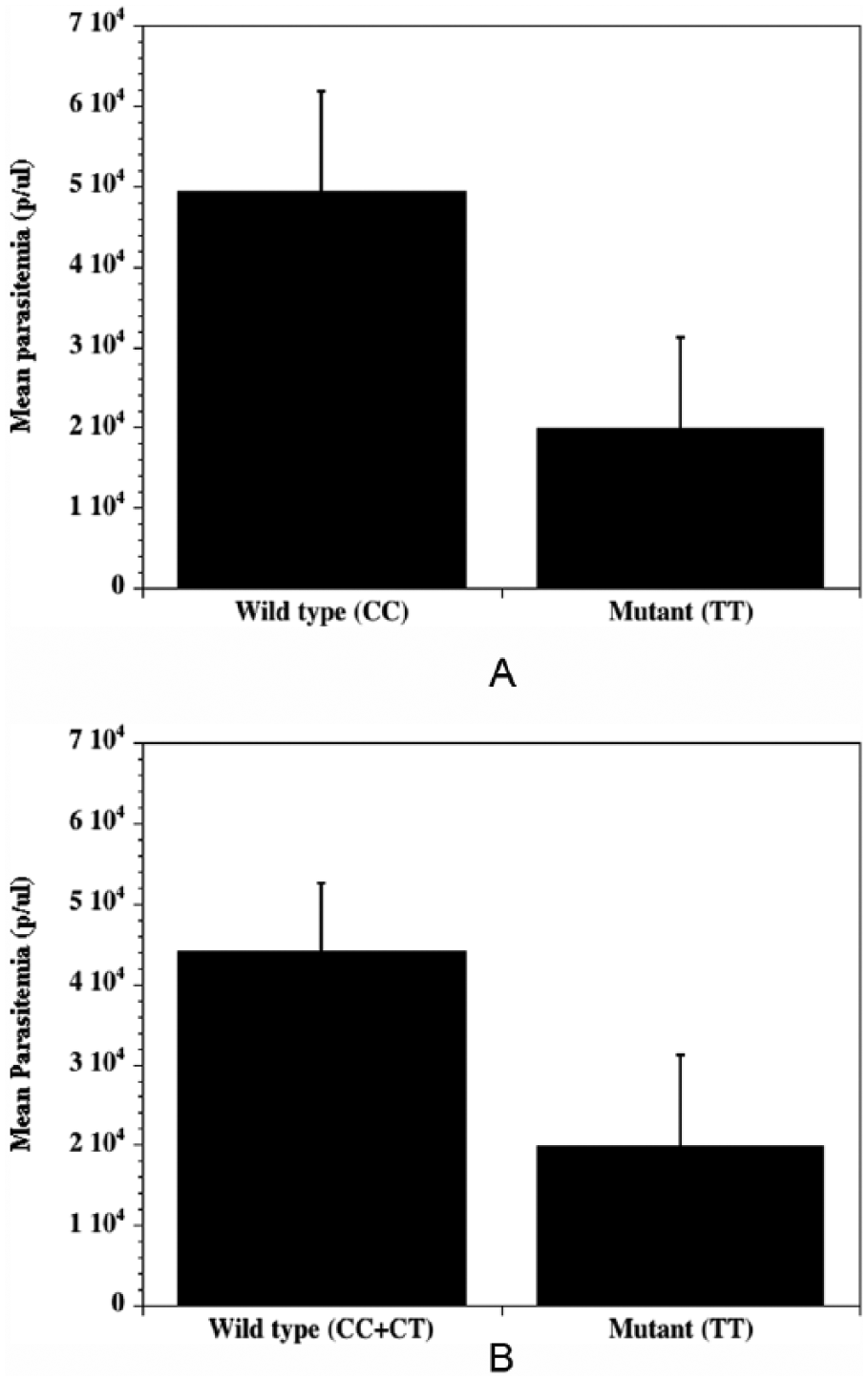
Mean parasite distribution between patients with malaria with either (A) wild-type (−159 C/C) or mutant (−159 T/T) variants and (B) wild-type (−159 C/C+C/T) versus mutant (−159 T/T) variants of the CD14 gene promoter polymorphism (snp −159 C/T; rs2569190).
Discussion
Malaria is still a disease that confounds a lot of researchers globally and the millions who fall sick to this infection constantly, especially considering the possibility of multiple infections in a single year. Despite the intensified efforts toward designing a vaccine or new therapeutic regimen to replenish the stock of available antimalarials, thereby reducing the scourge of disease due to resistant parasites, there is a further need for deconvoluting the various factors contributing to the diversity in disease severity or susceptibility among endemic populations. One of such potential factors is CD14, a leucine-rich repeat surface protein linked to macrophage TLRs that functions as a receptor for bacteria lipopolysaccharide and microbial ligand recognition,14,18 as well as host response to viral and fungal infections.36,37 Its polymorphism has been shown to be a major regulator of innate immune response,
10
and its demonstrated expression in the brain is a critical factor in the development of cerebral malaria.
9
Significantly, it has been shown that CD14 genetic factors that affect differential expression of downstream genes would potentially have functional imperative in the immune response to infectious pathogens or immunological diseases.
10
To this end and as part of our ongoing work on immune regulatory patterns contributing to disease diversity and severity, we set out to determine the role, if any, of the genetic diversity of CD14 gene promoter polymorphism (snp −159 C/T; rs2519190) on markers of clinical disease or severity among and between
Even though there are a number of publications referencing CD14 and its regulatory or contributory role in several conditions and diseases, including allergy, just a handful is directly related to malaria. A signature publication on murine cerebral malaria showed that deletion of the CD14 gene confers protection against disease and a negative effect on parasite growth in transgenic mice challenged with
Oakley et al also reported on the beneficial role of CD14 in regulating parasitemia among CD14 transgenic and wild-type
In addition, although Wenisch et al 25 reported that they found no relationship between soluble CD14 and markers of disease severity in malaria infection, our present result states otherwise, based on genetic diversity analysis. Our results qualitatively delineate the potential reasons for the results obtained by Wenisch et al. This is an excellent study that should move this field further, especially among malaria-infected individuals from differing ethnic or endemic backgrounds or while examining severity of infection among nonimmune travelers from nonendemic regions. We recommend further studies with larger sample sizes and different population groups across sub-Saharan Africa.
The results in this study further confirm reports that CD14 polymorphism potentiates a TH1 differentiation and subsequent IFN-γ response leading to reduced malaria parasitemia. We believe that a variant CD14 would drive a deficient response leading to an immune cascade that could potentially make disease worse, leading to complications, compared with a normal immune response, in which infection can be adequately managed or controlled. Such complications could potentially include high-circulating immune complex levels,
1
extensive parasite diversity,
39
incomplete clearance of parasites posttreatment and potential for recrudescence,
2
or possibly development of drug-resistant strains. To this end, it is imperative that our results be extended by analyzing pro-inflammatory biomarkers in serum assays from similar patient groups as well as patients with cerebral or severe malaria, in addition to clarifying any contributory role TLR polymorphisms may play in regulating disease markers. In addition, we advocate strengthening awareness and understanding of CD14 regulatory effects on innate immune response, especially with malaria infection in sub-Saharan Africa, and potential to exacerbate disease outcome among infected individuals carrying the CD14 promoter gene mutant variant. We conclude that
Footnotes
Acknowledgements
The authors appreciate Maria Gazouli, University of Athens, Greece, and Iman Farid of Rochester Institute of Technology, Rochester, NY, USA, for technical assistance.
Peer review:
Three peer reviewers contributed to the peer review report. Reviewers’ reports totaled 1271 words, excluding any confidential comments to the academic editor.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was supported by the funding provided by a Research Laboratory Support and Faculty Development Awards, College of Health Sciences and Technology, Rochester Institute of Technology and an American Association of Immunology Undergraduate Faculty Travel Grant to present this work at the annual conference was provided (B.N.T.). Y.L. was supported by China Natural Science Foundation grants #31501002 and 31470070.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
BNT conceived, designed experiments, and drafted the manuscript. RIF, COF, and OO recruited patients; obtained consent; and collected blood samples. TJS, NA, and BNT performed genotyping and endonuclease digestion. YL, RIF, COF, OO, and BNT conducted statistical analyses. OO and COF contributed to scientific content. All authors read and approved the final version of the manuscript.
