Abstract
In these studies, we tested the effectiveness of combinatorial gene delivery to CCR5-expressing cell lines and primary cells to enhance resistance to HIV-1 infection. The transgenes used were chosen both to decrease membrane CCR5 and to inhibit HIV-1 replication, and were delivered using Tag-deleted SV40-derived vectors. rSV40s are very effective in transducing primary T cells and T cell lines. SV(RNAiR5), SV(RevM10.AU1) and SV(RNAiR5/RevM10.AU1) respectively deliver a small interfering RNA (siRNA) against CCR5, RevM10, to which a C-terminal AU1 epitope was added and which inhibits HIV-1 Rev, or both transgenes together. We used human cell lines (SupT1 and SupT1/CCR5) and primary human monocyte-derived macrophages (MDMs). We assessed RevM10 expression by flow cytometry (FACS). After transduction, we confirmed expression of each transgene by flow cytometry (FACS). Simultaneous delivery of both transgenes in a bifunctional vector protected both SupT1/CCR5 cells and MDMs from R5 tropic HIV-1Ba-L, better than either the monofunctional vectors, SV(RNAiR5) and SV(RevM10.AU1) individually. The bifunctional vector also protected from X4-tropic HIV-1, comparably to SV(RevM10.AU1). SV(RNAiR5) did not protect from X4-tropic HIV-1. Thus, combining these two transgenes in one vector protected from X4-tropic HIV-1 and provided enhanced protection for CCR5-bearing cells from R5-tropic strains of HIV-1. Combinatorial genetic therapy, by targeting of more then one HIV-1 function, may provide effective inhibition of HIV-1 replication.
Keywords
Introduction
Combinatorial antiretroviral therapies, such as highly active antiretroviral therapy (HAART) have been highly effective in prolonging life, but at best they keep HIV-1 infection in check while they are being administered. Drug-resistant viral mutants and drug side effects may develop after prolonged therapy.
Additional therapeutic approaches are therefore desirable. Intracellular immunization by gene transfer may be a promising supplemental approach to managing HIV disease. As with multiagent pharmacotherapy, combination genetic therapy targeting different HIV-1 functions has the attraction of likely increasing efficiency and decreasing the probability and rapidity with which resistant mutants arise.
Combination anti-HIV-1 genetic therapies requires that both transgenes be effective individually in inhibiting HIV-1, plus a vector that delivers these transgenes efficiently. With the discovery of RNA interference (RNAi), a new and more powerful tool has been added to the growing arsenal of anti-HIV transgenes. RNAi is highly potent in post-transcriptional gene silencing.1,2 Because of the high mutation rate of HIV and the specificity of RNAi, most studies using RNAi to protect from HIV-1 target cellular factors that are important in the HIV-1 replicative cycle.3–7 By targeting host genes critical for viral entry and replication, more sustained antiviral effects may be achievable.
More than any other anti-HIV transgene, RevM10, a defective variant of HIV-1 Rev has been used in experimental and clinical studies to inhibit HIV-1.8–14 HIV-1 Rev is a chaperone that is needed for export of singly spliced and unspliced HIV-1 transcripts from the nucleus. RevM10 is a transdominant inhibitor of Rev: it binds these transcripts but fails to export them from the nucleus. RevM10 can inhibit HIV-1 replication in T cell lines, primary T cells10,11,15 and primary blood mononuclear cells (PBMC). 16
Considering these issues, in the following proof of concept studies we used a Tag-deleted SV40-derived (rSV40) bifunctional vector, SV(RNAiR5-RevM10. AU1). A SV40-based vector was used in these studies because rSV40s transduce all HIV-susceptible target cell types used here highly efficiently, without selection and permanently. Using this combinatorial construct, we demonstrate high efficiency transduction, efficient gene delivery to MDMs, simultaneous expression of RevM10 and down regulation of CCR5 and enhanced protection from HIV-1 infection.
Materials and Methods
Plasmids and viral expression constructs
HIV-1NL4-3 HIV-1Ba-L, HIV-1JR-CSF and HIV-1 ADA-M, the strains of HIV-1 used in these studies, were obtained from the NIH AIDS Reference Reagent Repository Program (NIH-ARRRP) and passaged and titered according to standard techniques.
Construction of recombinant SV40 derivative viruses (rSV40) for gene transfer has been described previously. 17 We used the following Tag-deleted SV-40-derived vectors: SV(RNAiR5), SV(RevM10. AU1) and SV(RNAiR5-RevM10.AU1). RNAiR5 is an interfering RNA against CCR5 (siRNA vs. CCR5) driven by the adenovirus VA1 pol III promoter. 6 Details of its construction are published. 6 It targets sequences conserved among human, simian and murine CCR5 transcripts. Specifically RNAiR5 recognizes bases #748–769 of human CCR5 mRNA. RevM10.AU1 is RevM10 to which a carboxyl terminal AU1 epitope tag has been added. Its expression is driven by the CMV-IEP from strain AD169 of human cytomegalovirus. 18 The bifunctional vector, SV(RNAiR5-RevM10.AU1) is shown in Figure 1. The control virus for these studies, SV(BUGT), carries the cDNA for human bilirubin-UDP-glucuronosyl-transferase (BUGT). 19 This control vector is comparable to mock-transduction, and functions as a comparable control for studies involving inhibiting HIV-1 replication with transgenes driven by either pol II or pol III promoters. 6
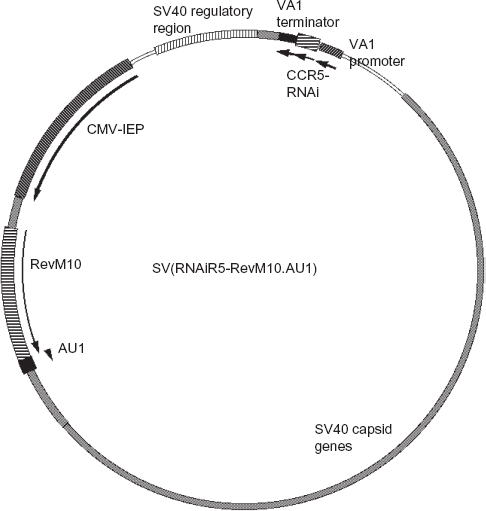
Map of SV(RNAiR5-RevM10.AU1). Key elements of the remaining SV40 genome are shown, as well as the orientation of the two transgenes. The SV40 regulatory region includes the SV40 origin of replication as well as the packaging sequences.
Production of rSV40 derivative virus
The techniques used to generate rSV40 virus from cloned rSV40 genomes have been described. 17 Briefly, virus genomes were excised from carrier plasmids, purified by agarose gel electrophoresis, recircularized and transfected into COS-7 cells, the packaging cell line used. No helper virus was used: COS-7 cells supply all necessary SV40 proteins, including Tag, in trans. Virus stocks were prepared as cell lysates, then band-purified as described elsewhere. 17 Resulting viruses were replication incompetent. Virus stocks were titered by Q-PCR. 20
Cell lines
COS-7 cells (ATCC) were maintained in Dulbecco's modified Eagle's medium (DMEM) with 2 mM L-glutamine adjusted to contain 1.5 g/l sodium bicarbonate, 4.5 g/l glucose, 1.0 mM sodium pyruvate, streptomycin (100 μg/ml), penicillin (200 U/ml), gentamicin (10 μg/ml), and 10% (vol/vol) newborn calf serum (NCS, Hyclone). SupT1 (NIH-ARRRP) and SupT1/CCR5 cells (kind gift of Dr. JA Hoxie, University of Pennsylvania) were grown in RPMI-1640 medium as described before. 6 All the cells were grown at 37 °C in humidified incubator with 5% CO2.
Primary cultures of monocyte-derived macrophages
PBMC were obtained by Lymphoprep (Sigma) density gradient centrifugation of leukocyte preparations from the TJU Blood Centre, from unindentified HIV-1 negative donors. Monocytes were enriched from PBMC fraction by 42.55% Percoll density gradient centrifugation, adherence to plastic overnight in RPMI 1640, supplemented with 10% heat-inactivated human AB serum that was negative for human immunodeficiency virus and hepatitis B antibodies (Fisher), and extensively washed without serum. Monocytes were cultured in RPMI 1640 supplemented with 10% heat-inactivated human AB serum and recombinant human granulocyto-macrophage and macrophage colony-stimulating factors (10 ng/ml each) (Sigma). Monocytes differentiated after 10 days into macrophages (MDM). Efficiency of differentiation was evaluated by FACS using antibody vs. CD14, and by morphological criteria.
Transduction
For transduction with SV40-derived viruses, SupT1 and SupT1/CCR5 cells were treated for 24 h at MOI of 10 in medium containing 2% NCS. This treatment was repeated twice, at MOI of 3, on sequential days. This approach, which we have used since initial studies of rSV40 gene delivery targeting HIV-’ functions. 21 and in many publications since then (reviewed, 22) produces effective transduction of virtually all the cells in a culture. While other approaches to transduction may be effective as well, we have used this approach because it is effective and to maintain consistency with and comparability to previously published studies. Primary cells, MDMs were transduced x 1, at MOI of 100. No selection was used. The rSV40-treated cells were then cultured for 5 days before further manipulation.
Antibodies
Antibodies specific for CCR5 were purchased from R&D systems Inc. FITC-labeled anti-AU1 antibody was obtained from Covance. PE labeled anti-CD14 antibody was purchased from BD Biosciences.
Flow cytometry analysis
For surface staining, cells were first treated with 10% normal mouse serum diluted in PBS for 30 min at 4 °C, then incubated with fluorophore-labeled antibodies diluted in PBS containing 1% BSA +0.1% sodium azide (FACS buffer) for 30 min at 4 °C. Cells were extensively washed using FACS buffer. For intracellular staining, cells were first permeabilized using PermFix reagent, following the manufacturer's recommendations (BD Pharmingen). Cells were blocked for 30 min at 4 °C in 10% normal mouse serum in PBS, then incubated with for 30 min at 4 °C with FITC-conjugated anti-AU1 antibody, diluted in FACS buffer. Cells+ were analyzed on Becton Dickinson FACSsort using Cellquest software.
Immunostaining
Cells were fixed and permeabilized with cold 100% ethanol for 10 min at 4 °C. Then, nonspecific signal was blocked with the normal 10% mouse serum for 30 min at room temperature. Anti-AU1 FITC labeled antibodies were added for 30 min at 4 °C. Slides were washed extensively and mounted.
HIV-1 infection assays
The HIV-1 susceptible T cells (SupT1 and SupT1/ CCR5) and MDMs transduced with SV(RNAiR5-RevM10.AU1), SV(RNAiR5), SV(RevM10.AU1) or SV(BUGT) were challenged with 100, 500 or 1000 IU (for MDM) of cell-free HIV-1 strains NL4-3, BaL, JR-CSF or ADA-M. After overnight incubation at 37 °C, cells were extensively washed with prewarmed (37 °C) PBS and maintained in growth medium. Every 2-3 days, the supernatant was changed and aliquots were taken for HIV-1 p24 analyses. HIV-1 p24 antigen levels in the supernatants were determined by ELISA (Zeptomatrix).
Quantitation of CCR5 mRNA in SupT1/ CCR5 cells and MDM by real-time reverse transcriptase PCR assay
Total RNA was extracted from SupT1/CCR5 cells and MDM, transduced with SV40-derived vectors by using standard protocols (Qiagen). Total RNA was subjected to reverse transcription using oligodT primers SuperScript III Reverse Transcriptase, following the manufacture's instructions (Invitrogen). Resulting cDNA was used as a template for real-time PCR amplification. Primers specific for CCR5 (forward: 5‘-GCCTTCTCCAGACAAACCAG-3’ and reverse: 5‘-GTTTTAGCCATCCCCCAAAT- 3’) (IDT, Coralville, IA) were used to determine trascript levels while GAPDH (forward: 5‘-CAACGAATTT-GGCTACAGCA-3’ and reverse: 5‘-AGGGGTC-TACATGGCAACTG-3’) primers were used as an internal control. The Stratagene detection system (Stratagene Mx3000P) was used for real-time PCR analysis. Thermal cycling conditions were: initial denaturation at 95 °C for 15 min followed by 40 cycles of 95 °C for 30 seconds, annealing at 55 °C for 60 s and extention at 72 °C for 30 s and 1 cycle of denaturation and annealing of 95 °C for 30 s and 55 °C for 60 s, respectively. At the end of each PCR run, data were automatically analyzed by the system and amplification plots were obtained. For each PCR, 1 μl of cDNA template was added to 24 μl of QuantiTest SYBR Green PCR Master Mix (Qiagen). To generate a CCR5 RNA standard curve to quantify CCR5 mRNA in SupT1/CCR5 cells and MDM, known amounts of the CCR5 cDNA standard were 10-fold serially diluted and then amplified in the same plate under identical conditions. The quantity of CCR5 mRNA in the samples was automatically calculated by the Stratagene Mx3000P system based on the data obtained from the standard curve. To control for the integrity of RNA and normalize CCR5 mRNA levels, GAPDH RNA was amplified using the same conditions. Reaction products were electrophoresed using 1% agarose gels and appropriate bands were visualized using UV light.
Results
Transgene expression following transduction with SV(RNAiR5-RevM10.AU1)
The effect of rSV40-derived siRNA on cell surface CCR5 and expression RevM10 were determined by flow cytometry and immunostaining. SupT1/ CCR5 cells, which stably express human CCR5, were transduced with SV(RNAiR5-RevM10.AU1). Control cells were mock-transduced. Expression of RevM10 was detected by FACS using antibody against the appended carboxyl terminal AU1-epitope, and was verified by fluorescence microscopy. Transduction efficiency, as assessed by flow cytometry, was >98% (Fig. 2). Analysis of three independent cytofluorimetry experiments showed 50%–60% reduction in cell surface CCR5 protein in rSV40-transduced SupT1/CCR5 (as illustrated in Fig. 2c).
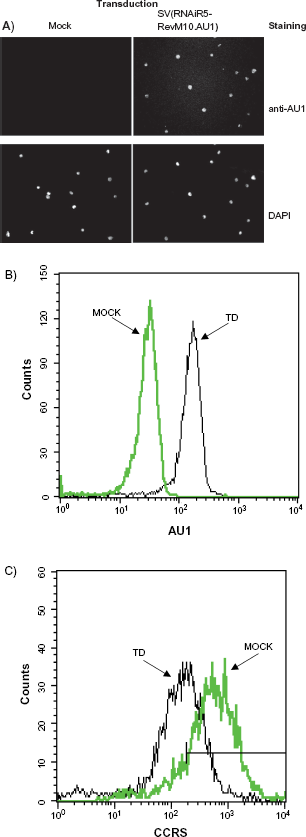
Transduction of SupT1/CCR5 cells using SV(RNAiR5-RevM10. AU1). SupT1/CCR5 cells were transduced with SV(RNAiR5-RevM10. AU1), or mock-transduced. rSV40-derivered transgenes expression were detected by immunostaining for AU1 A); by FACs analysis for the AU1 epitope appended to the carboxyl terminus of RevM10 B), and for cell membrane CCR5 C).
Monocyte-derived macrophages (MDMs) were transduced with SV(RNAiR5-RevM10.AU1). Control cells in these studies were mock-transduced. Expression of RevM10, was ascertained by FACS and fluorescence microscopy, and was comparable to that observed for SupT1/CCR5 cells (data not shown). Approximately 50% reduction in cell surface CCR5 was noted in SV(RNAiR5-RevM10.AU1)-transduced MDM (data not shown).
Decreased CCR5 mRNA in SupT1/CCR5 cells after transduction with SV(RNAiR5-RevM10.AU1)
To confirm that the down regulation of CCR5 coreceptor as seen by FACS analysis was due to reduced levels of CCR5 mRNA, SupT1/CCR5 cells and MDMs were transduced with SV(RNAiR5-RevM10. AU1), SV(RevM10.AU1), SV(RNAiR5) or (negative control) SV(BUGT). One week later, levels of CCR5 mRNA were analyzed by Q-RT-PCR. As an internal control, GAPDH mRNA was used. Cells transduced with SV(RNAiR5) or SV(RNAiR5-RevM10.AU1) showed considerably less CCR5 mRNA than either mock-transduced or cells transduced with SV(BUGT) or SV(RevM10.AU1) (illustrated for SupT1/CCR5 cells in Fig. 3 and quantified for both SupT1/CCR5 cells and MDMs in Table 1).
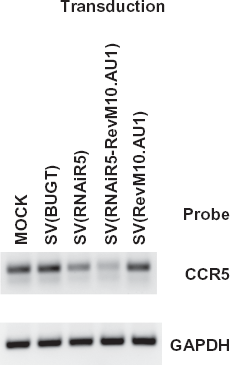
RT-PCR detection of CCR5 mRNA downregulation in SupT1/ CCR5 cells after transduction with SV40 vectors. SupT1/CCR5 cells were transduced with SV(RNAiR5-RevM10.AU1), SV(RNAiR5) or SV(RevM10. AU1). Controls were mock transduced SupT1/CCR5 cells and cells transduced with SV(BUGT). RT-PCR products were amplified to detect the coreceptor transcripts in SupT1/CCR5 cells in mock-transduced cells (lane 1); cells transduced with SV(BUGT) (lane 2); SV(RNAiR5) (lane 3); SV(RNAiR5-RevM10.AU1) (lane 4) and SV(RevM10.AU1) (lane 5).
Quantitation of CCR5 mRNA in SupT1/CCR5 and MDM cells transduced with SV40-derived vectors.
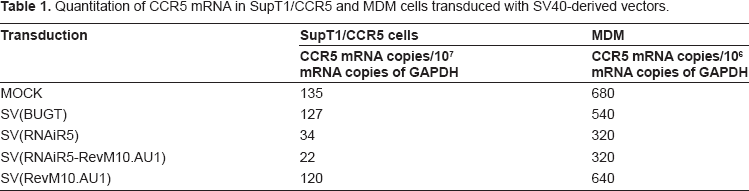

Slightly more effective targeting of CCR5 mRNA was seen using the bifunctional vector, compared to SV(RNAiR5) alone. These differences are within the variability of transgene effectiveness that is seen using SV(RNAiR5) alone. 6
Inhibition of HIV replication in SupT1 and SupT1/CCR5 cells
To test the efficacy of anti-CCR5 siRNA and RevM10 in protecting from R5- and X4-tropic HIV infection, SupT1 and SupT1/CCR5 cells were transduced with these several vectors: the bifunctional SV(RNAiR5-RevM10.AU1) or its monofunctional relatives, SV(RNAiR5) and SV(RevM10.AU1). Control cells were either transduced with SV(BUGT) or mock-transduced. Transgene expression was confirmed as above, and cells were challenged with the X4-tropic HIV-1NL4-3 and the R5-tropic HIV-1BaL.
SupT1 cells (X4 only) were protected from the X4-tropic strain, HIV-1NL4-3, virtually identically by SV(RevM10.AU1) and SV(RNAiR5-RevM10. AU1). As anticipated, CCR5 downregulation by SV(RNAiR5) alone did not protect SupT1 cells from this strain of HIV (Fig. 4a). By contrast, SupT1/CCR5 cells (bearing X4 and R5) were protected from HIV-1Ba-L (R5-tropic) by SV(RNAiR5), SV(RevM10) and the bifunctional vector (Fig. 4b). Protection by the latter was slightly greater than by either of the monofunctional vectors.
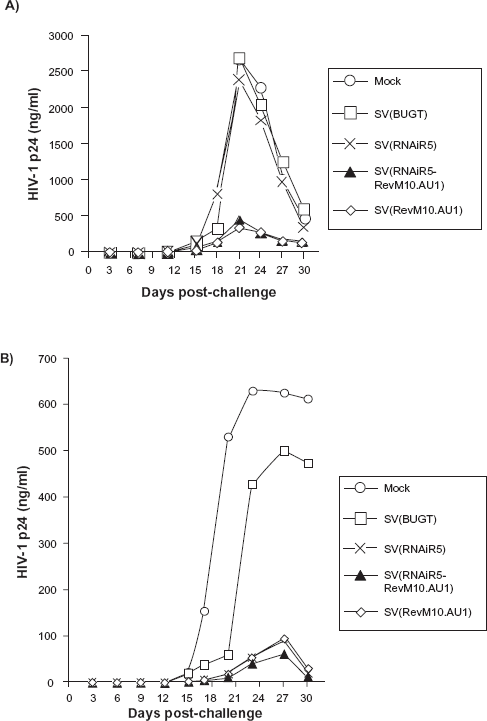
Inhibition of HIV replication in SupT1 and SupT1/CCR5 cells after transduction with SV40 vectors. SupT1 and SupT1/CCR5 cells were transduced with SV(RNAiR5-RevM10.AU1), SV(RNAiR5) or SV(RevM10.AU1). The same cell types transduced with SV(BUGT), or mock-transduced, were used as negative control populations. Five days after transduction SupT1 A) and SupT1/CCR5 B) cells were challenged with 100 infectious units of HIV-1 NL4-3 or 100 IU HIV-1 Ba-L respectively. HIV replication was measured by ELISA for p24 antigen in culture supernatants. results shown are representative of at least three independent experiments for each condition.
Inhibition of R5-tropic HIV-1 strains in MDMs by mono- and bifunctional rSV40s
Above, we determined that SV(RNAiR5-RevM10. AU1) could almost completely protect from both R5- and X4-tropic strains of HIV-1 at the challenge doses used, and that its level of protection was comparable at those challenge doses, to that afforded by either of the monofunctional vectors tested against the respective HIV-1 strains. We then sought to evaluate whether this bufunctional vector could protect primary MDMs. Further, we sought to to discriminate the protection provided by the combination of transgenes from that provided by each individually. Higher challenge doses were therefore used to overcome, at lest partially, the protection afforded by each transgene alone in order to visualize any augmentation of protection by combining the two transgenes in a single vector.
We therefore tested the effectiveness of SV(RNAiR5-RevM10.AU1) in protecting from HIV replication, compared to the individual monofunctional vectors, SV(RevM10.AU1) and SV(RNAiR5), in human MDMs. Primary MDM were transduced with SV(RNAiR5), SV(RevM10.AU1) or SV(RNAiR5-RevM10.AU1). Mock-transduction or transduction with SV(BUGT) were used as negative controls. Five days after transduction the MDM cells were challenged with 1000 IU HIV-1 Ba-L or 500 IU HIV-1 JR-CSF or ADA-M (Fig. 5).
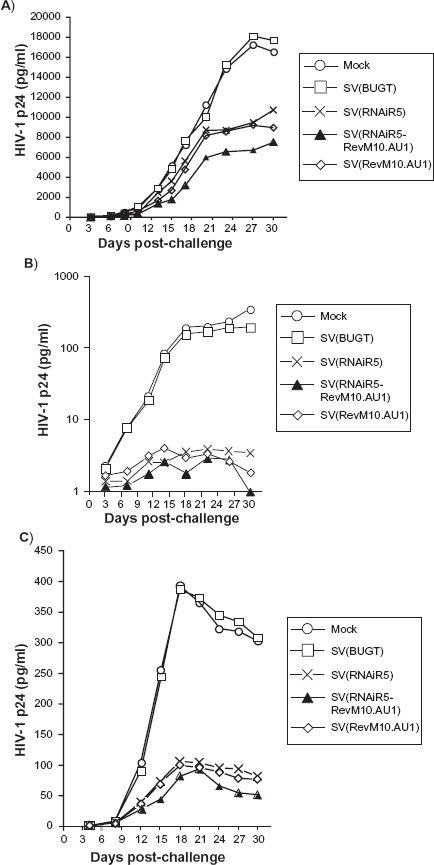
Inhibition of HIV replication in MDM cells after transduction with SV40 vectors. MDM cells were transduced with the same vectors as in Figure 4 Five days after transduction, cells were challenged with 1000 infectious units of HIV-1 Ba-L A) or 500 infection units of HIV-1 JR-CSF B) and HIV-1 ADA-M C). p24 antigen was measured in culture supernatants by ELISA. The results of these challenge assays on MDMs, shown here, are representative of at least three independent experiments involving MDMs from different donors.
In all cases, the bifunctional vector protected MDMs better than either monofunctional vector alone (Fig. 5). The magnitude of difference was most evident in MDM cultures challenged with HIV-1 Ba-L, the most virulent of the strains tested, and was least evident in MDM challenged with HIV-1 JRC-SF, which strain replicated least efficiently. The improved protection from HIV-1 ADA-M with bifunctional vector was intermediate, as was the virulence of this strain of HIV-1.
Discussion
Macrophages and T cells are major cellular targets for HIV-1 infection. 23 Macrophages represent a key reservoir for HIV and are a source of newly replicating HIV-1 throughout the infection.
Most clinically transmitted HIV-1 initially infects monocytes and their derivatives, macrophages and dendritic cells. However, over time virus tropism changes and T lymphocytes became its major targets. Any therapy for HIV infection should thus protect in both monocytes/macrophages and T lymphocytes. At the same time, the ability of HIV-1 to mutate and so escape therapies that solely target viral gene products suggests that cellular functions that are obligatory for HIV-1 replication should be considered attractive targets for therapeutic manipulations.
HIV-1 entry into human CD4+ cells requires the participation of certain cell membrane chemokine receptors that function as coreceptors for HIV. CCR5 and CXCR4 are two major coreceptors used by macrophage-tropic and T-cell-tropic HIV strains, respectively. 24 The CCR5 chemokine receptor is important for most clinical strains of HIV to establish infection and to disseminate throughout the body. It appears not to be essential to cell physiology, due to functional redundancies in the chemokine receptor family. 25 Individuals who lack functional CCR5 protein due to a homozygous 32 base-pair deletion (CCR5δ32) in the CCR5 gene are both apparently healthy and relatively resistant to HIV infection.26–31 Heterozygous individuals with approximately 50% less CCR5 surface expression tend to have lower plasma HIV viral loads and a generally prolonged course of disease. 32 In addition, once infected by HIV-1, their progression to AIDS is generally much slower than is the case for homozygous for wild-type CCR5. 33 Approximately 1%–2% of the Western European Caucasian population is homozygous for a 32-bp deletion in the coding region of the CCR5 gene, rendering them less susceptible to HIV infection. 34 Although one cannot be completely sure that decreasing CCR5 would be without consequences, these observations suggest that lowering CCR5 could help to protect high-risk individuals from HIV infection and/or slow disease progression once HIV infection is established. CCR5 is thus an attractive therapeutic target.
RNA interference is a highly specific mechanism of posttranscriptional gene silencing mediated by double-stranded siRNAs 19 to 24 nucleotides in length, that can be targeted to any mRNA of interest.1,2,35,36 Silencing occurs via a RNA-induced silencing complex (RISC)-associated RNase III-like endonuclease that cleaves the target homologous mRNA. A number of studies using siRNAs have targeted HIV genes as well as the cellular molecules critical for HIV entry, namely CD4, CXCR4 and CCR5.3,5–7,37
We approached antiretroviral genetic therapy by combining two transgenes, each targeting a different facet of HIV-1 replicative cycle. Most studies of multiple gene delivery have used combinations of multiple vectors, each carrying one transgene. 14 However, many vectors have been used to deliver polycistronic constructs.38–40 Recombinant gene transfer vectors derived from SV40 (rSV40s) are usually made by deleting large T antigen and are potentially of great utility in inhibiting HIV-1, since these vectors: a) transduce all types of lymphocytes, monocytes and their derivatives and CD34+ bone marrow cells very efficiently whether the cells are resting or dividing; b) integrate into cellular DNA to provide permanent transgene expression that does not change over time; and c) are transparent to immune system and so can be administered multiple times.14,41–43 rSV40 vectors are not detectably immunogenic in that neither does the rSV40 particle elicit neutralizing antibody responses nor do transduced cells elicit immune responses.
These studies were designed to test our ability to render CCR5-expressing cell lines and primary cells resistant to HIV infection by delivering transgenes designed to decrease membrane CCR5 and to express RevM10. RevM10 has been used in clinical trials in humans, and there has been no evidence of immunologic or other adverse reaction to the protein. As a nuclear protein, it is unlikely to elicit as strong an immune response as some cytoplasmic or cell membrane proteins might. Further, rSV40 vectors tend to produce proteins a lower levels than most other vectors, further decreasing the likelihood of immunogenicity.
In addition to RevM10, SV(RNAiR5-RevM10. AU1) carries a RNAi that targets CCR5. This vector was used to transduce CCR5-expresing SupT1 cells, as well as primary monocyte-derived macrophages. 44 In SupT1/CCR5 cells, SV(RNAiR5-RevM10.AU1) provided transgene expression in over 95% of cultured cells. Expression of RevM10 was evidenced by immunodetection of AU1, and decreased cell membrane CCR5, compared to control cells. In addition, simultaneous delivery of these transgenes in this combinatorial vector protected from HIV-1 replication: HIV-1 p24 levels in culture supernatants were decreased for both SupT1 and SupT1/CCR5 cell lines infected respectively with either X4-tropic HIVNL4-3 or R5-tropic HIV-1BaL. Since RevM10 is effective against both X4- and R5-tropic strains, all cell types should be protected from HIV by this vector. We found that combining these two transgenes in one vector provides a degree of enhanced protection for CCR5-bearing cells. The parallel curves of HIV-1 replication in protected and unprotected cells represent the nature of each individual HIV strain. Replication curves represent the sensitivity of these gene-modified (or control) cells to the challenge doses of the HIV-1 strain used. HIV-1 p24 production decreases when susceptible cells die. In unprotected cultures, very few viable cells remain at the end of the culture period. However, cultures expressing anti-HIV transgenes contain abundant cells at the end of the culture period.
Gene transfer to MDMs has been difficult to achieve using conventional vector systems because these cells are terminally differentiated and do not divide, and also because their phagocytic functions lead to degradation of phagocytosed vector particles. However, we have demonstrated both efficient gene transfer to MDMs using rSV40 vectors here and previously. 45 Unlike many other viruses, SV40 enters cells via lipid rafts. Virions are then transported unaltered to the nucleus via microtubules, bypassing the phagocytic process and so preserving viral particles intact.46–49 It may be for this reason that rSV40 vectors transduce MDMs so effectively. In this study, we also have shown that replication of R5-tropic HIV-1 strains Ba-L, JR-CSF or ADA-M was strongly inhibited in MDMs transduced with SV(RNAiR5-RevM10.AU1).
Effective augmentation of therapeutic efficacy by transduction with combinatorial construct requires that the several transgenes attack the disease process differently. To test this, an assay system must involve challenge doses that overcome the protection conferred by a single therapeutic agent, in order to assess the effectiveness of a combination by more rigorous criteria.
Levels of protection by the combinatorial construct varied among the several challenge strains, and better protection from R5-tropic HIV-1 was always seen with the bifunctional construct than with either transgene individually. Further, the level of protection from R5-tropic strains of HIV-1 generally exceeds the magnitude of CCR5 downregulation, an observation we have reported previously. 45 We described previously that multiple transgenes, delivered consecutively by different rSV40 vectors, confer increased protection against challenge with very high doses of HIV-1. Levels of protection afforded by these combinations varied, and could not necessarily be predicted from protection conferred by each transgene individually. The modest reduction of HIV-1 replication in MDM cells reflects the high challenge dose of HIV-1 (500 or 1000 infectious units) that was used for these experiments, and also the reduction of membrane CCR5 after treatment with rSV40s.
In order to evaluate the effectiveness of combinations of protective genes, it is necessary to challenge transduced cells with doses of HIV that overwhelm the protection provided by either gene individually. Hence, the high challenge doses of HIV-1 used here are necessary to evaluate the extent to which combinatorial gene therapy enhances protection compared to single agent therapy.
We have previously reported that the extent of protection from R5-tropic strains of HIV-1 is disproportionate to the downregulation of cell membrane CCR5 achieved. Thus, even modest reduction in plasma membrane CCR5 receptor density is accompanied by substantial protection from HIV-1 challenge. 34
Thus, targeting of more than one HIV-1 function simultaneously improves inhibition of HIV-1 replication. Combinations of transgenes directed against both cellular and HIV-1 gene products, such as we have used here, may provide high levels of protection from X4- and R5-tropic strains of HIV-1.
Disclosures
The authors report no conflicts of interest.
Footnotes
Acknowledgments
The authors acknowledge and appreciate the input and encouragement of colleagues, especially Drs. Scott Cairns, Harris Goldstein, Roger J. Pomerantz and the late Nava Sarver. These studies were supported by NIH grants MH70287, A141399 and A148244. This research has been approved by all institutional review committees.
