Abstract
This paper concerns a new method for identifying aberrant signal transduction pathways (STPs) in cancer using case/control gene expression-level datasets, and applying that method and an existing method to an ovarian carcinoma dataset. Both methods identify STPs that are plausibly linked to all cancers based on current knowledge. Thus, the paper is most appropriate for the cancer informatics community. Our hypothesis is that STPs that are altered in tumorous tissue can be identified by applying a new Bayesian network (BN)-based method (causal analysis of STP aberration (CASA)) and an existing method (signaling pathway impact analysis (SPIA)) to the cancer genome atlas (TCGA) gene expression-level datasets. To test this hypothesis, we analyzed 20 cancer-related STPs and 6 randomly chosen STPs using the 591 cases in the TCGA ovarian carcinoma dataset, and the 102 controls in all 5 TCGA cancer datasets. We identified all the genes related to each of the 26 pathways, and developed separate gene expression datasets for each pathway. The results of the two methods were highly correlated. Furthermore, many of the STPs that ranked highest according to both methods are plausibly linked to all cancers based on current knowledge. Finally, CASA ranked the cancer-related STPs over the randomly selected STPs at a significance level below 0.05 (
Introduction
Microarray technology is providing us with increasingly abundant gene expression-level datasets. For example, the cancer genome atlas (TCGA) makes available gene expression-level data on cases and controls in five different types of cancer. Translating the information in these data into a better understanding of underlying biological mechanisms is of paramount importance to identifying therapeutic targets for cancer. In particular, if the data can inform us as to whether and how a signal transduction pathway (STP) is altered in the cancer, we can investigate targets on that pathway.
An STP is a network of intercellular information flow initiated when extracellular signaling molecules bind to cell-surface receptors. The signaling molecules become modified, causing a change in their functional capability and affecting a change in the subsequent molecules in the network. This cascading process culminates in a cellular response. Consensus pathways have been developed based on the composite of studies concerning individual pathway components. Figure 1 shows a portion of the signaling pathway of human primary naive CD4 T cells, downstream from CD3, CD28, and LFA-1 activation. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway 1 is a collection of manually drawn pathways representing our knowledge of the molecular interaction and reactions for about 136 pathways. Signaling pathways are not stand alone, but rather it is believed there is inter-pathway communication. 2

A portion of the STP of human primary naive CD4 t cells, downstream from CD3, CD28, and LFA-1 activation.
Many aberrant STPs have been associated with various cancers.3–9 For example, we now know that the ErbB, PI3K-Akt, and Wnt pathways are associated with breast cancer. To develop optimal treatments for cancer patients, it is important to discover which STPs are implicated in a cancer or cancer-subtype.
The phosphorylation activity state of each protein in an STP corresponds to the information flow on the STP. However, protein phosphorylation assays are slow, relatively expensive, and can be performed for a tiny but important fraction of the genome. Protein expression level (abundance) is correlated with activity, and gene expression level (mRNA abundance) is associated with protein abundance (correlation coefficient of 0.4–0.6). So, it seems gene expression data should be loosely correlated with activity. Furthermore, as mentioned above, microarray technology is providing us with increasingly abundant gene expression-level datasets. So, researchers developed techniques that investigate which STPs are implicated in a cancer by analyzing gene expression datasets. Initially, techniques such as over-representation analysis10–12 were employed. These techniques simply determine which genes are differentially expressed in the sample groups. Such methods ignore the topology of the network, and so do not account for key biological information. That is, if a pathway is activated through a single receptor and that protein is not produced, the pathway will be severely impacted. However, a protein that appears downstream may have a limited effect on the pathway. Recently, researchers have developed methods that account for the topology of an STP when analyzing gene expression data to determine whether the STP is implicated in a cancer.13–15 Signaling pathway impact analysis (SPIA) 13 is a software package (http://bioinformaticsprb.med.wayne.edu/SPIA) that analyzes gene expression data to identify whether a signaling network is relevant in a given condition that combines over-representation analysis with a measurement of the perturbation measured in a pathway.
However, the correlation of gene expression with activity is not well established. Some studies show that protein expression level (abundance) is often not positively correlated with activity 16 and that gene expression level is often not correlated with protein abundance. 17 Thus, gene expression levels might at most be loosely correlated with activity, which means that the causal structure of an STP might not be represented by the relationships among gene expression levels. More fundamentally, it remained an open question as to whether there even are causal relationships among the gene expression levels of genes coding for proteins on an STP. Neapolitan et al. 18 investigated this question. Specifically, they used a Bayesian network (BN) model to study whether the expression levels of genes that code for proteins on a given STP are causally related and whether this causal structure is altered when the STP is involved in a particular cancer. The results of their study supported that there is a causal structure and that it is altered.
The technique used in the investigation in Ref. 18 provides us with a new method for analyzing whether an STP is implicated in a cancer using gene expression data. In this paper, we present this technique. Then we apply both this technique and SPIA 13 to the analysis of the ovarian carcinoma dataset provided by TCGA. We obtained highly correlated results using the two methods, and we identified biologically plausible STPs as being the ones to be most likely implicated in ovarian carcinoma.
Method
As our method applies BNs to modeling STPs, we first review BNs.
BNs
A BN19–21 consists of a directed acyclic graph (DAG)
Figure 2 shows a BN representing the causal relationships among a subset of the variables related to lung cancer. Using this BN, we can determine conditional probabilities of interest using a BN inference algorithm.
19
For example, we can determine
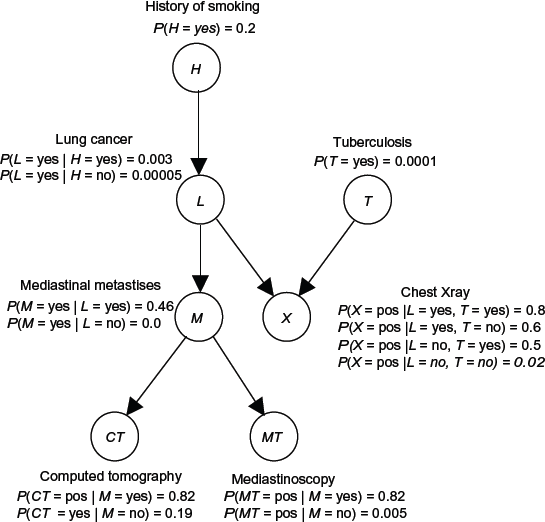
A BN containing a subset of the variables related to lung cancer.
A BN DAG model consists of a DAG
In the constraint-based approach,
22
we learn a DAG model from the conditional independencies that the data suggested are present in the generative probability distribution 
When learning a DAG model from data, we can only learn a Markov equivalence class of DAG models rather than a unique DAG model. Two DAGs are called
Many biological processes have been modeled using BNs including molecular phylogenetics, 25 gene regulatory networks,26–28 genetic linkage, 29 genetic epistasis,30–34 and STPs.35–39
STPs modeled as BNs
If we represent the phosphorylation activity state of each protein in an STP by a random variable and draw an arc from
Causal analysis of STP aberration (CASA)
In what follows for simplicity we will say that a gene coding for a protein on an STP is on the STP itself. We assume we have two sets of data. The first set contains the gene expression levels of all (or at least most) genes in a set of cases (tumors) and the second set contains the gene expression levels of all genes in a set of controls. Let
There are two models. Model 


The larger the value of
We call the method CASA.
Jiang et al.
18
used Equations (2) and (3) to analyze 5 STPs associated with breast cancer, 10 STPs associated with other cancers, and 10 randomly chosen STPs, using a breast cancer gene expression-level dataset containing 529 cases and 61 controls. They obtained significant results indicating that
Application to ovarian cancer
TCGA makes available datasets concerning breast cancer, colon adenocarcinoma, glioblastoma, lung squamous cell carcinoma, and ovarian carcinoma. Each dataset contains data on the expression levels of 17,814 genes in cases (tumorous tissue) and controls (non-tumorous tissue). Table 1 shows the number of cases and controls in each of these datasets.
The number of cases and controls in the five TCGA datasets.

These datasets are highly unbalanced in that there are many more cases that controls. Difficulties can occur with unbalanced datasets. For example, predictive accuracy, a method often used for evaluating the performance of a classifier, is not appropriate when the data are unbalanced. 40 However, our application is discovery, not prediction. The BDeu score, which we employ, automatically incorporates both the number of cases and controls into the resultant score. Too few data items would make it more difficult to distinguish models, but not produce an inappropriate measure. Furthermore, to increase the number of controls, we used the controls from all five datasets, resulting in a total of 102 controls.
We investigated the ovarian carcinoma dataset. We analyzed 20 cancer-related STPs to see which ones the data indicate are involved in ovarian cancer. We also analyzed six arbitrary STPs to see whether STPs involved in cancer in general have more implicating scores than arbitrary STPs. These pathways were selected at random from the KEGG pathways list after removing the cancer-related pathways. The STPs analyzed appear in Table 2.
The STPs analyzed using the TCGA ovarian carcinoma dataset.

Using the KEGG database, we identified all the genes related to each of the 26 pathways. We extracted gene expression profiles for the 591 ovarian carcinoma cases and 102 controls in the TCGA database. By mapping the gene names of the genes in the gene sets identified using KEGG pathways and the gene names in TCGA data, we were able to extract the gene expression profiles for each of the 26 pathways for the 591 cases and 102 controls. All expression levels were discretized to values
Using the resultant datasets, we used CASA to learn Bayes factors and SPIA to determine
Results
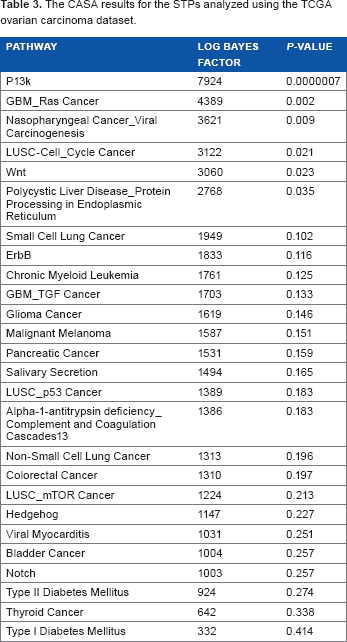
Table 3 shows the results for CASA, and Table 4 shows the results for SPIA. The
The Casa results for the STPs analyzed using the TCGA ovarian carcinoma dataset.
The SPIA results for the STPs analyzed using the TCGA ovarian carcinoma datasets.

Notice that in both tables, the cancer-related pathways in general are near the top. Based on a two-sample
We combined the
The combined results for the STPs analyzed using the TCGA ovarian carcinoma dataset. There is an X in the far right columns if Casa or SPIA separately found the STP noteworthy.

Many of the results obtained are plausible according to current knowledge. PI3 K, which is “probably one of the most important pathways in cancer metabolism and growth,”
43
has
Discussion
We developed CASA, which is a BN-based method for investigating whether STPs are implicated in cancer using case–control gene expression datasets. We applied both CASA and another topology-based method, SPIA, to the TCGA ovarian carcinoma dataset to analyze 20 cancer-related STPs and 6 randomly selected STPs. The results of the two methods were highly correlated. CASA ranked the cancer-related STPs over the randomly selected STPs at a significance level below 0.05 (
These results open up avenues for future research. In particular, we can analyze all 136 pathways in KEGG pathway with the purpose of identifying undiscovered pathways related to ovarian cancer. This analysis will require a good deal of manual effort to develop the individual STP datasets from the manually drawn pathways and the TCGA datasets. Second, we can analyze the remaining four cancers in the TCGA datasets, and perform a pan cancer analysis, looking for STPs involved across all cancers.
Conclusion
We conclude that our study supports that both CASA and SPIA can identify aberrant STPs in cancer using case/control gene expression-level data. These results open up avenues for discovering cancer-related STPs across different types of cancers.
Author Contributions
XJ conceived and designed the experiments. XJ processed the data, developed the datasets representing pathways, and analyzed the data. RN wrote the first draft of the manuscript. XJ contributed to the writing of the manuscript. RN and XJ jointly developed the structure and arguments for the paper. Both authors reviewed and approved the final manuscript.
