Abstract
Epistasis helps to explain how multiple single-nucleotide polymorphisms (SNPs) interact to cause disease. A variety of tools have been developed to detect epistasis. In this article, we explore the strengths and weaknesses of an information theory approach for detecting epistasis and compare it to the logistic regression approach through simulations. We consider several scenarios to simulate the involvement of SNPs in an epistasis network with respect to linkage disequilibrium patterns among them and the presence or absence of main and interaction effects. We conclude that the information theory approach more efficiently detects interaction effects when main effects are absent, whereas, in general, the logistic regression approach is appropriate in all scenarios but results in higher false positives. We compute epistasis networks for SNPs in the
Introduction
Genome-wide association studies (GWAS) are used to identify single-nucleotide polymorphisms (SNPs) associated with complex diseases such as cancer. 1 However, most GWAS analyze the main effects of SNPs. Epistasis is observed when the effect of an SNP is modified by other SNPs.2–4 Epistasis between SNPs helps to explain how multiple SNPs interact to cause disease. For example, epistasis between genes has been associated with hypertension, 5 sporadic breast cancer, 6 and several other diseases. 7 Epistasis also plays a subtle part in explaining missing heritability.8,9 Thus, identifying epistatic SNP interactions is of interest to better understand disease etiology. Furthermore, some studies suggest that, if the epistatic variance is larger than the additive variance, more power can be achieved to detect SNPs by searching for epistasis between SNPs rather than evaluating only the main effects. 10
A variety of tools have been used to detect epistasis, such as regression,11–14 Bayesian methods,15–20 and artificial intelligence algorithms.21–27 For higher order interactions, where regression methods are not suitable, several machine learning methods such as multifactor dimensionality reduction, 28 tree-based methods, 25 and entropy-based methods23,29 have been proposed, as they use classifiers and feature selection to reduce the computational burden.
In this article, we use simulations to explore the strengths and weaknesses of an information theory approach
29
for detecting epistasis compared to the logistic regression approach. We perform studies in which we simulate SNPs with and without the main effects. We also consider three types of interaction patterns and two types of linkage disequilibrium patterns. Finally, we demonstrate the practical application of these methods to identify an epistasis network. We use data from a head and neck cancer GWAS of the
Materials and Methods
We used a case-control study design to introduce the approaches to epistasis network analysis; however, the methods are also applicable to continuous phenotypes. The case-control status is denoted by a binary indicator Y, which takes the value of 1 or 0, corresponding to the categorization of the individual as being among the cases or the controls. The epistasis networks are networks in which the nodes are SNPs and the edges between the nodes correspond to the interaction between the SNPs. Hereafter, we define the two approaches for developing epistasis networks.
Information Theory Approach
For ease of presentation, we consider epistasis between two SNPs, A and B. Each SNP can have three possible genotypes: AA, Aa, and aa, which are coded as 0, 1, and 2, respectively, and where a is the minor allele. In the information theory approach, the association of the disease with an SNP or with the interaction between a pair of SNPs is quantified by assigning weights referred to as mutual information when a single SNP is studied and information gain when the interaction between SNPs is studied.
30
In the regression framework, these weights correspond to the respective odds ratios of the main or interaction effects. Specifically, mutual information between two variables provides a measure of the reduction in randomness in a variable when information about another variable is available. The mutual information of SNP A and the case-control status Y (the main effect of SNP A) is defined as



The mutual information
Given a pair of SNPs A and B, the information gain of A and B (interaction effect) is defined as

The information gain takes values between −1 and 1. A positive value indicates interactions that explain a part of the phenotypic variance; a zero value indicates interactions that do not explain any phenotypic variance; and a negative value indicates that modeling the interactions will be redundant because the information is already contained in the main effects (ie, modeling would possibly lead to multicollinearity). In this analysis, an information gain greater than zero was considered to represent a significant interaction.
Logistic Regression Approach
In standard logistic regression modeling, interactions between SNP A and SNP B are evaluated by testing the significance of β

We used Bonferroni correction to account for multiple comparisons. For an epistasis network of
Simulations
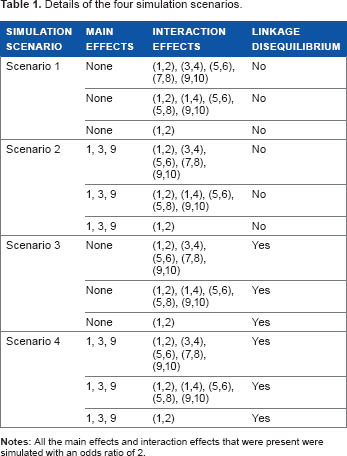
We performed simulation studies to investigate the performance of the methods. We considered several scenarios to simulate the SNPs involved in an epistasis network: scenarios with different linkage disequilibrium patterns and scenarios with presence or absence of main and interaction effects. In scenarios 1 and 2, all the SNPs were in linkage equilibrium, whereas in scenarios 3 and 4 the SNPs were in linkage disequilibrium. We used the linkage disequilibrium pattern of the 

Epistasis networks for the four scenarios simulated on the basis of network 1. (

Epistasis networks for the four scenarios simulated on the basis of network 2. (

Epistasis networks for the four scenarios simulated on the basis of network 3. (
Details of the four simulation scenarios.
Results
We analyzed the simulated data from the four simulation scenarios using the information theory approach and the logistic model approach as described previously. For each simulation scenario, the results are presented for the three interaction networks in Figures 1A, 2A, and 3A (referred to as networks 1, 2, and 3, respectively).
Simulation Scenario 1
In simulation scenario 1, the SNPs were in linkage equilibrium and had interacting effects, but no main effects. For the simulation based on network 1 (Fig. 1A), the information theory approach exactly identified all five interaction effects without any false positives (Fig. 1B). The logistic regression approach also identified all five simulated interaction effects; however, it also falsely identified several interactions that were not simulated (Fig. 1C). In the simulation using network 2 (Fig. 2A), which included two SNPs (SNP 5 and SNP 1) that were common in two independent interactions, the information theory approach identified only two of the five interactions simulated (Fig. 2B), whereas the logistic regression approach identified all five interactions; however, it also identified several false positive interactions (Fig. 2C). In the simulation using network 3 (Fig. 3A), which involved only the interaction between SNP 1 and SNP 2, the information theory approach and the logistic regression approach identified the true simulated interaction; however, both approaches also identified a few false positive interactions (Fig. 3B and C).
Simulation Scenario 2
In, simulation scenario 2, all the SNPs were in linkage equilibrium and had both interaction and main effects. For the simulation based on network 1 (Fig. 1A), the information theory approach identified only two of the five interaction effects that were simulated (Fig. 1D). In contrast, the logistic regression approach identified all the simulated interactions; however, it additionally identified several false positives (Fig. 1E). In the simulation using network 2, the information theory approach identified only two of the five interactions simulated (Fig. 2D), whereas the logistic regression approach identified the simulated interactions as well as several false positives (Fig. 2E). In the simulation with network 3, the information theory approach failed to identify the true simulated interaction and identified two false positive interactions (Fig. 3D). In contrast, the logistic regression approach identified the true simulated interaction (Fig. 3E); however, it also identified several false positive interactions.
Simulation Scenario 3
In simulation scenario 3, the SNPs were in linkage disequilibrium and had interaction effects, but no main effects. In the simulation using network 1, the information theory approach exactly identified all five interaction effects that were simulated (Fig. 1F), whereas the logistic regression approach identified interactions that were not simulated in addition to the simulated interactions (Fig. 1G). In the simulation using network 2, the information theory approach identified three of the five true simulated interactions, whereas the logistic regression approach identified several false positives in addition to the simulated interactions (Fig. 2F and G). In the simulation using network 3, both approaches identified the true simulated interaction without any false positives (Fig. 3F and G).
Simulation Scenario 4
In simulation scenario 4, the SNPs were in linkage disequilibrium and had both interaction effects and main effects. For the simulation based on network 1, the information theory approach identified only two of the five interaction effects that were simulated (Fig. 1H), whereas the logistic regression approach identified several false positives in addition to the five simulated interactions (Fig.1I). For the simulation based on network 2, the information theory approach identified only one of the five true simulated interactions, whereas the logistic regression approach identified several false positives in addition to the five simulated interactions (Fig. 2H and I). For the simulation based on network 3, the information theory approach failed to identify the true simulated interaction, whereas the logistic regression approach identified the true simulated interaction (Fig. 3H and I).
Head and Neck Cancer Data
We applied both approaches to data from a GWAS of head and neck cancer. The study participants were patients at The University of Texas MD Anderson Cancer Center (UT MD Anderson) with newly diagnosed, histologically confirmed, previously untreated head and neck cancer, including cancers of the oral cavity, pharynx, and larynx. The study genotyping was performed in two phases. The data from phase 1 included 2,696 individuals: 1,154 head and neck cancer patients and 1,542 controls. The data from phase 2 included 3,996 individuals: 1,031 cases and 2,965 controls. The institutional review board at UT MD Anderson approved the case-control study, and all participants provided written informed consent.
In this analysis, we developed epistasis networks for SNPs within the
We computed the epistasis network for the phase 1 data and used the phase 2 data to validate the epistasis network. The epistasis networks we developed for the phase 1 data by using the information theory approach and the logistic regression approach are shown in Figure 4A and B, respectively. The epistasis network based on the information theory approach identified the interaction between SNP

Epistasis network for the phase 1 head and neck cancer GWAS. (
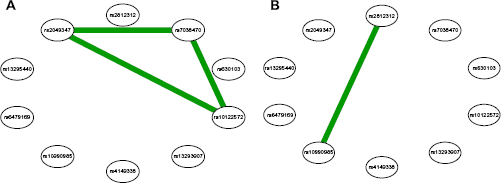
Epistasis network for the phase 2 head and neck cancer GWAS. (
Discussion
In this paper, we compare the information theory approach and the logistic regression approach for modeling epistasis networks. We used simulations to explore the strengths and weaknesses of the two approaches. We considered several simulation scenarios to simulate SNPs involved in an epistasis network with varying degrees of linkage disequilibrium patterns and the presence or absence of main and interaction effects.
The information theory approach accurately identified the epistasis network when there were no main effects. However, in the presence of only main effects, the interactions that included SNPs without main effects were not identifiable using this approach. In contrast, the logistic regression approach always included the true simulated interactions; however, it also included a higher number of false positives compared to the information theory approach. The higher number of false positives could be due to the fact that the logistic regression was performed using a single interaction at a time instead of including all the interactions in a single multivariable regression model. This would lead to model misspecification in the logistic regression framework. Importantly, covariates can be easily incorporated into the logistic regression approach, whereas inclusion of covariates is not straightforward in the information theory approach. The presence of SNPs in low linkage disequilibrium (
We applied the two approaches to develop epistasis networks for the head and neck cancer genetic data collected in two phases. The discrepancies between the logistic regression approach and the information theory approach were due to SNP
In summary, we have provided insights into the construction of epistasis networks using the information theory approach and the logistic regression approach. We concluded that the information theory approach more efficiently detects interaction effects when main effects are absent. In general, the logistic regression approach is appropriate in all scenarios but results in higher false positives. An understanding of the various strengths and weaknesses of these approaches provides insight for developing novel sophisticated methods to identify epistasis networks.
Author Contributions
Conceived and designed the experiments: RT, SS. Analyzed the data: RT, SS. Wrote the first draft of the manuscript: RT, SS. Contributed to the writing of the manuscript: RT, SS. Agree with manuscript results and conclusions: RT, SS. Jointly developed the structure and arguments for the paper: RT, SS. Made critical revisions and approved final version: SS. Both authors reviewed and approved of the final manuscript.
Footnotes
Acknowledgments
We thank Lee Ann Chastain for editing the manuscript. Some of the controls used for the analyses described in this manuscript were obtained from dbGaP at ![]() through dbGaP accession number phs000092.v1.p. Funding support for the Study of Addiction: Genetics and Environment (SAGE) was provided through the NIH Genes, Environment and Health Initiative (GEI) (U01 HG004422). SAGE is one of the genome-wide association studies funded as part of the Gene Environment Association Studies (GENEVA) under GEI. Assistance with phenotype harmonization and genotype cleaning, as well as with general study coordination, was provided by the GENEVA Coordinating Center (U01 HG004446). Assistance with data cleaning was provided by the National Center for Biotechnology Information. Support for collection of datasets and samples was provided by the Collaborative Study on the Genetics of Alcoholism (COGA; U10 AA008401), the Collaborative Genetic Study of Nicotine Dependence (COGEND; P01 CA089392), and the Family Study of Cocaine Dependence (FSCD; R01 DA013423). Funding support for genotyping, which was performed at the Johns Hopkins University Center for Inherited Disease Research, was provided by the NIH GEI (U01HG004438), the National Institute on Alcohol Abuse and Alcoholism, the National Institute on Drug Abuse, and the NIH contract “High-throughput genotyping for studying the genetic contributions to human disease” (HHSN268200782096C).
through dbGaP accession number phs000092.v1.p. Funding support for the Study of Addiction: Genetics and Environment (SAGE) was provided through the NIH Genes, Environment and Health Initiative (GEI) (U01 HG004422). SAGE is one of the genome-wide association studies funded as part of the Gene Environment Association Studies (GENEVA) under GEI. Assistance with phenotype harmonization and genotype cleaning, as well as with general study coordination, was provided by the GENEVA Coordinating Center (U01 HG004446). Assistance with data cleaning was provided by the National Center for Biotechnology Information. Support for collection of datasets and samples was provided by the Collaborative Study on the Genetics of Alcoholism (COGA; U10 AA008401), the Collaborative Genetic Study of Nicotine Dependence (COGEND; P01 CA089392), and the Family Study of Cocaine Dependence (FSCD; R01 DA013423). Funding support for genotyping, which was performed at the Johns Hopkins University Center for Inherited Disease Research, was provided by the NIH GEI (U01HG004438), the National Institute on Alcohol Abuse and Alcoholism, the National Institute on Drug Abuse, and the NIH contract “High-throughput genotyping for studying the genetic contributions to human disease” (HHSN268200782096C).
