Abstract
The current work addresses the unification of Electronic Health Records related to cervical cancer into a single medical knowledge source, in the context of the EU-funded ASSIST research project. The project aims to facilitate the research for cervical precancer and cancer through a system that virtually unifies multiple patient record repositories, physically located in different medical centers/hospitals, thus, increasing flexibility by allowing the formation of study groups “on demand” and by recycling patient records in new studies. To this end, ASSIST uses semantic technologies to translate all medical entities (such as patient examination results, history, habits, genetic profile) and represent them in a common form, encoded in the ASSIST Cervical Cancer Ontology. The current paper presents the knowledge elicitation approach followed, towards the definition and representation of the disease's medical concepts and rules that constitute the basis for the ASSIST Cervical Cancer Ontology. The proposed approach constitutes a paradigm for semantic integration of heterogeneous clinical data that may be applicable to other biomedical application domains.
Introduction
Lack of uniformly structured data affects many areas of biomedical research, including drug discovery, systems biology, and individualized medicine, all of which rely heavily on integrating and interpreting data sets produced by different experimental methods at different levels of granularity, for which both the discovery and characterization is extremely difficult. 1 The heterogeneity and “isolation” of biomedical resources has been identified as the key research challenge, 2 towards the realization of translational medicine. 3
Genetic association studies constitute a significant scientific approach that may lead to a more comprehensive and holistic insight on the origin of complex diseases, 4 aiming to identify association between one or more genetic variants (e.g. polymorphisms) and a trait, which might be some quantitative characteristic, a discrete attribute, or a disease. 5 A genetic variant is genotyped in a population for which phenotypic information is available (such as disease occurrence, or a range of different trait values). If a correlation is observed between genotype and phenotype, there is an association between the variant and the disease or trait. 6
Although the number of studies elaborating on phenotype-genotype associations for common diseases is rapidly increasing, several studies show variation in the underlying association between genotype and outcome among the populations studied, resulting in questionable findings. It is evident that reliable association studies require large sets of patient phenotypic and genotypic data, all provided in a structured format. 7 Thus, methodologies for data and system interoperability, assisted by the appropriate technologies, will alleviate the problems mentioned above. While standardisation of phenotypic data and clinical practices seems unrealistic, semantic annotation and transformations to widely agreed classification schemes may provide workable solutions.
In this regard, Semantic Web technologies, 8 which are based on common formats that support aggregation and integration of data drawn from diverse sources, seem a key-enabler towards navigation and meaningful use of digital resources by employing automatic processes. The benefits promised by the Semantic Web include aggregation of heterogeneous data using explicit semantics, simplified annotation and sharing of findings, the expression of rich and well-defined models for data aggregation and search, easier reuse of data in unanticipated ways, and the application of logic to infer additional insights. 9 The potential of this technology endeavor has been particularly highlighted in biomedicine with initiatives such as the Semantic Web Health Care and Life Sciences Interest Group (HCLSIG) established by W3C (World Wide Web Consortium), which is chartered to explore and support the use of Semantic Web technologies to improve collaboration, research and development, and innovation in the information ecosystem of the health care and life science domains. Especially in biomedicine, it is envisioned that the use of Semantic Web technologies will improve the productivity of research, help raise the quality of health care, and enable scientists to formulate new hypotheses inspiring research based on clinical experiences.
Ontologies, considered as the backbone of the Semantic Web, play an important role in biomedical research through a variety of applications. 10 They provide the controlled vocabulary required for the annotation of biological datasets, the biomedical literature and patient records, facilitating the retrieval of and, more generally, access to information. Such standardization also facilitates the exchange of information and contributes to semantic interoperability among systems. By providing a representation of a domain, ontologies are also used in the mediation approach to integrating datasets. 11 Finally, many applications use ontologies as a source of computable domain knowledge, including natural language processing applications and decision support systems. 12 Ontologies are also critical to hypothesis generation and knowledge discovery in a data-driven approach to biomedical research. 13
The concept of supporting in a systematic way the conduction of multi-center association studies has been elaborated in ASSIST (Association Studies aSsisted by Inference and Semantic Technologies) by adopting inference and semantic technologies. 14 ASSIST is an EU-funded research project that aims to facilitate the design and execution of genetic association studies on cervical precancer and cancer (CxPCa). Toward this goal, ASSIST provides the means for seamless integration and virtual unification of distributed and heterogeneous CxPCa data repositories, as well as the underlying mechanisms to undertake the entire process of expressing and statistically evaluating medical hypotheses based on the collected data, in order to generate medically important associations. ASSIST aims to be used primarily by biomedical researchers for the identification of new markers of risk, diagnosis and prognosis, and possibly treatment of cervical precancer and early cancer via the implementation of case-control association studies.
Cervical cancer is an important cause of women mortality throughout the world. Current evidence suggests that specific subtypes of a certain virus, namely, the Human Papilloma Virus (HPV), cause genital infection that predisposes to cervical cancer development later in life. 15 Under research are other cofactors of cervical precancerous lesions. Tobacco use, number of sexual partners, number of deliveries, use of hormonal contraception, are characteristics that can put individuals at risk for CxPCa. 16 Moreover, genetic polymorphisms, such as Methy-leneTetraHydroFolate Reductase (MTHFR), p53 codon 72, Glutathione-S-Transferase (GST) T1, GST M1, and cytochrome P1 (CYP1) A1, are currently under investigation about their role in the pathogenesis of this disease.17–21
The current paper presents the knowledge elicitation approach followed towards the definition and representation of medical concepts related to diagnosis and disease severity that constitute the basis for the ASSIST CxPCa semantic model, emphasizing on the data unification procedure applied in three busy clinical sites across Europe. The paper is structured as follows. First, the origin and nature of the CxPCa data repositories considered in ASSIST are described. Then, the major CxPCa semantic medical concepts elaborated in the knowledge engineering phase of the project are presented. The common coding towards the semantic unification of CxPCa data repositories is then introduced, as far as the major clinical interventions related to CxPCa are concerned. The approach followed to infer the disease severity based on semantic rules associating the clinical findings of the abovementioned medical interventions is then elaborated, while aspects related to the implementation of the semantic model and an example usage scenario are presented. Finally, a discussion on related works as well as on the virtue of the proposed approach and its potential conclude the paper.
Materials and Methods
Cervical cancer data repositories
The ASSIST project aims to help medical researchers and clinicians to perform association studies with the use of information technologies (IT) within an open, integrated platform, allowing access to existing patient records from three different European clinics. More specifically, ASSIST currently integrates patient data originated from three gynecology clinics located in Belgium (Ghent University Hospital), Germany (Charité Hospital, Berlin) and Greece (Papageorgiou General Hospital, Thessaloniki). Each data repository has its own autonomous schema and captures patient information related to CxPCa, according to the local clinical procedures followed. An ethical committee approach was obtained as appropriately from each site of data collection before the utilization of patients' data. In addition, those data were anonymised prior to their availability via the ASSIST platform.
Personal data included in the databases were: date of birth, age at presentation, place of living, gravidity, parity, tobacco use (cigarettes per day and smoking years), use of oral contraceptives (years), use of intrauterine contraceptive device (years). Data regarding cervical HPV exposure included the age at the date of 1st HPV testing, whether HPV presence was detected in each visit, whether a low risk or a high risk HPV type was detected, as well as the kind of testing (e.g. PCR, hybrid capture, specific commercial kits, etc.). PAP smears were used for HPV genotyping. Cytology, colposcopy, and—when available—histology results were recorded (details given in the next paragraph). In cases where an intervention (i.e. punch biopsy of the cervix, large loop excision of transformation zone (LLETZ), conectomy or hysterectomy) was necessary, the kind, the date, and the result of the intervention were also available. All data are in form of discrete categorical values, i.e. no vital signal and image data are included.
Furthermore, genetic test results for polymorphisms possibly associated with CxPCa, i.e. p53 codon 72 polymorphisms (arginine/arginine omozygotes, arginine/prolin heterozygotes, prolin/prolin omozygotes), MTHFR polymorhisms (cytocine/cytocine omozygotes, cytocine/thymine heterozygotes, and thymine/thymine omozygotes), GSTT1, GSTM1, and CYP1A1 polymorphisms (wild type, heterozygotes, and null genotype), were also considered when available. For SNP analysis, either PAP smear or blood sample was used.
Data recruitment from the three different busy clinical sites across Europe, increased number of patients enrolled in the project, and virtual unification of these data, are all factors that increase the power and quality of association studies minimizing the limitations of the outcomes. At this point, records from 2,038 patients (more than 3,000 visits) have been made accessible to ASSIST by the three clinics. Using information from these medical records, research studies can be implemented to extract associations between CxPCa and viral, lifestyle and genetic factors aiming to advance knowledge on the origin and progress of the disease.
ASSIST virtually unifies these heterogeneous data sources syntactically, by defining a common data schema, and semantically via a domain ontology for precancerous and cancerous cervical lesions that encapsulates the appropriate medical knowledge to model the disease towards inferring its severity via the available diagnostic information (cytology, colposcopy and histology) and HPV infection results, as well as to express other disease-related information such as putative genetic polymorphisms associated with CxPCa, lifestyle and personal information. The main challenge has been to unify the three types of diagnostic results, coded according to locally used schemes, and infer disease severity expressed in a uniform and medically sound way. Thus, uniform coding has been elaborated for each type of diagnostic test result, so that the underlying heterogeneity in each clinic site is addressed. Interpretation to the limited existing standardised coding schemes is also supported when possible. This uniform classification of exam results is then employed in the context of medical rules for the inference part of the system, so as to identify the disease severity.
Cervical cancer semantic concepts
The typical approach for diagnosis and treatment of precancerous cervical lesions and/or invasive cancer includes anamnesis, routine clinical examination, smear test sampling (cytology), disease-specific clinical examination (colposcopy) and, if clinically necessary, biopsy sampling from suspicious for cancer sites (histology). Contemporary medical knowledge and clinical cost-effectiveness dictate that where cytology and colposcopy are normal, no further interventions are required. On the most of other occasions, especially in the presence of moderate or severe cytological and/or colposcopical abnormalities, histology is mandatory in order to exclude or to verify the disease. 22 Furthermore, HPV testing may be advised in certain cases and lifestyle related questions may be asked to the patient.
In current clinical practice there is no uniform way of describing cytology, colposcopy and histology findings. It was therefore realized that a need was eminent for common interpretation of similar conditions and diseases, and for common clinical protocols, if possible. The challenge of aggregating medical information lying in disperse databases, though not easy, could enable the reuse and exploitation of existing medical data for clinical and research purposes.
It has to be noted that no genetic factor has been established so far as to be a risk factor for the development of cervical neoplasia; hence, genetic testing is not part of clinical practice for the disease. However, genotype information on certain polymorphisms may exist in medical records as part of research.
ASSIST coding
Towards a common interpretation of the diagnostic test results, a scheme for uniform coding has been defined for the crucial fields of cytology, colposcopy and histology. These test results express the severity of cervical disease although they vary in the level of detail and type of information. ASSIST introduces a uniform coding scheme for all three tests consisting of 4 distinct classes (Table 1): the first one includes test results with normal findings, the second class indicates Low Grade Cervical Intraepithelial Neoplasia (LCIN), the third class indicates High Grade Cervical Intraepithelial Neoplasia (HCIN) and, finally, the fourth class corresponds to Invasive Cervical Cancer. The proposed coding scheme (from now on referred as “
ASSIST coding of data derived from screening for cervical cancer and/or from surveillance of women with precancerous lesions or cervical cancer.

ASSIST coding of data derived from screening for cervical cancer and/or from surveillance of women with precancerous lesions or cervical cancer.

In particular, regarding cytology results, there are two classification systems widely used, namely, the Bethesda and the Munich system (an extension of the Papanicolaou (PAP) classification), parallel to the detailed description of findings.23–27
Regarding colposcopy results,
Similarly, for histology results,
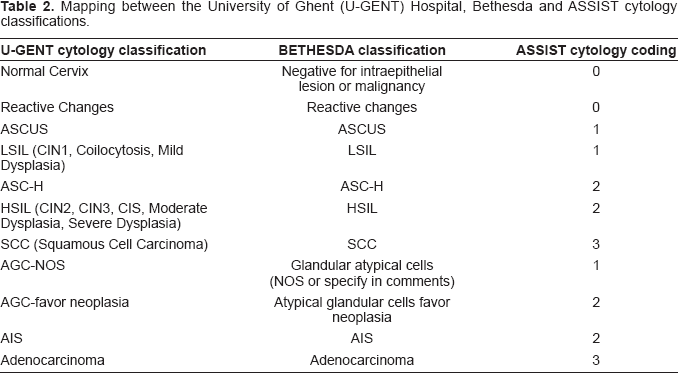
Since ASSIST coding consists of 4 classes, it poses limitations when mapping test results expressed with higher level of detail. Unfortunately, there is no standardization related to colposcopy and histology results, therefore, clinics tend to use locally defined coding schemes. However, cytological results may be described using the two established classification systems, namely, Bethesda and Munich, in which case they are preserved by ASSIST. Specifically, if cytological results coming from a specific clinic use either Bethesda or Munich classes (or a subset of them), then ASSIST will keep this classification, in addition to the ASSIST mapping. However, if cytological results are expressed in merged Bethesda or Munich classes, then the ASSIST classification will be used instead. As a result, it is optional to request and retrieve cytological results expressed uniformly either in Bethesda or in Munich (provided that the original records contain the necessary information) or in the ASSIST coding scheme. Tables 2 and 3 depict two examples of cytology results integration. In case 1, the University Hospital of Ghent uses a cytology classification with a subset of distinct Bethesda classes; cytology records are therefore integrated in ASSIST in two ways: i) preserving Bethesda classification and ii) mapped to ASSIST coding. In case 2, however, the Charité Hospital classifies cytology results in merged classes from both the Bethesda and Munich system. Specifically, although most classes are distinct Munich classes, the HSIL class used (which is a Bethesda term) cannot be mapped to a distinct Munich class. In this case, integration is performed using the ASSIST coding. Cytology results can always be mapped to the four distinct ASSIST codes that are related to clinical decision making during management of women with normal cervical screening results, precancerous lesions or invasive cervical cancer.
Mapping between the University of Ghent (U-GENT) Hospital, Bethesda and ASSIST cytology classifications.
Mapping between the Charité Hospital, Munich and ASSIST cytology classifications.

The amount of the information collected and expressed by using the ASSIST coding, is finally represented by an index, called “
In order to define the SI based on the results of cytology, colposcopy and histology, all these results have to be first unified and interpreted in the ASSIST coding. Every medical record is essentially associated to a patient
ASSIST medical rule and procedural logic for CxPCa Severity Index definition.

ASSIST medical rule and procedural logic for CxPCa Severity Index definition.
Severity Index distinction in 4 classes similar to ASSIST coding, appears to be clinically acceptable because it is closely related to major clinical outcomes and subsequent major clinical decisions. The SI is exclusively depended on the clinical results. Today, the status of a cervix is defined as: a) normal, b) with low grade lesion, c) with high grade lesion or d) with cancer. In this regard, the SI (from 0 to 3) directly corresponds to this stratification, and therefore is considered to be a tool enhanced by clinical significance.
Figure 1 depicts the overall data unification scheme employed in ASSIST that leads to the identification of the SI for each patient. The abovementioned semantic concepts and rules on CxPCa have been encoded formally and incorporated in the ASSIST Cervical Cancer Ontology that is described in the following.

Heterogeneity in biomedical data related to cervical diagnostic examinations in individual patients and their virtual unification.
ASSIST semantic model
The ASSIST Cervical Cancer Ontology, being the first one to ever semantically model CxPCa concepts, was designed and constructed following the medical knowledge elicitation phase described above. It encapsulates basic EHR (Electronic Health Record) information, examination results and genetic information together with the medical rules defined, so as to enable the mapping of diverse terms originated from each CxPCa data repository into a common schema and finally to infer the SI for CxPCa of each patient.
An interesting property of the ASSIST Cervical Cancer Ontology is that it provides great level of extensibility. The concept hierarchy is expanded from rather generic medical entities to specific CxPCa ones. This way, it can virtually incorporate any cancer domain knowledge and can even be further extended to other medical domains as well. This could provide potential means for representation of diseases and medical concepts and research on interrelation of diseases.
As noted in section 2.4, computation of SI is based in the notion of Case. It is, therefore, natural for the conceptual model that is built around the
Entities constituting the ASSIST Cervical Cancer Ontology are essentially of two types:
Figure 2 depicts the major concepts of the ASSIST Cervical Cancer Ontology, as well as their associations via appropriate properties.

Schematic representation of the basic concepts and their relations in the ASSIST Cervical Cancer Ontology.
As far as reasoning is concerned, this is based on two types of rules, expressed as Description Logics restrictions, 28 which produce inferred types for proper characterization of asserted knowledge. The first type of restrictions is basically employed to achieve the semantic unification of examination results in terms of the ASSIST Coding scheme. Figure 3 depicts the example of a restriction employed to infer that a cytology has indicated findings within normal limits, i.e. ASSIST cytology Code = 0, using the rules as defined in Tables 2 and 3. The second type of restrictions aims at the aggregation of unified examination results, to produce verdict on the SI of the corresponding case, as defined in Table 4. Patients categorized as having SI = 1 are inferred by employing the restriction depicted in Figure 4.

Example restriction rule employed to achieve the semantic unification of cytology results that correspond to findings within normal limits in terms of the ASSIST coding scheme.
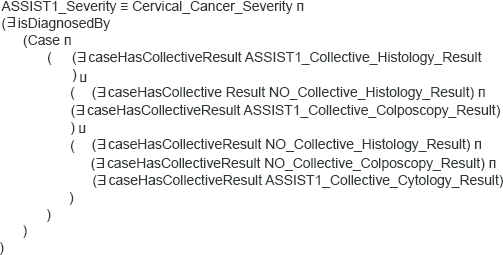
Example restriction rule employed to aggregate unified examination results so as to infer SI = 1 for the corresponding case.
The ASSIST Cervical Cancer Ontology was developed in Protégé knowledge modeling tool, 29 while the knowledge representation language employed was OWL-DL. 30 Sesame (http://www.openrdf.org/) and OWLIM 31 were chosen for storage and reasoning, respectively.
Furthermore, a simple medical-researcher oriented ontology editor was developed, in order to ensure easy and secure maintainability and incorporation of new knowledge. This tool permits easy rule insertion and concepts/properties manipulation requiring no knowledge engineering expertise, while assuring that vital concepts and interrelations remain unaffected.
The ASSIST Cervical Cancer Ontology TBox currently consists of 163 classes, 21 object properties, 33 datatype properties and 26 equivalent class axioms, while current instantiation of the ontology ABox contains about 60,000 individuals.
Example usage scenario
Suppose a researcher wishes to investigate whether there is an association between the MTHFR polymorphism and cervical precancerous lesions. While planning the study, he/she consults ASSIST to see, if there are shared data from patients with LCIN, HCIN and healthy patients for whom MTHFR genotype data are also available. The ASSIST User Interface offers the option to use the inferred ASSIST Cervical Cancer SI to express LCIN or HCIN, which is defined on the basis of the medical rules explained in section 2.4. Selecting a query to view all patients by inferred SI, the user can rapidly check the type of patients for whom data are shared through the ASSIST infrastructure (Fig. 5).

The user can express the severity of the disease through the results of cytology or colposcopy or histology or any combination of those. The results of those exams are expressed through the unified ASSIST coding. Using combined queries, the researcher asks for patients with genotypic data for the MTHFR polymorphism and for whom the staging of precancerous lesions is expressed through cytological results coded in the ASSIST unified cytology coding (Fig. 6). The graphical representation of the results indicates that the distribution of the MTHFR genotypes is different in the three stages of the disease. The user can continue with more queries involving additional patient parameters and further statistical tests on the extracted dataset. ASSIST offers a user friendly environment to express queries in a “language” easily understandable by the medical researcher and to also retrieve patient records that may use different internal coding schemes.

Lately, there is an increasing research interest in genetic association studies, emphasizing in quality aspects. The potential of reusing existing patient data can lead to high number of samples, thus, increasing power of the tests, while at the same time reducing the cost for the implementation of such studies. 32 Several projects are under development, aiming to provide integration tools that will allow accessibility and interpretation to a wealth of clinical records in disperse repositories. For example, i2b2 (https://www.i2b2.org/) is an NIH-funded research project developing a scalable IT framework that will bridge clinical research data and the vast data banks arising from basic science research, in order to better understand the genetic bases of complex diseases. The UK-founded CLEF project (http://www.clinical-escience.org/) focuses on generic methods for the capture, processing, and dissemination of information about cancer patients and their care using EHR data integrated with support for clinical and basic research in the biosciences. It aims to incorporate this information in formal knowledge management systems within a Grid framework. In the same fashion, the EU-funded NeuroWeb project (http://nuke.neurowebkc.eu/) aims to improve healthcare delivery related to neurosciences, achieving knowledge-based, personalized diagnosis and therapy through vertical integration of existing clinical and genetic databases. It stimulates the sharing of knowledge on cerebrovascular diseases using a Web-based platform. The Health-e-Child project (http://www.health-e-child.org/) aims at developing an integrated healthcare platform for paediatrics, providing seamless integration of traditional and emerging sources of biomedical information, primarily focusing on individualized disease prevention, screening, early diagnosis, therapy and follow-up of paediatric heart diseases, inflammatory diseases, and brain tumours. In most of these efforts, semantic technologies have been employed as an enabling technical endeavor for biomedical data integration and knowledge modeling. It has to be noted that, in such efforts, ethical and regulatory issues need also to be addressed in combination with privacy and security aspects.
ASSIST moves in a parallel course with the abovementioned research initiatives, aspiring that its platform will function as an IT tool enabling association studies linked with cervical cancer research by establishing a collaborative environment and allowing any medical group active in this area to use its facilities and/or contribute their own data/results. Cervical cancer, the second most common cancer in women, is recently subject of a tremendous interest by the scientists all over the world, especially because of the new knowledge concerning the causal role of HPV in the pathogenesis of the disease, and the new perspectives of disease prevention through prophylactic vaccination against viral infection. However, the interactions between HPV infection, environmental factors and host genetic background have not been yet fully elucidated and still reflect a field of continuing research interest. To this end, ASSIST aims to offer a new technological solution that will virtually unify multiple patient record repositories (physically located at different laboratories, clinics and/or hospitals) thus, enabling researchers to utilize existing genotypic and phenotypic patient data from several clinics and perform association studies in a low-cost and time-efficient way. Currently, a unified CxPCa data repository has been constructed, as a result of the ASSIST semantic integration methodology described above. With regards to genetic data, the current version of the ASSIST Cervical Cancer Ontology includes genotypic information of polymorphisms suspected to cause cervical cancer. Such data are currently available in research databases and can directly be exploited through the ASSIST platform and produce association study results. As genomic sequencing results will become available in the following years, it is evident that the ASSIST Cervical Cancer Ontology has to be expanded to incorporate such kind of information.
Following a generic design, the ASSIST system may be expanded in terms of its underlying knowledge model, in order to facilitate genetic association studies for other diseases, e.g. colon cancer and cardiovascular diseases. As presented in this paper, in ASSIST the main challenge has been to unify diagnostic results and severity index which are expression of disease severity. The presented approach constitutes a paradigm for semantic integration of heterogeneous clinical data that may be applicable to more diseases. The ultimate goal for ASSIST is to foster the biomedical research community by providing an open, integrated and collaborative framework to facilitate genetic association studies.
Disclosure
The authors report no conflicts of interest.
Footnotes
Acknowledgment
This work was supported in part by the IST-2004-027510 project, entitled “Association Studies Assisted by Inference and Semantic Technologies (ASSIST)”, funded by the Commission of the European Community (CEC).
