Abstract
Amphibian defensive skin secretions and reptile venoms are rich sources of bioactive peptides with potential pharmacological/pharmaceutical applications. As amphibian and reptile populations are in rapid global decline, our research group has been developing analytical methods that permit generation of robust molecular data from non-invasive skin secretion samples and venom samples. While previously we have demonstrated that parallel proteome and venom gland transcriptome analyses can be performed on such samples, here we report the presence of DNA that facilitates the more widely-used applications of gene sequencing, such as molecular phylogenetics, in a non-invasive manner that circumvents specimen sacrifice. From this “surrogate” tissue, we acquired partial 12S and 16S rRNA gene sequences that are presented for illustration purposes. Thus from a single sample of amphibian skin secretion and reptile venom, robust and complementary proteome, transcriptome and genome data can be generated for applications in diverse scientific disciplines.
Introduction
On a global scale, amphibian and reptile populations have suffered widespread declines and extinctions in recent decades [1,2]. While some declines are clearly due to factors including ultraviolet radiation, predation, habitat modification, environmental acidity and toxicants, disease, changes in climate or weather patterns and interactions among these factors, in many others, the reasons are not clear [3,4,5,6]. As part of the overall “biodiversity crisis”, recent reports suggest that many amphibians and reptiles are undergoing range reduction and extinction, such as the Costa Rican golden toad
Aside from the aesthetic value and functional roles these animals serve in their environs, they may offer properties of benefit to human welfare, such as bioactive peptides. The structural diversity of polypeptides in amphibian skin granular gland and reptile venom gland probably reflects a plethora of different roles, either in the regulation of physiological functions, defence against predators (amphibians) or in prey submission and capture (reptiles) [11,12,13,14]. These venom and granular gland secretions are released following attack and stress (amphibians) or in striking prey (reptiles) and, in addition to polypeptides, they also contain various components (proteins, biogenic amines) resulting in a wide spectrum of bioactivities [15,16,17,18]. Despite intensive study of frog skin peptides and lizard peptides for several decades, the vast majority remain uncharacterized and the vast majority of amphibian species remain to be studied. The extraordinary complexity and diversity of frog skin and lizard venom peptides renders intriguing resources for novel drug lead discovery–-a fact that is particularly relevant at present as interest in peptides as therapeutics undergoes a renaissance in the pharmaceutical industry [19,20]. In the past, the study of frog skin secretions and lizard venoms necessitated the sacrifice of the specimens and extraction of the dissected skin or venom gland, respectively. This procedure was highly destructive and often required several tens, hundreds or even thousands of specimens and the choice of chemical extraction medium was highly selective in terms of component solubility [15,21]. The advent of the non-invasive mild transdermal electrical stimulation technique for frog skin secretion [22] and a modified gentle acquisition technique for lizard venom [23,24], obviated the need for endangered specimen sacrifice and produced a more denned and molecularly complete product for proteomic analysis. However, molecular studies related to cloning of messenger RNAs encoding bioactive peptides required sacrifice of specimens and library construction from excised tissues [25,26,27,28]. Recently, our research group has reported the presence of granular gland and venom gland transcriptomes in these secretions [29,30]. Consistent with contemporary biological science ideology, we have further examined and modified the technologies employed in the study of these secretion samples.
Here we describe the identification and isolation of genomic DNA from such with original data presented on the partial structure of 12S and 16S ribosomal RNA genes from the skin secretion of the Chinese toad
Materials and Methods
Acquisition of toad and lizard venom
Two adult specimens of large-webbed bell toads,
Adult captive specimens of the Mexican beaded lizard (Heloderma horridum) were housed in the Arizona-Sonora Desert Museum. After properly restraining the animal [23], venom was collected by inserting a plastic reinforced rubber hose into the mouth and then collecting venom in a glass beaker as it was expressed from the glands into the oral cavity via the bite pressure of the lizard [24]. The beaker was then chilled and subsequently snap-frozen with liquid nitrogen prior to lyophilization.
Both techniques mentioned above caused no harm and minimal stress to the animals. As
DNA amplification and sequencing
Total genomic DNA for PCR was extracted from twenty milligrams of lyophilized toad venom using a Wizard® genomic DNA purification kit (Promega Corporation, U.S.A).
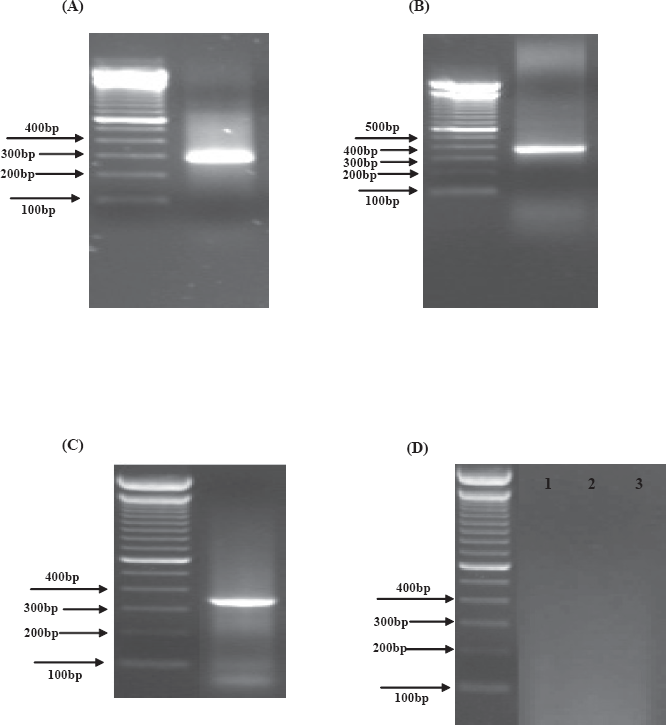
Gel electropherograms of PCR products generated using primers to (
Total genomic DNA for PCR was extracted from twenty milligrams of lyophilized lizard venom using a Wizard® genomic DNA purification kit (Promega Corporation, U.S.A). The
Results
Approximately 50–250 ng of genomic DNA was present in a PCR appropriate format in amphibian and reptile lyophilized venoms. The novel nucleotide sequences of a 315-bp segment of the 12S rRNA gene and a 379-bp segment of the 16S rRNA gene were determined by the PCR-direct sequencing method from the venom-derived library of
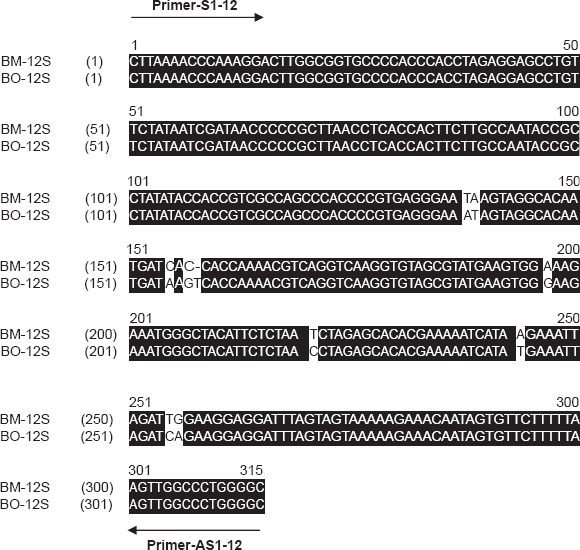
Nucleotide sequence alignments of Bombina maxima 12S rRNA (BM-12S) and Bombina orientalis 12S rRNA (B0-12S). The AlignX programme of the Vector NTI suite was employed. The sense (S) and antisense (AS) primer binding sites are indicated.

Nucleotide sequence alignments of
From
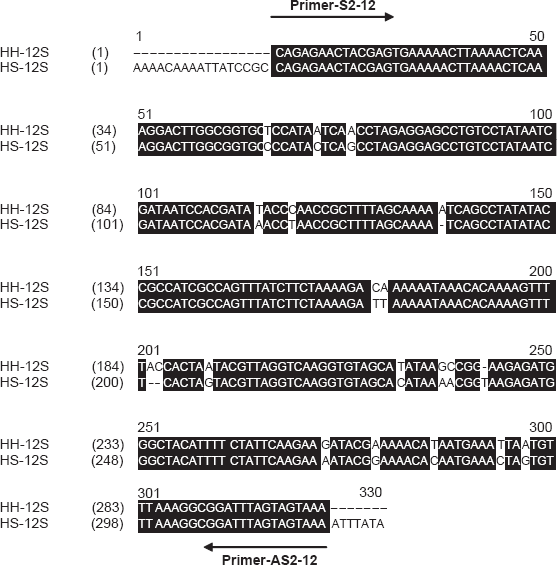
Nucleotide sequence alignments of Heloderma horridum 12S rRNA (HH-12S) and Heloderma suspectum 12S rRNA (HS-12S). The AlignX programme of the Vector NTI suite was employed. The sense (S) and antisense (AS) primer binding sites are indicated.
Discussion
Molecular cloning of bioactive polypeptide mRNAs from amphibian skin or reptile venom gland provides much useful information on endogenous propolypeptide convertase specificities, co-encoded peptides and can be a vital step in the initiation of studies designed to map genomic organization of respective genes [16,20]. Until now, whilst isolation and structural characterization of proteins/peptides could be achieved using lyophilized venom, construction of transcriptome cDNA and genomic DNA libraries for the purpose of molecular cloning of precursors and gene mapping, respectively, required the use of tissue samples that necessitated sacrifice of living specimens [25,26,27,28]. Amphibians and reptiles are two groups of vertebrates that are suffering global declines due to a combination of factors, including habitat destruction and pollution on one hand and pathogenic viral and fungal diseases on the other [1,2]. These factors however, do not explain all observed population crashes [6,32,33]. The acquisition of a wide range of amphibian/reptile species for venom studies is not trivial and every effort has to be made to address the well-being of rare captive specimens. We were thus faced with the problem of obtaining structural information on skin/venom gland peptide cDNAs and genomic DNA without recourse to specimen sacrifice. To achieve this goal, we attempted to obtain this information by a novel and highly speculative route–-cloning from the skin secretion or venom itself–-essentially using the secretion or venom as a “surrogate” tissue for this purpose.
In initial studies performed on several different species, we unambiguously demonstrated that the skin or venom gland transcriptomes are present in stimulated defensive secretions or venoms as evidenced by the successful cloning and sequencing of an array of full-length polyadenylated mRNAs encoding venom gland secretion peptides [29,30].
In this study, we focused our attention to determine if genomic DNA was present in a PCR appropriate format in such lyophilized venoms. It was indeed found to be and approximately 50-250 ng of genomic DNA was present. We successfully obtained partial but expected (from primers employed) DNA nucleotide sequence of the mitochondrial gene for 12S rRNA and 16S rRNA from a single lyophilized sample of amphibian skin secretion and reptile venom. The skin secretion of large-webbed bell toads
Conclusions
To our knowledge, this study is the first reported where a single sample of amphibian skin secretion and reptile venom has been shown to contain elements of the animal's genome that when amplified and sequenced, can provide useful information for a variety of applications such as in molecular taxonomy. This is of course, in addition to the global venom gland proteome and transcriptome. Such a finding offers a rapid, non-invasive and non-lethal approach to obtaining molecular genetic data in a manner that does not compromise scientific robustness and that additionally, offers the possibility of performing and serially-repeating such studies on the same individual specimens. The development of this technique and hopefully its adoption by researchers in the field, is a significant contribution to the evolving ethical framework regarding the usage of live animals for scientific research, especially those that are representative of threatened biodiversity. Thus the application of technologies such as described in the present study should preserve biodiversity for future biotechnological exploitation and systematic studies.
Footnotes
There is no conflict.
Acknowledgements
Hang Fai Kwok was in receipt of a Vice Chancellor's Overseas Research Scholarships from the University of Ulster during his PhD studies. We also thank the herpetologists of the Arizona-Sonora Desert Museum for acquisition of venom from the Gila monster
