Abstract
This work examines the feasibility and implementation of information service-orientated architecture (ISOA) on an emergent literature domain of human papillomavirus, head and neck cancer, and imaging. From this work, we examine the impact of cancer informatics and generate a full set of summarizing clinical pearls. Additionally, we describe how such an ISOA creates potential benefits in informatics education, enhancing utility for creating enduring digital content in this clinical domain.
Introduction
Worldwide, head and neck squamous cell carcinoma (HNSCC) is the sixth most common type of cancer. 1 As last reported in 2010, there is an estimated 600,000 cases of HNSCC reported each year around the globe. 2 Unfortunately, only 40–50% of these patients will survive more than five years. 3 Based on epidemiological projections, head and neck human papillomavirus-positive (HPV+) cancers are estimated to comprise over 25% of all known HNSCC. 4 These data suggest that approximately 150,000 new cases are affected by HPV+ head and neck cancer around the world each year. Currently, a subset of HNSCC, oropharyngeal squamous cell carcinoma, accounts for approximately 10% of all HNSCC worldwide. 2
In the United States alone, HPV+ cancers represent around 63% of all oropharyngeal cancers. 5 However, some reports also speculate that HPV and HNSCC could be as large as 90% in some demographics. 2 There are approximately 40,250 new cases of cancer of the oral cavity and pharynx diagnosed each year, as reported by the American Cancer Society in 2012. 6 Additionally, according to the North American Association of Central Cancer Registries and from data collected by the National Cancer Institute's SEER program and the CDC's National Cancer Registries during the 1999–2008 period, the incidence rates of HPV+ oropharyngeal cancer have increased dramatically by 4.4% per year, particularly among white males. 6
In recent publications, it has been postulated that the overall decreasing incidence of oral cavity/pharyngeal cancer may reflect reductions in smoking and alcohol consumption among all race/sex groups. But decreases in smoking-related HNSCC are offset by the trend of rising HPV+ cancers among other demographics, which may reflect changes in sexual practices since the mid-1960s, particularly among white males. 7 Mounting evidence suggests increased radiosensitivity of HPV+ HNSCC. 8 Because of the changing demographics of HPV+ HNSCC, some clinical investigators, including our group, have now proposed that it is feasible to lower radiation dose in patients with HPV+ cases.
A phase II trial is currently underway that seeks to identify patients with HPV+ oropharyngeal cancer. Early evidence suggests that patients with HPV-associated HNSCC may not require the full radiation dose given in a standard regimen of Intensity-Modulated Radiation Therapy (IMRT). 9 In this study, 46 patients had a complete clinical response with a lower-than-standard radiation dose, at 54 Gy, 9 with standard treatment doses being 66–70 Gy.
A reduction in radiation dose could potentially reduce late-term morbidity of radiation therapy in some cases and could change the standard of treatment for patients with HPV+ HNSCC.
8
Nevertheless, it is still too early to determine the correct dose to apply in radiation therapy or the appropriate imaging follow-up just from a single small study. Typical numbers for a phase II trial would be between 100 and 300 patients.
10
Because of the small number of participants in the ECOG trial (
Recognizing challenges for informatics for HPV and head and neck cancer
We are inspired by several targets that, despite the attempts of some evidence-based services, remain unmet in the status quo. 11 These unmet challenges in literature management and informatics include the failure to recognize an information need, physicians not being prepared for where a particular field may be heading and a preference for choosing the most convenient clinical paths rather than most appropriate choices. Users are further hindered by the excessive time and effort required to find answers to appropriate questions from resources, the difficulty of navigating overwhelming bodies of literature, and the inadequacy of current literature search technologies.
Our goal is to construct a literature informatics management education system (LIMES) for implementation in an academic teaching hospital. Unfortunately, organization and knowledge collection, such as what we are attempting to do with our LIMES, are unfortunately often relegated to “Cinderella” roles, assumed to be automatic to perform and often difficult to measure. Thus, we have chosen to focus on a small but emergent literature set, for which our case includes the use and role of imaging for HPV head and neck cancers, which is the primary interest and expertise of our institution. The proposed LIMES not only archives the articles, but also employs a newly recommended technique to systematically summarize and categorize content in a digital repository that is semantically mineable.
Another compounding challenge is the organization of information across different departmental clinical teams (eg, diagnostic radiology, radiation therapy and otorhinolaryngology). Such coordination can currently take years to establish communication between different departments. However, communication among different specialties could potentially benefit from informatics tools. The idea is to lay down what is considered as known and what is not known in the current published literature on HPV-HNSCC imaging and have multiple disciplines represented in the literature management process. Because of physician time constraints, we also seek to potentially share the burden of search, summarization, categorization, and organization with medical trainees.
Unfortunately, to our knowledge, the study/discussion of data collection by individuals is often not, if ever, described in published literature. Yet, the process of reading and reviewing in research, as well as learning how best to do these things, is often a very cumbersome process. For example, Ely has described how there appears to be a trend of increasing time demands on clinicians and less time to find literature resources. 12 Coumou and Meijman have described systematic methods by which primary care physicians search for answers to clinical questions. 13 They describe how physicians often first choose to consult colleagues whom they know well. 13 This practice of asking colleagues has not changed much in recent years, despite the proliferation of electronic publications made available worldwide.
Physicians in the past had limited options for finding answers, sometimes reduced to only the use of an intermediary person such as a librarian. 13 In this work, we propose, and have tested, a structured approach that generates clinical roadmaps and potential “pearls” from abstract information found in HPV head and neck cancer literature, using what we have called information service oriented architecture (ISOA). Essentially, we use an ISOA to break down the application functions of a large organization into a set of interoperable services. In our case, trainees, ie, medical school or premedical school students, will evaluate and structure the elements into our content management system (CMS) prior to review by attending physicians. We conjecture that this process will benefit trainee education.
Background on systems that reduce content with literature
Much of the focus on automated content extraction from literature management centers is on Natural Language Processing (NLP).14,15 NLP methods are most effective when they have a wide body of open literature to train on. In medicine, a successful NLP-based program was able to examine electronic clinical notes in women with early-stage incident breast cancers and identify whether and when recurrences were diagnosed. This was performed on clinical notes from 1,472 patients in which the NLP-based system correctly identified 92% of recurrences. 16 The area of medical informatics that implements NLP has strong roots in the domains of knowledge engineering and management. Knowledge engineering has been defined as the process of building tools for programing automation, including advances in machine learning, data analysis, rule induction, and analytic learning algorithms. 17
In our ISOA, our focus has been on collection from trainees and oversight by the attending physician. We have considered four primary concerns: (1) content management, (2) classification, (3) summarization, and (4) evidence-based tools. However, because of the numerous effort and ubiquity of proposed automated tools, we aim to incorporate the use of automated tools once they can achieve consistent results that approach 90% precision and recall. 18
CMSs are technologies that support the collection and management of published content. For literature, CMSs are used to store Adobe Systems Portable Document Format (PDF) files and potentially share this software. This includes both free and commercial packages such as Endnote, RefManager, Mendeley, Qiqqa, and Quosa among many more reference management software. 19 The first two, Endnote and RefManager, have roots in desktop software and are purchasable by those who have financial resources. Mendeley, Qiqqa, and Quosa are supported by premium options, such as space, sharing requirements, and/or are advertisement driven, and have a concern in the social sharing and citing of articles. All the above systems are superb for the management of content (PDFs) and highlighting. These tools are designed mainly for managing large libraries of resources, but currently do not have sufficient summarizing and classification/indexing tools within and across articles that we seek to achieve in our LIMES.
In informatics, classification can usually be referred to as indexing. 20 Medline indexers are employees who are required to have at least a bachelor's degree in a biomedical science. 21 A few university centers and the National Library of Medicine among others have put efforts into NLP tools for indexing. Hliaoutakis, in 2009, was able to demonstrate algorithmic improvements over the MMTX indexing package in speed and accuracy, using a C-value/NC-value method with the AMTex package. 22 However, both these software programs rely heavily on the publically available packages, such as Metamap, Medline and other important UMLS libraries. 15 Moreover, the NLM has been tasked to and continues to improve their own indexing tools 23 on a daily basis and continues to surpass many other indexing engines in the medical domain to our knowledge.
Text summarization (TS) methodology includes work by Luhn going as far back as 1958. 24 More recently, TS has been applied successfully to the medical field.23,25,26 Some additional current research in biomedical domains includes investigative work using abstractive-oriented and extractive paradigms. 26
Presently, evidence- based clinical services such as “UpTo-Date” are all that attending physicians have available to them for current evidence-based reviews. These services provide clinical topical reviews written by recognized physicians who provide evidence-based reviews with recommendations. These services are purchasable.
Improving and sharing information in the clinical environment is vital for enhancing patient care. There continues to be daily proliferation of new journal articles and Internet digests that describe emergent health issues. Unfortunately, physicians and researchers have limited ways to ensure that they remain up to speed on this changing information. In the past, this was strictly done by browsing abstracts. However, current methods of updating their knowledge include Google searching and/or the use of evidence-based services. All the three methods (browsing abstracts, Google searching, and evidence-based services) while having some benefits for searching also have some pitfalls.
We have provided a limited review of the literature on automation, as it in large part motivates our design. If one could create a mineable LIMES service, then other clinical investigators could build upon that work. However, we are focused on the ISOA as a method to provide a systematic support for literature management. Automation, while having the potential of extreme benefit, has only partial or limited efficiency in terms of language processing; extracting medical problems from narrative text still continues to evolve. Regardless, human experts still have to rank and rate automation services as well as provide “training” data even to evaluate the efficacy of automation. We believe that the development and use of this ISOA can assist in the education of our medical trainees.
Methods
HPV-HNSCC is an emerging field that was brought to our attention by several of our attending physicians. Dr. Alleman is the Bob G. Eaton Chair in Radiological Sciences and is boarded by both the American Board of Radiology and American Board of Otolaryngology as well as serves on several national Radiation Therapy Oncology Group (RTOG) trials. Dr. Chance Matthiesen is an Assistant Professor of Medicine and Radiation Oncologist at the University of Oklahoma Health Sciences Center and serves as a scientific reviewer for various national journals. He presently serves on the RTOG Head and Neck Steering Committee. Gynecological cancer is a prime concern in Oklahoma, and the University of Oklahoma has five board-certified gynecologic oncologists and currently in a National Cancer Institute endorsed clinical trial site.
To obtain the medical articles for our research, we performed a search in PubMed using “HPV head and neck cancer” as our search terms, which gave us 2187 articles. Because we wanted to focus on the imaging of HPV+ HNSCC, we added “imaging” to our original search criteria and ended up with an initial of 22 articles that matched our search. From these 22 articles, only two articles had to be discarded. One of these articles was not currently available in English and the other focused on HPV+ head and neck cancer in animals. The 20 remaining articles were used to initiate our research in the late 2013. Note, however, that as of May 2014, performing the same search now produces 37 results that could be potentially useful for a second study.
Approximate incidence of HPV+/- HNSCC/oropharyngeal cancer both worldwide and in the Us as estimated from publication reports. 2

Our first goal was to find appropriate trainees because they were essential to the task. Medical students are extremely busy studying for board examinations and other tests, and there is very little time to work on research projects. Our team felt that, as time was limited, the LIMES system would be easier to implement with less-formalized training and could be performed more on-the-job. For example, learning MESH terms and controlled vocabulary would likely go beyond the time commitments possible for physicians to manage and potentially for the trainees to perform. Our early estimate was a time commitment of about 10–20 hours per attending physician and about a 20–40 hours commitment for students to perform this work. Both may seem to be small numbers for researchers, but in reality had to fit into an academic hospital schedule of practicing clinical physicians.
For this project, we employed one medical student and one pre-medical student. We assessed their ability to understand a selected set of 20 medical articles about HPV-HNSCC imaging. To evaluate the performance of the ISOA, we developed a simple instrument, as shown in Table 2. We defined an “item” in each cell of a spreadsheet (it represents one block of text). Additionally, topic accuracy (multi) is when there are multiple topics for a single “item”.
Evaluation instrument used by physician mentors.

The trainees each created a chart for each of the 20 articles. Within each tab, the trainee placed information extracted from each article that they deemed to be of significance, as well as rewriting the text they extracted from the article into simpler terms and then categorizing this information using our Clinical Classification Schema (Fig. 1). The attending physicians “graded” the work of the trainees using these spreadsheets and the information provided in Table 2. This table represents the 12 specific questions the attending physicians used to “grade” the trainees’ work. The “items” that the table refers to are specific cells within the spreadsheets that relate to the 12 questions the attending physicians used for their grading.
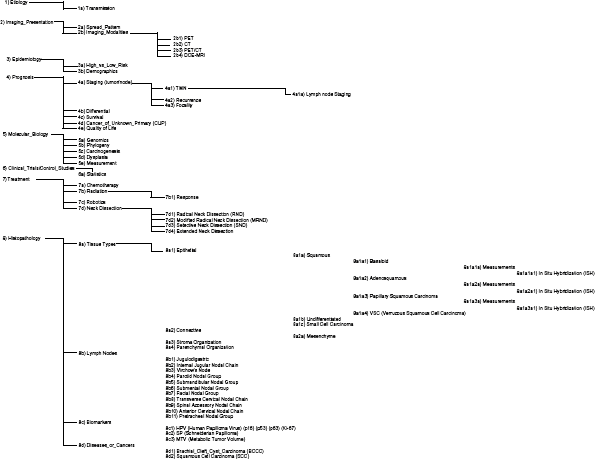
Our Clinical Classification Schema, which was used by trainees to classify the items they extracted from the 20 articles into the proper place.
The 12 questions are broken down into three different categories: (1) item-level questions, (2) topic-level questions, and (3) article-level questions. Item questions refer to the specific items trainees put into the cells. This could be the actual text that was extracted directly from the article or the condensed version of that same text.
Topic questions relate each row of text the trainee inputted on a classification level. The goal of this level was to classify the articles correctly. The attending physicians graded the trainees in the five classification areas: classification accuracy, sub-classification accuracy, multiple classification accuracy, classification introduction (adding a new classification to our Clinical Classification Schema), and classification educational value. The final category of questions is the article questions. These questions relate to each article, specifically determining their importance toward improving the knowledge of HPV-HNSCC literature and the educational benefit for trainees. Finally, for the entire project, attending physicians would then select the most relevant clinical pearls 27 among the abstracted text from both an instructional and those having informatics value. The goal of this work was to present clinical “pearls” that could be used to help future clinicians/researchers follow the field.
Results
Our clinical “Pearls” represent the data that were extracted from the 20 original articles and found to be important by the trainees. The attending physicians, when reviewing the articles, narrowed these “Pearls” into a smaller expert opinion “Pearl” list. This list is shown in Table 3. These “expert Pearls” represent key information that is deemed helpful by physicians for other physicians toward the demographics, role of imaging, and key therapeutic information.
Clinical pearls generated from this LIMES study.

Our experts’ gradings of the two trainees’ work is shown in Figure 2. As seen, our two trainees scored an average of 4 (top 20%) on the eight questions that were numerically based. Questions 8 and 9 were qualitative, not quantitative, so there was no statistical information to collect. Since questions 11 and 12 were answered once per article, their statistical data are not of value.
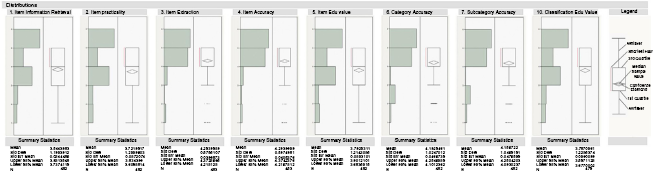
The means and standard deviations of the scores for the eight numerically represented questions. Note that these values are aggregated across all physicians and trainees.
Figure 3 illustrates trends from several of the questions, revealing visible relationships between trainees 1 and 2. We noted that there appeared on average some reasonable agreement at least on the item level; this seems to become much weaker at the classification level of questions (classification-level assessment appears to be less consistent than item-level assessment).
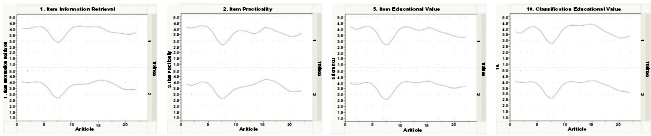
Selected trend values between the two trainees on item information retrieval, item practicality, item education value and classification education value.
Finally, because of the low number of trainees and physician mentors, only one of the instrument questions appeared to have a low statistical correlation value of 0.2254.

Item extraction (instrument question 3) illustrated a correlation value of 0.22564 on item extraction “grades” between the two physician experts.
Discussion
This work focuses on informatics by working with practicing academic clinicians (Drs. Alleman from Diagnostic Radiology and Matthiesen from Radiation Oncology) who are managing real medical trainees to curate and create a resource on HPV and head and neck cancer. This work is less concerned with the significantly more complex task of reducing the textual extraction to automation tools. Instead, we have created what we termed as an ISOA. This ISOA relies on a small group of early career professionals and trainees, overseen by senior attending physicians, to collect, manage, and organize this information. We believe that our ISOA is a feasible solution, as we have shown it to work successfully within the confines of a clinical environment. Additionally, it is mainly resourced by medical students and attending physician experts who do not require extensive technical training. However, based on the feedback from our attending physicians, we have already considered future steps.
Our physician reviewers provided these actual responses prior to the entire team finally assembling to reflect and summarize our results as a community. Dr. Alleman shared the following: the area of HPV and HNSCC and imaging “is a very dynamic body of literature that is diversified between radiology, head and neck, radiation therapy, and with ties to gynecological cancer, epidemiology, and even immunology. So, this is a diverse and dynamic body of literature that is difficult for a single individual to assimilate”. The main benefit, according to Dr. Alleman, is exposure and training for informatics provided to medical students and pre-medical students.
According to our Radiation Oncologist expert, Dr. Matthiesen, “The student summarizations were accurate and of use, but the selection of information to include in the summary needs work. However, the project showed the students often struggled to grasp the main concepts of many articles. Occasionally they appeared to get lost in the details of the manuscripts. Students also struggled to categorize information properly. I would attribute this likely to lack of general knowledge of the disease and lack of exposure. Because of this, more advanced radiation oncology residents or fellows might be better suited as test subjects in a future project for the categorization portion of the ISOA.”
In summary, we have concluded that initiating a repository with pre-medical students and medical students is feasible at the “item” level. Summary accuracy for each item is consistent and accurate enough to be useable by our attending physicians. However, more experienced students, or additional preparation or training, may be needed prior to the second phase. Perhaps, a second phase could include integration with medical residents/fellows who can also benefit from this exercise. In the long run, we hope to deploy an automated NLP system.
We sought to engage critical thinkers on the topic of HPV and head and neck cancer and prioritized efforts to recruit these physicians to participate in the mentorship of students. However, physician time is understandably focused on the direct care of patients, and this fact limited the participation of a large number of mentors in this project. We do believe that an ISOA could be designed to ft even the hectic schedules of physicians due to its simplicity and the large amounts of potential users. Additionally, we hope such a tool can better communication of information between different departments such as Radiation Oncology and the Radiology. This work has the potential to be extended not just at single institution, but also to provide a training method for future generations of medical professionals in this emerging issue. We are limited by our focus on emergent literature, and so may not necessarily provide a method that is targeted for research involving large numbers of publications, but we feel that it is timely and necessary to plan and implement a multicenter trial. Because of the low overall prevalence of HPV-HNSCC case studies, such tools may be helpful for greater efficiency of clinical trial design. Both physicians and our team agree that the creation of enduring content is one of the biggest benefits, as it is a snapshot in time that can be built upon by future informatics and knowledge workers, whether they expand this directly in their medical practice or organize these data as training material for future automatic processes in natural language processing constructs.
Author Contributions
Conceived the concepts: DHW. Analyzed the data: DHW, ALF. Wrote the first draft of the manuscript: DHW. Contributed to the writing of the manuscript: DHW, CLM, AMA, ALF, TCG. Agree with manuscript results and conclusions: DHW, CLM, AMA, ALF, TCG. Jointly developed the structure and arguments for the paper: DHW, CLM, AMA, ALF, TCG. Made critical revisions and approved final version: DHW, CLM, AMA, ALF, TCG. All authors reviewed and approved of the final manuscript.
Footnotes
Acknowledgments
We thank Shari C. Clifton, Prasanna Vaduvathiriyan, and Phill Jo for their assistance in the search for the referenced articles used in this publication.
