Abstract
Objective
To quantify the impact of International Classification of Disease 10th Revision Clinical Modification (ICD-10-CM) transition in cancer clinical trials by comparing coding accuracy and data discontinuity in backward ICD-10-CM to ICD-9-CM mapping via two tools, and to develop a standard ICD-9-CM and ICD-10-CM bridging methodology for retrospective analyses.
Background
While the transition to ICD-10-CM has been delayed until October 2015, its impact on cancer-related studies utilizing ICD-9-CM diagnoses has been inadequately explored.
Materials and Methods
Three high impact journals with broad national and international readerships were reviewed for cancer-related studies utilizing ICD-9-CM diagnoses codes in study design, methods, or results. Forward ICD-9-CM to ICD-10-CM mapping was performing using a translational methodology with the Motif web portal ICD-9-CM conversion tool. Backward mapping from ICD-10-CM to ICD-9-CM was performed using both Centers for Medicare and Medicaid Services (CMS) general equivalence mappings (GEMs) files and the Motif web portal tool. Generated ICD-9-CM codes were compared with the original ICD-9-CM codes to assess data accuracy and discontinuity.
Results
While both methods yielded additional ICD-9-CM codes, the CMS GEMs method provided incomplete coverage with 16 of the original ICD-9-CM codes missing, whereas the Motif web portal method provided complete coverage. Of these 16 codes, 12 ICD-9-CM codes were present in 2010 Illinois Medicaid data, and accounted for 0.52% of patient encounters and 0.35% of total Medicaid reimbursements. Extraneous ICD-9-CM codes from both methods (Centers for Medicare and Medicaid Services general equivalent mapping [CMS GEMs, n = 161; Motif web portal, n = 246]) in excess of original ICD-9-CM codes accounted for 2.1% and 2.3% of total patient encounters and 3.4% and 4.1% of total Medicaid reimbursements from the 2010 Illinois Medicare database.
Discussion
Longitudinal data analyses post-ICD-10-CM transition will require backward ICD-10-CM to ICD-9-CM coding, and data comparison for accuracy. Researchers must be aware that all methods for backward coding are not comparable in yielding original ICD-9-CM codes.
Conclusions
The mandated delay is an opportunity for organizations to better understand areas of financial risk with regards to data management via backward coding. Our methodology is relevant for all healthcare-related coding data, and can be replicated by organizations as a strategy to mitigate financial risk.
Introduction
The United States was originally scheduled to transition from the International Classification of Disease 9th Revision Clinical Modification (ICD-9-CM) to ICD-10-CM on October 1, 2014. 1 However on April 1, 2014, the Protecting Access to Medicare Act of 2014 was enacted with delay in ICD-10-CM implementation until October 1, 2015. 2 In 2012, Centers for Medicare and Medicaid Services (CMS) estimated that a 1-year delay in ICD-10-CM implementation could cost $306 million 3 ; a more recent estimate by the American Health Information Management Association estimated an additional $1 billion to $6.6 billion in excess of already incurred costs. 4 The vast majority of these costs are estimated to arise from financing ICD-10-CM implementation including technology upgrades, biller and coder training, and clinical documentation enhancement. However, less attention has been paid to post-implementation costs of, and management strategies for, data accuracy within longitudinal databases spanning both ICD-9-CM and ICD-10-CM.
While ICD-10-CM has greater specificity and promises to be more effective at comprehensive public health data reporting, ICD-10 implementation in other countries has resulted in considerable disruption in data reporting with demonstrable impact on relative risk estimates for death 5 and reported differences in comorbidity coding within 5 years of transition. 6 While the original intent for ICD codes was for global standardization of epidemiological reporting to the World Health Organization (WHO), ICD-9-CM codes are widely utilized in the US for billing and reimbursement purposes, in addition to measuring safety and quality of medical care, designing delivery symptoms and setting healthcare policy, resource utilization, performance measures, healthcare research and clinical trials, and public health reporting. Given that coded data is essential for public health disease tracking, epidemiological healthcare research, cancer registries, clinical trials, healthcare utilization patterns and resource allocation, and payer reimbursements, the impact of discontinuity in data reporting has significant financial impact. Data collection spanning October 2015 will require both forward (ICD-9-CM to ICD-10-CM) and backward (ICD-10-CM to ICD-9-CM) coding to ensure data congruency. Currently, there is a dearth of published literature on the sensitivity and specificity of backward coding methods, and no standard ICD-9-CM and ICD-10-CM bridging methodology for accurate coding in longitudinal data analyses.
The CMS has created a reference bidirectional GEM code translation system which is widely utilized and publically available. CMS GEMs files provide forward and backward translations in a numerical and tabular format7,8 without further information about ambiguous or complex code mappings. The Translational Health Informatics group at the University of Illinois at Chicago (UIC) has developed an unbiased network modeling methodology (Motif web portal ICD-9-CM conversion tool, or Motif web portal tool at http://www.lussierlab.org/transition-to-ICD10CM) to visually map ICD-9-CM and ICD-10-CM code conversions, and quantitatively predict problematic ICD-9-CM to ICD-10-CM bidirectional mappings. 9
Per prior work, 36% of ICD-9-CM to ICD-10-CM translations are associated with complex mappings, and 1% of ICD-9-CM codes have no corresponding ICD-10-CM codes. 9 Despite prior work suggesting hematology–oncology would be the least impacted medical specialty based on a lower ICD-10-CM to ICD-9-CM code ratio, and smaller frequency of complex mappings, 9 recently published data showed forward mapping of commonly utilized ICD-9-CM hematology-oncology diagnoses within an academic cancer center's database, and Illinois Medicaid database resulted in information loss affecting upwards of $500,000, and 6% of total billing costs. 10
The objective of our study was to quantify the impact of ICD-10-CM transition in cancer clinical trials by comparing coding accuracy and data discontinuity with backward ICD-10-CM to ICD-9-CM mapping via two tools, and to develop a standard ICD-9-CM and ICD-10-CM bridging methodology for use in longitudinal data analyses.
Methods
Data Collection
The research project was approved by the University of Illinois Institutional Review Board 20120773. ICD-9-CM diagnosis codes were collected from three high impact medical journals, and utilization data from an Illinois Medicaid dataset.
Identification of Articles Utilizing ICD-9-CM Diagnoses
Three high impact medical journals with broad national and international readerships were selected by the authors for analysis, including the New England Journal of Medicine (NEJM), Journal of Clinical Oncology (JCO), and Blood. All published articles between January 1, 2013 and December 31, 2013 were reviewed manually and independently by two University of Illinois hematology–oncology physicians for utilization of ICD-9-CM diagnoses codes in study design, methods, or results (Fig. 1). Upon final review, only cancer-related studies based in the United States utilizing ICD-9-CM codes were included. While SEER-Medicaid, SEER-Medicare, and Veterans Administration databases (only those which confirmed usage of ICD-9-CM) were included, publications utilizing WHO ICD9, WHO ICD10, ICD-O, National Death Index, and SEER were not included because ICD-9-CM codes were not utilized. Review articles, editorials, correspondence, commentaries, and grand rounds were also excluded. In cases of disagreement or confusion regarding ICD-9-CM usage, corresponding authors of the publication were contacted via email for confirmation. Two contact attempts were made within 4 weeks, and articles without responses were excluded. For articles confirmed to utilize ICD-9-CM codes, all ICD-9-CM diagnoses and the numbers of associated patients were recorded (Table 1).

Selection of articles utilizing ICD-9-CM codes within study design, methods, and/or results.
Final articles and ICD-9-CM codes, with cancer-specific and Klabunde-specific categories.

ICD-9-CM to ICD-10-CM Forward Coding
Each ICD-9-CM code was mapped forward to its associated ICD-10-CM code(s) via the Motif web portal tool (version 1.2) per previously described methodology 9 (Table 1).
ICD-10-CM Gap Analysis
The generated list of ICD-10-CM codes was again reviewed by two hematology-oncology physicians to confirm clinical relevance to the original article. A gap analysis was conducted by comparing generated ICD-10-CM codes to a complete listing of all 2014 ICD-10-CM codes (http://www.icd10data.com) to identify clinically relevant ICD-10-CM codes missing from the initial generated list. This step was necessary as 1% of all ICD-10-CM codes do not have any associated mapping, and to be representative of how a researcher would view ICD-10-CM data. Missing ICD-10-CM codes which were categorized as relevant and important for the study were added to the initial generated ICD-10-CM list to create a comprehensive ICD-10-CM list.
ICD-10-CM to ICD-9-CM Backward Coding and Comprehensive Analysis
The comprehensive list of ICD-10-CM codes (generated and additional relevant codes) was coded backward to ICD-9-CM through two methods, including the Motif web portal tool (v1.2) 9 and publically available existing CMS GEMs files (http://www.cms.gov/Medicare/Coding/ICD10/2014-ICD-10-CM-and-GEMs.html).
To assess data fidelity and identify the method with the highest backward coding accuracy, original ICD-9-CM codes from the seven final articles were compared to the generated ICD-9-CM codes from the Motif web portal method and the CMS GEM method.
Cost Analysis
Original ICD-9-CM codes that were missing from the final lists generated by both backward coding methods were categorized as “lost in translation.” Additional ICD-9-CM codes generated by both methods were categorized as “extraneous.” An Illinois Medicaid database was assessed for the presence of ICD-9-CM codes that were lost in the final translation back to ICD-9-CM per previously described methodology.9,10 Illinois Medicaid data consisted of reimbursement data and associated ICD-9-CM diagnosis codes for calendar year 2010 for all patients assigned to a UIC primary care physician as of April 2011. In total, Illinois Medicaid 2010 data consist of 1,466,581 patient encounters, 299 institutions, 38,644 patients, and $382 million in Medicaid reimbursements. For “lost” and “extraneous” ICD-9-CM codes, associated reimbursements and patient visits were tallied to provide an estimate of potential financial risk associated with ICD-9-CM transition with regards to longitudinal data analysis.
Results
In total, 1,567 original articles were reviewed from the NEJM (247), Blood (908), and JCO (412). Initially, 68 articles were identified as utilizing administrative databases. Of these, 18 were excluded due to being non-US-based studies with the use of country-specific ICD codes; eight were excluded due to being non-cancer related; four were excluded due to lack of email response from corresponding authors; 31 were excluded due to utilization of other diagnoses codes or administrative databases. Seven articles were included in the final evaluation (Fig. 1).
In total, 412 discrete ICD-9-CM codes were identified from all seven articles (Table 1), of which 245 ICD-9-CM codes were associated with the Klabunde comorbidity classification. The Klabunde comorbidity index incorporates diagnostic and procedure data contained within Medicare physician claims to incorporate comorbidities recorded on both inpatient and outpatient claims, and is based on Charlson comorbidity conditions.11,12 A total of 54 ICD-9-CM codes were associated with malignancy: prostate (2), secondary neoplasm of lymph nodes (8), secondary neoplasm of bone and bone marrow (1), colo-rectal-anal (15), breast (11), trachea and lung (7), and lymphoma (10).
ICD-9-CM to ICD-10-CM Forward Coding
In total, 412 ICD-9-CM codes mapped forward to 1437 ICD-10-CM codes; six of these ICD-9-CM parent codes did not map forward to any ICD-10-CM codes. Parent codes are ICD-CM codes that do not map forward to ICD-10-CM, but are still utilized for diagnoses, reimbursement, and clinical research.
ICD-10-CM Gap Analysis
Given that 1% of ICD-10-CM codes are without associated mappings, we felt it necessary to compare the generated ICD-10-CM codes with a manual review of the comprehensive 2014 ICD-10-CM CMS GEMs diagnoses list. This step is necessary for retrospective research utilizing ICD-9-CM and ICD-10-CM codes. Upon manual review of the complete ICD-10-CM CMS GEMs diagnoses list, an additional 441 ICD-10-CM codes were found to be clinically relevant to the initial articles. These additional 441 ICD-10-CM codes were not identified during forward mapping from the original ICD-9-CM codes. In total, 1,864 discrete ICD-10-CM codes were included in comprehensive ICD-10-CM list.
ICD-10-CM to ICD-9-CM Backward Coding, and Comprehensive Analysis
Mapping backward from 1864 ICD-10-CM codes via both the CMS GEMS and Motif web portal methods resulted in a higher number of ICD-9-CM codes than the original list of 406 ICD-9-CM (excluding parent codes). While the Motif web portal method yielded 652 ICD-9-CM codes including 406 original ICD-9-CM codes, the CMS GEMS method yielded 551 ICD-9-CM codes including 390 of the original 406 ICD-9-CM codes. The 16 original codes missing from the CMS-GEMS backward map method are listed in Table 2; none were cancer related, and 12 codes were associated with the Klabunde comorbidity index. The Motif web portal tool demonstrated higher sensitivity for identifying initial ICD-9-CM codes in comparison to the CMS-GEMs method (p < 0.0001 using Wilcoxon matched pairs); the Motif web portal tool sensitivity was 0.985 and specificity was 0.983, compared to CMS GEMs sensitivity 0.947 and specificity of 0.989, respectively (Fig. 2).
ICD-9-CM codes lost in backward translation, and associated fiancial information.

included in Klabunde comorbidity.
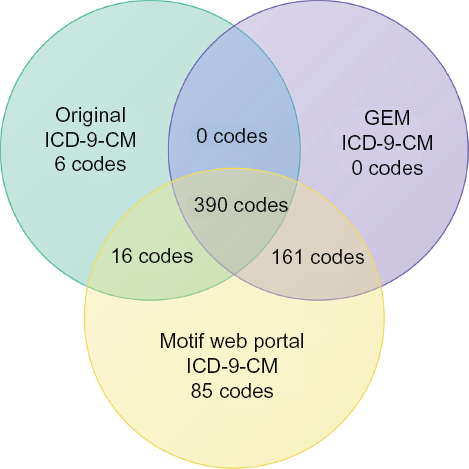
Venn diagram of original ICD-9-CM and backward generated ICD-9-CM codes. The sensitivity of CMS GEMs method was 0.946 and specificity was 0.989. The sensitivity of the Motif web portal tool was 0.985 and specificity was 0.983.
Cost Analysis
The 16 ICD-9-CM codes that were “lost in translation” during backward coding from ICD-10-CM via CMS GEMs method to ICD-9-CM accounted for 3.9% of the original 412 ICD-9-CM codes from all seven articles. Of these 16 codes, 12 were present in the Medicaid data. Medicaid reimbursements associated with the 12 ICD-9-CM codes accounted for a total of 7,489 patient encounters and a cost of $1,134,999 million, or 0.52% of total patient encounters and 0.35% of total Medicaid reimbursements from the 2010 Illinois Medicare database (Table 2).
“Extraneous” ICD-9-CM codes generated by the Motif web portal method (n = 246) accounted for a total of 32,989 patient encounters, and $13,498,800 in Medicaid Reimbursements. “Extraneous” ICD-9-CM codes generated by the CMS GEMs method (n = 161) accounted for a total of 30,560 patient encounters, and $11,396,400 in Medicaid Reimbursements. In total, extraneous ICD-9-CM codes from both methods (CMS GEMs, n = 161; Motif web portal, n = 246) in excess of original ICD-9-CM codes accounted for 30,560 and 32,989 or 2.1% and 2.3% of total patient encounters and $11,395,000 and $13,498,000 or 3.4% and 4.1% of total Medicaid reimbursements from the 2010 Illinois Medicare database. Dollar amounts are rounded to 5 significant digits.
Discussion
Immediate Impact upon Longitudinal Data Analyses
While the majority of ICD-10-CM transition has revolved around implementation challenges, little discussion has been devoted to the impact of coded data discontinuity for clinical trials and cancer registries, public health disease tracking, epidemiological healthcare research, healthcare utilization patterns and resource allocation, and payer reimbursements. Based on international experiences with ICD-10 transition,5,6 we anticipate substantial challenges with backward coding from ICD-10-CM to ICD-9-CM for databases spanning the ICD-10-CM transition, and ensuring the backward codes are consistent and inclusive of the initial ICD-9-CM codes utilized in administrative databases. Based on our evaluation of seven cancer-related studies, we present a specific bridging methodology and analytic tools which can replicated for retrospective longitudinal data analyses to ensure data fidelity between original ICD-9-CM, ICD-10-CM codes, and generated ICD-9-CM via backward coding. We confirm the necessity for a manual review of ICD-10-CM codes after the initial forward coding is completed to ensure a comprehensive and relevant ICD-10-CM list is being utilized, based on our findings that manual review yielded an additional 23% of clinically relevant ICD-10-CM codes absent with forward coding alone. Finally, we show significant differences in backward coding methodology in yielding original ICD-9-CM codes with greater sensitivity and full coverage with the Motif web portal method compared to the CMS GEMs files method. The 16 missing ICD-9-CM codes accounted for a substantial potential financial impact of greater than $1 million, while extraneous ICD-9-CM codes accounted for greater than $24 million in the 2010 Illinois Medicaid database, highlighting significant financial risk of data and diagnoses discontinuity with impact on reimbursements and business analytic forecasting.
International experience with ICD-10 transition provides some insight and limited guidance into the types of challenges we may face in the US with ICD-10-CM transition. In a Swiss analysis evaluating comorbidity indices scoring for three hospitals between 1999 to 2003 following ICD-10 introduction, sensitivity estimates improved by greater than 5% to a sensitivity of 43% over 5 years, suggesting that improved data accuracy was related to a coding “learning curve”. 6 A Canadian publication evaluated 32 medical conditions with ICD-10 diagnoses within 4,008 charts, and subsequently recoded the medical conditions to ICD-9-CM for data comparison; researchers found that ICD-10 data had lower sensitivity for 7 of 32 conditions assessed relative to ICD-9-CM data, and that validity differed between coding versions with sensitivity varying between 9% and 80%. 13 A publication assessing the impact of ICD-9 to ICD-10 transition on relative risk estimates found sensitivity as low as 26%, and reported that inconsistencies in mortality outcomes as classified by ICD-9 and ICD-10 could bias and substantially impact relative risk estimates. 5
However, the majority of these data utilized dual coding rather than backward coding as a means for comparison. There are little published data reporting on the impact of backward coding for longitudinal databases. A recent US-based publication assessed the impact of Medicare payments to hospitals utilizing 2009 Medicare data, through converting ICD-9-CM diagnoses to ICD-10, and converting ICD-10 via CMS GEMs to ICD-9-CM. The study found that backward mapping resulted in change in diagnoses for 3.66% of patients with moderate financial impact with decreases in up to 0.46% of payment distributions to all hospitals. 14 This is consistent with our findings with regards to the substantive impact for “missing” ICD-9-CM codes, impacting more than 7,000 patient encounters and more than $1 million in 2010 Illinois Medicaid reimbursements (0.3% of all bills and 0.5% of all visits). This highlights the importance of supplementing the initial ICD-10-CM codes generated by forward coding with a manual review of a comprehensive ICD-10-CM diagnoses list; given that 1% of ICD-10-CM codes will not have associated ICD-9-CM codes, the potential for data loss is greater without this additional step. Additionally, our study reports a significant number of “extraneous” ICD-9-CM codes with backward coding; extraneous ICD-9-CM codes are potential data confounders with substantial financial impact, affecting 4.4% of total patient encounters (>60,000 patient encounters) and 7.6% of total Medicaid reimbursements (>$24 million) in the 2010 Illinois Medicaid dataset. There is likely greater financial risk associated with backward coding than has previously been reported. Missing or extraneous diagnoses may present as reimbursement differences and significantly impact future practice financial analyses, business forecasting, and resource allocation. Missing diagnoses are more likely to result in decreased reimbursements, while extraneous codes may result in resource misallocation. A reliable methodology for backward coding may mitigate financial risk by identifying areas of data discrepancy and anticipating financial impact.
Notably, our study demonstrated significant differences in data quality through backward coding via two methods, the Motif web portal and CMS GEMs. Both results yielded more ICD-9-CM codes than the original list, but the Motif web portal method successfully yielded all of the initial 412 codes (missing six parent codes), whereas the CMS GEMs method missed 16 of the original 412 codes. Twelve of these 16 codes were relevant for the Klabunde comorbidity classification, demonstrating the potential risk for information loss for comorbidity indices and impact on incorrect scoring. A substantive body of health services research and clinical trials research focuses on specific comorbidities, and comorbid illness is of significant concern for patients with cancer. Widely utilized comorbidity measures include the Charlson index, the Klabunde index (incorporating both inpatient and outpatient diagnoses and procedure codes from Medicare data), the Adult comorbidity Evaluation 27 15 ; disease-specific comorbidity measures are also frequently utilized, and most have been adapted for use with ICD-9-CM administrative databases. 16 It will be necessary to devise new ICD-10-CM coding algorithms for defining comorbidities to improve data accuracy, similar to the Canadian experience.17,18
Long-term Challenges with Longitudinal Data Analyses
Backward coding will be most relevant in the first few years when ICD-9-CM data are still being utilized within clinical research, disease tracking, hospital reimbursements, and practice utilization and resource forecasting. In addition to the implications for clinical research as detailed above, from an organizational financial health and private sector perspective, it is vital to understand comparative data trends from a quarterly to an annual perspective. Without careful attention to data fidelity between ICD-9-CM diagnoses prior to October 1 2015, and ICD-10-CM diagnoses on and after October 1 2015, it is likely that organizations will be at significant financial risk with regards to forecasting resource utilization, understanding differences in reimbursements, and predicting future revenue cycle management. Given that ICD-CM-10 transition has been delayed until at least October 2015, and that ICD-11-CM transition will likely occur within a few years of WHO ICD-11 presentation in 2017, it is even more essential that equivalent comparative data is being utilized for financial decision-making.
Limitations and Future Directions
Limitations of our study include our manual review of initial articles, our use of one forward mapping methodology, our use of a Medicaid dataset, and lack of real-world financial forecasting examples taking into account missing and extraneous diagnoses resulting from backward coding. While manual review was accurate and detailed, development of programming for automated review and quality control would increase our method's applicability toward larger literature reviews, epidemiological trends, and for use in private practices. Based on prior work, we chose to proceed with forward mapping utilizing the Motif web portal methodology rather than the CMS GEMs method; repeating the analysis with the CMS GEMs method may have shown more potential areas of data discontinuity. Medicaid dataset was characterized by a high volume of primary care and pediatrics-related diagnoses; use of a Medicare dataset would likely have yielded a more thorough financial analysis with likely more associated visits and billing costs. Based on prior international experience, current reimbursement practices, and business analytic models, we are concerned about substantial financial risk ensuing from missed and extraneous diagnoses but are unable to parse out the relative financial risk from each grouping. Precise financial risk modeling would be helpful to further quantify financial impact.
Conclusion
In conclusion, there are significant implications for data discontinuity during the ICD-10-CM transition that has not been adequately explored. The mandated delay is an opportunity for organizations to better understand areas of financial risk with regards to data management via backward coding. Based on our evaluation of seven cancer-related studies, we present a broadly applicable ICD-9-CM and ICD-10-CM bridging methodology. These analytic tools can be replicated by organizations and clinical researchers for retrospective longitudinal data analyses to ensure data fidelity between present ICD-9-CM codes, associated ICD-10-CM codes, and the backward mapping of those ICD-10-CM codes to ICD-9-CM codes. We confirm the necessity for a manual review of ICD-10-CM codes after the initial forward coding is completed to ensure a comprehensive and relevant ICD-10-CM list is being utilized. Finally, we show significant differences in backward coding methodology in yielding original ICD-9-CM codes with greater sensitivity and full coverage with the Motif web portal method compared to the CMS GEMs files method. Despite our focus on cancer-related journals, our methodology is widely applicable and relevant for all healthcare-related coding data, and can be replicated by organizations as a strategy to mitigate financial risk.
Author Contributions
Conceived the concepts: NKV, ADB, AS, PD. Analyzed the data: NKV, ADB, AS, PD. Wrote the first draft of the manuscript: NKV, ADB, AS, PD. Contributed to the writing of the manuscript: NKV, ADB, AS, PD. Agree with manuscript results and conclusions: NKV, ADB, AS, PD. Jointly developed the structure and arguments for the paper: NKV, ADB, AS, PD. Made critical revisions and approved final version: NKV, ADB, AS, PD. All authors reviewed and approved of the final manuscript.
