Abstract
Micro ribonucleic acids (miRNAs) represent a class of small noncoding RNAs that play important roles in multiple biological processes by degrading targeted mRNAs or by repressing mRNA translation. In the case of algal lineages, especially dinoflagellates, knowledge regarding the miRNA system is still limited and its regulatory role remains unclear. In the present study, a computational approach was employed to screen miRNAs from the expressed sequence tags (ESTs) of
Introduction
Micro ribonucleic acids (miRNAs) are small (≈19–23 nt) noncoding RNAs that are abundant in most plant and animal species. 1 miRNAs have been shown to post-transcriptionally regulate the genes affecting many different biological processes.2–5 miRNAs are normally identified through direct sequencing of small RNA libraries with high-throughput sequencing methods. Moreover, many miRNAs from a broad range of species have been predicted by fast-developing computational approaches.6–8 Because most of the miRNA-predicting algorithms are mainly based on whole genome sequences, the prediction of miRNAs is limited to model species, such as wormwood and some bilaterian animals.9–11 Expressed sequence tags (ESTs) can be another resource for identifying miRNA precursors, owing to the fact that pre-miRNAs (precursors of miRNAs) always share a similar evolutionary conserved secondary hairpin structure. Several efficient pipelines have been developed to identify miRNAs in numerous species without available genome sequences, including porcine, peach, horsegram, and certain species of insects.9,12–15 Therefore, the computational approach using EST data has already become a widely-used method for miRNA prediction. Based on the survey of the miRNEST database, 16 9,980 miRNA candidates were predicted from ESTs of 420 species. 17
Recently, several studies have focused on miRNAs in protists taxa; their results indicated that miRNA machinery originated from these early-diverging eukaryotes.18,19 miRNAs have also been identified in some species of protists alga; for instance, 13 miRNAs have been identified from the diatom
Dinoflagellates represent a large and diverse eukaryotic flagellated microalgae group that constitutes important primary producers in marine and freshwater ecosystems.
25
Some dinoflagellate species, such as
Materials and Methods
Dataset of miRNAs and Alexandrium tamarense ESTs
All 21,643 previously known mature miRNAs were obtained from the miRNA Registry Database (release 18.0, November 2011).31,32 These miRNAs were defined as the reference miRNA dataset. A total of 10,885
Prediction of putative miRNAs and their precursors
The procedure used to search for putative miRNAs in the present study was modified from previously reported EST-based approaches,9,15 which is summarized in Figure 1. First, the PatScan software (Argonne National Laboratory, Argonne, IL, USA) was used to identify the matched patterns between all known mature miRNAs and the ESTs of
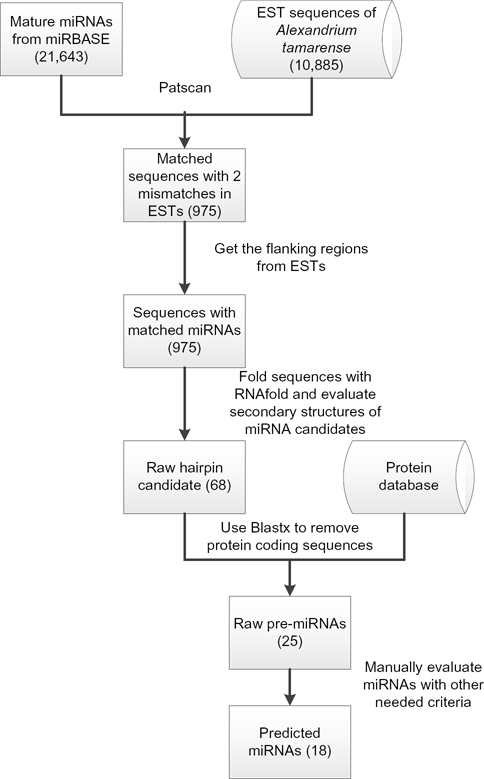
Schematic representation of the search procedures used to predict the miRNA candidates. The numbers in the parentheses indicate the sequence numbers for each step.
Three parameters, minimum fold energies (MFEs), minimal free energy indices (MFEIs), and G+C content were employed to distinguish miRNAs from other types of coding and noncoding RNAs. MFEs of all candidate miRNA precursors were generated by the MFOLD 3.5 program, and the MFEIs were calculated by the following equation:

Candidate miRNAs were considered as potential miRNA only if they fit the following criteria: 1) an RNA sequence could fold into an appropriate stem-loop hairpin secondary structure; 2) a mature miRNA sequence site was present in one arm of the hairpin structure; 3) a mature miRNA sequence had less than six mismatches with the opposite miRNA* sequence (miRNA* refers to the small RNA processed from the hairpin arm opposite the mature miRNA) in the other arm; 4) no loops or breaks were found in miRNA* sequences; and 5) the predicted secondary structures had higher MFEIs and negative MFEs.
Target prediction for identified miRNAs
psRNA Target
36
was used to screen potential target genes to predict the possible function of identified potential miRNAs. The transcriptomes from two sequenced diatom species,
Results and Discussion
Identification of putative miRNAs from Alexandrium tamarense
In this study, a strategy using a combination of software and bioinformatics methods based on homology searching and secondary structure evaluation was employed to screen for potential miRNAs from
List of miRNAs predicted from ESTs of Alexandrium tamarense.

Presently, a total of 99 miRNAs have been identified from several algal lineages, including diatoms, brown algae, red algae, and green algae.20–23,38 The components of RNAi machinery have also been identified in some algal species,
24
suggesting the existence of an miRNA system in algae. However, the miRNA system does appear to be present in all algal groups. For example, homologs of all the key miRNA biogenesis components in the genome of
Characteristics of the identified putative miRNAs
The 18 putative miRNAs were classified into 12 families on the basis of sequence similarities. There were five members in the miR-4492 family, two members in the miR-4508 and miR-4266 families, and only one member in the nine other families (Table 1). It has been reported that dinoflagellates experienced tertiary endosymbiosis events during their evolution, as their nuclear genomes are the largest among those of eukaryotes and their genes are always arranged in tandem arrays. 25 Gene duplication is considered to be a major source of emergence of novel miRNA genes.39,40 Considering the complexity of the evolutionary history of dinoflagellates, it can be deduced that their multimember miRNA families could have arisen through gene duplication events.
The sequences of miRNAs predicted in
The length of
Characteristics of miRNA precursors of Alexandrium tamarense.

Notably, none of the predicted miRNAs from
Targets of the identified miRNAs and their putative roles
The functional importance of an miRNA is ultimately defined by its target genes and the regulation it exerts on the expression of these genes. In order to determine the probable roles of the putative miRNAs in the present study, target genes were predicted with the psRNATarget software (Table 3). As the genome information of dinoflagellates is still unavailable, the transcriptomic information from two diatom species (
Predicted targets for newly identified miRNAs in Alexandrium tamarense.

It has been reported that most of the miRNA target genes encoding transcription factors, signal transduction factors, and metabolic transporters are involved in the regulation of growth and development.15,45 Similar functional biases were also observed in this study. For example, ata-miR-1275 targeted a zinc finger gene in
Conclusion
Our preliminary results based on in silico analysis not only demonstrate the existence of an miRNA system in dinoflagellates, but they also provide clues for understanding the function and evolution of these miRNAs in algal lineages.
Footnotes
Acknowledgments
We are grateful to all the laboratory members for their technical advice and helpful discussions. We also thank three anonymous reviewers for their valuable suggestions.
Author Contributions
Conceived and designed the experiments: DG, LQ, LS. Analyzed the data: DG, LQ, ZH, QZ, JW. Wrote the first draft of the manuscript: DG, QG. Contributed to the writing of the manuscript: DG, LQ, QG, LS. Agree with manuscript results and conclusions: DG, LQ, ZH, QZ, JW, QG, LS. Jointly developed the structure and arguments for the paper: DG, LQ, ZH, QZ, JW, QG, LS. Made critical revisions and approved final version: DG, LQ, ZH, QZ, JW, QG, LS. All authors reviewed and approved of the final manuscript.
Disclosures and Ethics
As a requirement of publication the authors have provided signed confirmation of their compliance with ethical and legal obligations including but not limited to compliance with ICMJE authorship and competing interests guidelines, that the article is neither under consideration for publication nor published elsewhere, of their compliance with legal and ethical guidelines concerning human and animal research participants (if applicable), and that permission has been obtained for reproduction of any copyrighted material. This article was subject to blind, independent, expert peer review. The reviewers reported no competing interests.
