Abstract
Lipases are enzymes that catalyze the hydrolysis of lipids. Based on protein structures and sequences, lipases can be classified into different protein families. The majority of conventional mammalian lipases are members of the protein super-families of serine esterases and alpha-beta hydrolases. Differential expression of lipases and related alpha-beta hydrolases in tumor cells has been observed. The physiological or patho-physiological functions of these tumor related enzymes are largely unknown. However, lipases are not only involved in energy metabolism but also in the metabolism of bioactive molecules, e.g. phosphatidic acid or arachidonic acid, suggesting that tumor-specifically expressed lipases might be interesting targets for the development of future treatment strategies. Moreover, independent of the patho-physiological function, tumor associated lipases can serve as targets for immunological treatment strategies. In addition, lipases with exclusive expression in single tumor entities can serve as potential diagnostic targets.
Keywords
Introduction
Lipases are essentially involved in energy metabolism and the generation of second messengers. Since the first description of lipases in the 19th century 1 several other lipolytic enzymes have been described. Based on sequence and structural similarities it was recognized that these enzymes are members of large protein families. Not all members of these super-families are necessarily lipolytic enzymes, e.g. thyroglobulin (TG) and carboxyl ester lipase (CEL) are both members of the carboxylesterase family but TG has lost catalytic activity. 2 Protein structure analysis indicated the existence of a large lipase/esterase super-family. 3 Members of this super-family are characterized by a structural motif, the so called α/β fold. 3 The human genome encodes several genes that are known or predicted members of this super-family. The natural substrates, functions, and evolutionary relationships of these proteins have not been completely resolved. However, differential expression of α/β hydrolases has been implicated in a variety of physiological and patho-physiological processes including cancer.4,5 Especially phospholipases are involved in multiple tumor relevant signaling pathways. Consequently, activities of phopholipase C,6–8 phospholipase D9–11 or phospholipase A212–14 have been shown to be linked with tumor development and progression. Several lipases and other members of the α/β hydrolase super-family are serine hydrolases that are characterized by a conserved GxSxG motif. Strategies for the development of selective inhibitors for this class of enzymes have been developed. 15 Therefore, the pharmacological targeting of these molecules seems feasible and tumor associated lipases might allow the development of novel treatment strategies. Moreover, lipases with high expression in tumor cells can be considered as potential targets for tumor directed immune responses. In general, tumor antigens can be divided into tumor-specific antigens (TSA, e.g. point mutated cellular genes, onco-fusion proteins, or clone-specific idiotypes)16–22 and tumor-associated antigens (TAA). In contrast to TSA that are not present in normal cells, the majority of known tumor antigens are derived from normal cellular genes that are expressed at least under certain conditions in normal cells. These TAA include differentiation antigens, e.g. tyrosinase or Melan-A,23,24 as well as so-called cancer/testis antigens (CTA) which are defined by their restricted expression in testis and tumor cells. 25 CTA were characterized on the basis of cytotoxic T cell responses against these antigens in cancer patients. 26 Other CTA were identified on the basis of expression profiles and database mining. 27 CTA are highly interesting as targets for tumor vaccination. For the majority of CTA no exact physiological or patho-physiological functions have been elucidated. Some CTA, e.g. members of the PRAME (preferentially expressed antigen in melanoma) family 28 and MAGE (melanoma antigen) family, 29 might be directly involved in the malignant phenotype of tumor cells.
In addition to the well known phospholipases involved in cancer relevant signal transduction events (see above), a number of lipase have been linked with cancer and will be discussed in the following sections.
Lipase I: A Ewing Tumor-associated Phospholipase and Cancer/Testis Antigen
Ewing tumor (ET) cells are sensitive for treatment with inhibitors with known activity for phospholipases (our unpublished observation). These inhibitors are relatively un-specific and it remains unclear whether lipases that are expressed in ET are responsible for the observed sensitivity. However, ET are tumors with high expression of enzymes with known phospholipase activity. One of these enzymes is lipase I (LIPI = membrane-associated phospolipase A1 beta). LIPI is a nice example for a tumor associated lipase that might be a potential diagnostic and/or therapeutic target. Based on the initial discovery, LIPI is also known as cancer/testis antigen 17 (CT17). We identified LIPI as ET associated cancer/testis antigen by using high-density DNA-microarray analysis.30,31 ET express highly tumor specific fusion proteins as a result of chromosomal rearrangements between members of the TLS-EWSR1-TAF15 (TET) protein family and members of the E26 (ETS)-family of transcription factors. 32 These fusion proteins act as aberrant transcription factors and are involved in the pathogenesis of ET. Therefore, molecular targeting of fusion proteins might be an interesting strategy for molecular therapy of ET.33–36 Theoretically, TET-ETS fusion proteins are tumor specific antigens and the high tumor specificity of these TSA renders the possibility likely that immunotherapeutic approaches might be successful with minimal side effect.37–40 However, peptides derived from the fusion region have only low binding activity for major histocompatibility (MHC) class I molecules 40 and other targets, e.g. cancer/testis antigens, might be required for efficient generation of cytotoxic T cells. In general, the expression of CTA in ET has been described as a rare event. 41 In addition to LIPI, expression of the CTA X antigen 1 (XAGE1)31,42 and testis expressed 15 (TEX15) 31 has been described. LIPI was described before as CTA on the basis of the typical expression pattern. 43 LIPI protein sequences have been detected only in mammals. 44 LIPI has been characterized as a phospholipase A1 iso-enzyme. 45 Despite the nearly exclusive expression in testis (and thyroid), 31 LIPI is a candidate gene for the lethal mouse mutant Ipd (lipid defect) which is characterized by fat accumulation in the liver, and polymorphisms in the human LIPI have been found to be associated with altered serum triglyceride levels. 46 Like other CTA, LIPI might be involved in reproduction and testicular function as suggested by the detection of LIPI on mature spermatids. 45 By its enzymatic activity LIPI produced lysophosphatidic acid (LPA), a potent modifier of several tumor related functions, e.g. cell transformation, migration, apoptosis inhibition, inhibition of differentiation, and angiogenesis.47–51 The relevant LPA receptor for LIPI signal transduction has not been identified until now. Available gene expression data suggest that among different tumors only ET express high amounts of LIPI. 31 The reason for the high expression of LIPI in ET is unknown. LIPI is located on chromosome 21 in close proximity to the centromere. Because many CTA are encoded by pericentromeric genes, it has been suggested that expression of these genes in germ cells reflects opening of the chromatin. 52 Whether expression of LIPI in ET is a consequence of oncogenic transformation or a reflection of the unknown histogenetic origin of ET has to be shown. In contrast to many other ET associated genes, regulation of LIPI expression by ET specific fusion proteins has not been described. The specificity of LIPI expression in ET suggests that RT-PCR for LIPI might be an interesting diagnostic tool. 31
Lipase H: A Second Lipase with High Expression in Ewing Tumors
The mammalian genome contains a second lipase with high homology to LIPI. This lipase, LIPH (lipase member H, phospholipase A1 alpha) is highly expressed in Ewing tumors but also in several normal tissues and therefore is no CTA.53,54 Like LIPI, LIPH has been characterized as phospholipase A1 iso-enzyme. 45 In contrast to LIPI, homologous proteins to LIPH have been found in other vertebrate species. 44 Mutations in LIPH are responsible for hair growth defects.55,56 The same phenotype is seen in patients with mutations of the receptor P2RY5,57,58 indicating that LPA-species produced by LIPH use this receptor for signaling.
Alpha-beta Hydrolase Domain containing 6: A Tumor/Transformation Associated α/β Hydrolase
LIPI represents an example for a lipase with nearly exclusive expression in a single tumor entity. Available expression data suggest that LIPI might be expressed in other tumor entities only with very low frequency. 31 As mentioned above, ET express other members of the lipase/esterase super-family (e.g. LIPH), but none of these genes has the same specificity as LIPI. Alpha-beta hydrolase containing 6 (ABHD6) is one of these genes. 59 In contrast to LIPI, ABHD6 seems to be regulated by EWSR1-ETS fusion proteins.59,60 Nevertheless, TET-ETS negative tumors have also been shown to express ABHD6. 5 Moreover, Epstein-Barr virus (EBV)-immortalized B-cells (so-called lymphoblastoid cell lines, LCL) express ABHD6. 61 Again, the physiological function of ABHD6 is unclear. ET show a very high expression of ABHD6, but the observation of differential expression of ABHD6 in diverse tumor entities and EBV-immortalized cells suggest that ABHD6 expression might be associated with cell transformation. DNA microarray data indicate differential expression of ABHD6 under conditions with disturbed lipid homeostasis, 62 aryl hydrocarbon receptor (AHR) signaling, 63 or peroxisome proliferator-activated receptor alpha (PPARA) signaling.64,65 ABHD6 can hydrolyze the endocannabinoid 2-arachidonoylglycerol (2-AG), 66 but it seems likely that ABHD6 uses additional physiological substrates. 2-AG has been shown to inhibit tumor invasion 67 and inhibition of 2-AG hydrolyzing enzymes might be an interesting therapeutic strategy. 2-AG binds and activates the cannabinoid receptors CNR1 and CNR2. 68 The same receptors were used by phytocannabinoids. 69 Phytocannabinoids and synthethic derivatives thereof can inhibit cancer cell proliferation, metastasis, and neo-angiogenesis. 69 Increasing the concentration of bioactive CNR1- and CNR2-ligands might have similar effects as the inhibition of 2-AG metabolizing enzymes. Whether cannabinoids are effective against ET cells should be investigated.
Alpha-beta Hydrolase Domain containing 2
DNA-microarray data from the Oncomine data base 70 indicate differential expression of lipases and related members of the alpha-beta hydrolase family in varying tumor entities. Multiple data sets from breast cancer samples indicate high expression of alpha-beta hydrolase containing 2 (ABHD2) in breast cancer in comparison to normal mamma.71,72 This enzyme, a putative peroxidase, also shows high signal intensities in melanoma. 73 High expression of ABHD2 was found in macrophages, 74 but the physiological function of ABHD2 is unknown.
Alpha-beta Hydrolase Domain containing 4 and 12
Another alpha-beta hydrolase which shows high signal intensities in microarray data from tumor samples is alpha-beta hydrolase containing 4 (ABHD4).75,76 ABHD4 shows high signal intensities in brain tumors. Like ABHD6, ABHD4 is involved in the metabolism of endocannabinoids. 77 Another lipase which is involved in endocannabinoid metabolism is alpha-beta hydrolase containing 12 (ABHD12). 66 Based on microarray data, this gene shows high expression in seminomas. 78
Lipoprotein Lipase (Lipase D)
Microarray data from brain tumors75,76 also indicate high expression of lipoprotein lipase (LPL, also known as lipase D, LIPD) in these tumors. Moreover, high LPL enzyme activity has been shown to be associated with shorter survival of patients with non-small cell lung cancer. 79 In contrast, inactivation of LPL by genomic deletions or epigenetic mechanisms was recently found in prostate cancer. 80 Because LPL has anti-inflammatory and pro-apoptotic activities, LPL might be a candidate for a tumor suppressor gene.80–82 The inverse correlation between LPL expression and survival in lung cancer patients might indicate that tumors with high LPL activity have acquired general anti-apoptotic mechanisms that override not only LPL activity but also other pro-apoptotic stimuli.
Alpha-beta Hydrolase Domain containing 9 (Epoxide Hydrolase 3), 14A, and 14B
The methylation level of alpha-beta hydrolase containing ABHD9 has been identified as predictive marker for prostate cancer. 4 ABHD9 is an epoxide hydrolase (EPHX3) with unknown physiological function. DNA-microarrays indicate up-regulation of this enzyme together with ABHD14A in bladder cancer. 83 The latter is also highly expressed in lung carcinoids 84 and (together with ABHD14B) in acute myeloid leukemia. 85 Seminomas also show high expression values for ABHD14B. 86 Again, the physiological function of these enzymes is unknown. The genes for ABHD14A and ABHD14B are located in close proximity in head to head orientation on chromosome 3 (Table 1). Studies from mice indicate that the expression of ABHD14A is regulated by the zinc finger protein ZIC1, 89 a factor involved in different cancer types.90–92
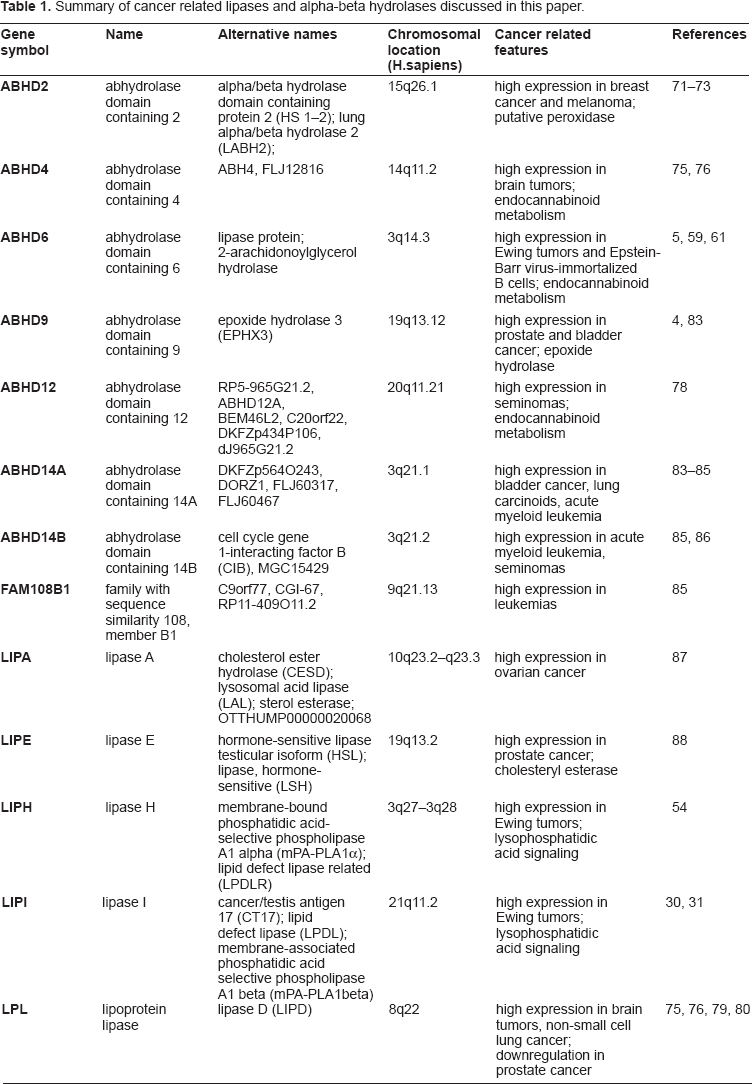
Summary of cancer related lipases and alpha-beta hydrolases discussed in this paper.
Lysosomal Acid Lipase (Lipase A) and Hormone sensitive Lipase (Lipase E)
Lysosomal acid lipase (LIPA) is up-regulated in response to chemotherapy in ovarian cancer cells. 87 LIPE (also known as hormone sensitive lipase, HSL) seems to be directly involved in the pathobiology of castration-resistant prostate cancer cells. 88 In these cells LIPE cleaves stored cholesteryl esters into cholesterol and free fatty acids including arachidonic acid. Free arachidonic acid is further activated by the activity of acyl-CoA synthetase long-chain family member 3 (ACSL3) and transported into mitochondria. Arachidonic acid activates transcription of genes involved in the transport of cholesterol into mitochondria. Finally, this cholesterol is used for androgen synthesis, rendering these cancer cells independent of exogenous androgen sources. Interestingly, gene fusions between ACSL3 and the transcription factor ETV1 (v-ets erythroblastosis virus E26 oncogene homolog E-26 variant 1) have been found in prostate cancer. 93 In contrast to many other gene fusions involving members of the ETS (E twenty six) family of transcription factors, 32 these gene fusions did not result in the expression of fusion proteins but lead to expression of truncated ETV1 proteins as a consequence of high activity of the ACSL3 promotor in the prostate. LIPE is also involved in cancer cachexia.94,95 The presence of factors that increase expression of LIPE in adipocytes in cancer patients has been observed 96 and inhibition of LIPE or LIPE-inducing factors might be a possible strategy for prevention of cancer cachexia.
Potential of Lipases as Targets for Tumor Therapy
Taken together, several lipases and related alpha-beta hydrolases are differentially expressed in cancer cells. The growing super-family of alpha-beta hydrolase domain proteins includes new protein families with unknown physiology and high expression in cancer cells, e.g. FAM108B1 which show high signal intensities in leukemia samples. 85 Tumor or stage specifically expressed lipases might be useful as diagnostic or prognostic markers. Inhibition of lipases or second messengers might be reasonable if the enzymatic activity of the lipase is involved in tumor pathology. Inhibition of LPA signaling can result in tumor inhibition, suggesting that LPA producing enzymes like LIPI might be interesting targets for pharmaceutical interventions. 97 LIPI activity is inhibited by vanadate 45 and might be responsible for the high sensitivity of ET cells for vanadate (our unpublished observation). However, the low specificity of vanadate renders the possibility likely that inhibition of additional ET associated enzymes contribute to this sensitivity. Combinatorial chemistry has been used for development of selective inhibitors for serine hydrolases. 98 Whether this approach can be used for development of LIPI inhibitors has to be shown. Alternatively, inhibition of the corresponding LPA receptor might be possible. The development of inhibitors with ABHD6 specificity has been described. 98 Moreover, ABHD6 is sensitive for the lipase inhibitor tetrahydrolipstatin (THL) 66 and ET cells are sensitive for treatment with THL (our unpublished observation). Again, low specificity of THL does not allow the conclusion that ABHD6 is the only lipase responsible for sensitivity of ET to THL. However, even if the enzymatic activity is dispensable for tumor growth, lipases with high expression in tumor cells can be used as target structures for the development of new treatment strategies (Fig. 1). These lipases can be considered as antigens for cytotoxic T cells. The fact that these antigens are expressed at least under certain conditions in normal tissues did not warrant the conclusion that tolerance for these antigens prevents mounting of effective immune responses. Several tumor-associated auto-antigens have been found to be recognized by cytotoxic T cells from cancer patients (see above) and naturally occurring auto-antibodies against lipases indicate that immune responses against these enzymes can be induced at least under pathological conditions.99,100 Finally, immunological tolerance is major histocompatibility complex (MHC) restricted and the adoptive transfer of allogeneic T cells with specificity for peptides derived from tumor-associated lipases in combination with allo-MHC might be possible in the future. 101
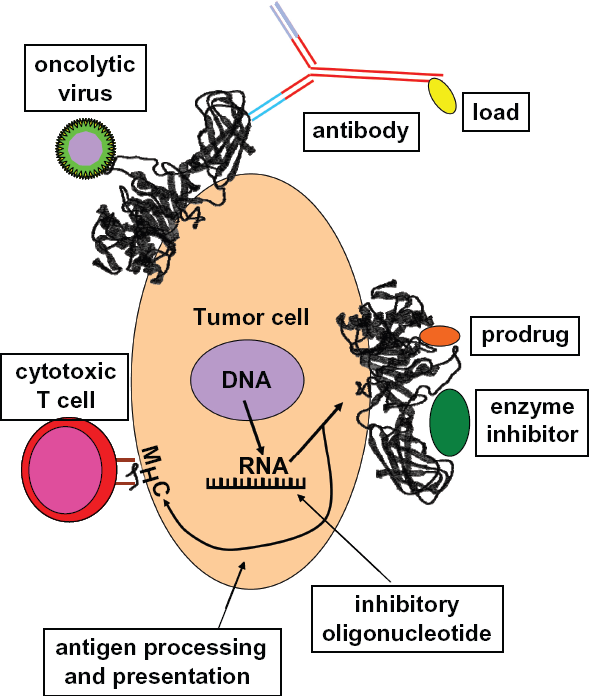
Lipases as therapeutic targets for cancer therapy. Lipases with high expression in tumor cells can be used as therapeutic targets. If the enzymatic function of the protein is required for tumor growth or metastasis, inhibition of the function (by enzyme inhibitors) or expression (e.g. by RNA interference, ribozymes, or anti-sense oligonucleotides) might be possible. If the function is irrelevant for the tumor cell, the enzyme activity can be used for prodrug activation. Independently of the enzymatic activity, antibodies can be used for delivery of toxins or radioisotopes. Oncolytic viruses can be developed which use membrane bound lipases as receptors for cell entry. Finally, cytotoxic T cells with specificity for lipase-derived peptides in combination with MHC class I molecules might be stimulated and kill tumor cells in vivo.
Conclusion
There is increasing evidence for the involvement of lipases and related molecules in cancer pathology. Table 1 summarizes the lipases discussed in the present paper. Pharmacological or immunological targeting of these lipases and the associated signaling pathways might allow the development of new treatment strategies. Further studies are advisable for the clarification of the patho-physiological function of lipases in cancer cells and for the development of future treatment strategies.
Disclosures
This manuscript has been read and approved by all authors. This paper is unique and is not under consideration by any other publication and has not been published elsewhere. The authors and peer reviewers of this paper report no conflicts of interest. The authors confirm that they have permission to reproduce any copyrighted material.
Footnotes
Acknowledgements
We thank Katja Utermoller-Staege (Lektorat Textwarte) and Vera Marks (communication skills) for copy-editing and critically reading the manuscript.
