Abstract
Renal disease is the eighth leading cause of death in the United States. Early diagnosis is usually based on the detection of proteinuria or elevated serum creatinine, a relatively poor biomarker that does not accurately predict renal disease progression. As a result, more predictive biomarkers of renal function are sought. We present preliminary data on three protein biomarkers, folate receptor alpha (FRA), mesothelin (MSLN), and megakaryocyte potentiating factor (MPF), currently being pursued for applications in oncology diagnostics, and evaluate serum and urine levels in subjects with renal disease. Compared to healthy subjects, a significant (
Keywords
Introduction
According to the Centers for Disease Control and Prevention, more than 10% of adults in the United States have some form of kidney disease. Chronic kidney disease (CKD), characterized by a progressive loss in renal function over a period of months or years, is the eighth leading cause of death in the United States. 1 Progression of CKD to end-stage renal disease (ESRD) occurs when the kidneys are no longer able to remove enough wastes and excess fluids from the body. Kidney transplantation and dialysis are the only available therapies for ESRD management. Early detection of renal disease using biomarkers may help to prevent and/or predict progression to CKD and ESRD resulting in better patient management and would be more cost-effective than highly invasive renal transplantation or dialysis procedures.
The initial diagnosis of renal disease is usually based on detection of proteinuria or an elevation of serum creatinine, freely filtered by the glomerulus, 2 and most commonly used to determine an estimated glomerular filtration rate (eGFR), an indicator of renal function. However, use of creatinine as a diagnostic marker has limitations. Serum creatinine varies by age, 3 race, 3 sex, 3 muscle mass,2,4 metabolism, 4 nutritional status, 4 co-morbid conditions, hydration status, and medication use5–9 and, consequently, significant renal disease can exist with minimal or no change in creatinine. 8 Therefore, markers of early injury, especially those that correlate with early fibrosis and progression, are needed and would prove beneficial in both diagnosis and patient management settings. An ideal renal disease biomarker should be accurate, reliable, and easy to measure with a standard non-invasive, reproducible, and sensitive assay. Three biomarkers, folate receptor alpha (FRA), mesothelin (MSLN), and megakaryocyte potentiating factor (MPF), described in detail below, were selected for evaluation as potential markers of renal disease. Each of these biomarkers is currently being evaluated as potential diagnostics in oncology and each is known to be filtered by the kidneys. As such, we reasoned that they may serve as viable candidate markers of renal function. Robust and reproducible assays are available for all three biomarkers.
FRA is a 38–40 Da glycosylphosphatidylinositol (GPI)-anchored protein that binds plasma folate (5-methyltetrahydrofolate) with high affinity (KD ~1nM) and transports it into the cell via endocytosis. In normal human tissues, FRA expression is restricted to polarized epithelial cells in a number of tissues,10–13 including high expression in the proximal tubules of the kidney where it has been shown to play a role in the tubular reabsorption of folate into the circulation. 14 Therefore, impairment of kidney function may prevent reabsorption of folate into the circulation and could cause folate deficiency, 15 which might then result in upregulation of FRA synthesis.16–20 FRA has also been shown to be expressed on a number of epithelial tumors including ovarian, endometrial, lung adenocarcinoma, renal clear cell cancer, and triple negative breast cancer.10–13,21–26
MSLN is a 70 kDa protein, which upon proteolytic cleavage results in a 32 kDa secreted product termed MPF27,28 and MSLN, a 40 kDa GPI-anchored glycoprotein that is also shed into the circulation.29,30 Serum MSLN is currently used for mesothelioma diagnosis, prognosis, and monitoring.31–33 MSLN is hypothesized to be involved in cell adhesion through interactions with MUC16 and intracellular signaling 34 and is highly expressed in ovarian cancer,34–37 mesothelioma,34,37–39 pancreatic cancer, 37 and a subset of lung adenocarcinoma. 37 MSLN also demonstrates restricted expression in normal tissues and is primarily expressed on mesothelial cells of the peritoneum, pericardium, and pleura.34,37,40 Little is known about the biological function of MPF. However, given that it is a cleavage product from the MSLN precursor protein, its expression should mimic that of MSLN. Owing to its value as a diagnostic for mesothelioma, some studies on MSLN relative to kidney function have been reported. Studies have shown that serum MSLN levels were increased in individuals with renal impairment and that these levels were dependent on the stage of CKD.17,31,41,42 Further, Hollevoet et al (2010) showed that an increase in MSLN correlated with an increase in creatinine levels. 31
While some literature on the variation of serum MSLN levels relative to kidney function exists, no such literature exists to our knowledge on serum MPF or FRA, and no comparison relative to the urinary levels of these markers is available. A urine-based assay would of course be ideal as a diagnostic or monitoring tool as it requires totally non-invasive procedures for sample acquisition. As such, we undertook a preliminary investigation of the potential clinical utility of measurements of FRA, MSLN, and MPF, in serum
Methods
Subject samples
This preliminary study included matched serum and urine samples from 200 subjects with varying stages of renal disease and 100 age-matched healthy subjects (Table 1). Samples were obtained from various commercial vendors with Institutional Review Board approvals and patient consent and were collected between 2009 and 2011 by standard techniques and processed/frozen within 30 minutes of collection. Urine samples represented spot collections collected at the time of blood draw and were centrifuged prior to freezing. All samples were stored at −80 °C, and thawed and aliquoted prior to analysis. Patient demographics, including sex, race, age, and stage (Table 1) were obtained from the suppliers.
Demographic and clinical characteristics of healthy subjects and renal disease subjects.

GFR was used in the determination of renal disease stage.
Electrochemiluminescence (ECL) assays
The ECL assays for FRA, MSLN, and MPF used in the present analyses have been described previously.43,44 Samples (serum, urine) from healthy or diseased subjects and standards were added to wells of 96-well plates previously coated with marker specific capture monoclonal antibody (MAb) and incubated at room temperature for two hours. The ruthenium labeled detection MAbs were diluted in assay buffer, added to washed plates, and incubated for an additional two hours at room temperature. Plates were washed, read buffer added, and signals measured using an MSD DISCOVERY WORKBENCH® (Mesoscale Discovery, Gaithersburg, MD). Optimal sample dilutions were: FRA (80-fold dilution of urine and a 20-fold dilution of serum), MSLN (60-fold dilution of urine and an 80-fold dilution of serum), and MPF (4-fold dilution of urine and a 20-fold dilution of serum).
Creatinine analysis
Creatinine concentrations in serum were determined using the QuantiChrom™ Creatinine Assay Kit (BioAssay Systems, Hayward, CA).
Statistical analyses
Pearson's correlation coefficient was performed to determine the correlation between the various biomarkers. Pairwise comparisons of biomarker levels between healthy subjects and stages of renal disease were performed using ANOVA followed by Tukey's post-hoc analysis. Receiver operating characteristic (ROC) analysis was employed to determine the performance of each marker by presence and stage of disease. ROC area under the curve (AUC) calculations were based on 95% confidence intervals. All comparisons were two-sided and a
Results
The ECL assays for FRA, MSLN, and MPF had intraday variability between 2–16% and excellent sensitivity with lower limits of detection (LLOD) of 1.22, 0.29, and 3.35 pg/mL, for FRA, MSLN, and MPF, respectively. 43
Matched serum/urine sample pairs from healthy subjects and subjects with renal disease were measured for FRA, MSLN, and MPF using the described ECL assays and for creatinine using a commercial assay. The patient cohort investigated in the present study was limited and skewed toward stage III disease making some comparisons difficult.
Importantly, all three protein markers were readily detected in both serum and urine matrices. A Pearson's correlation matrix for the serum and urinary levels of these protein biomarkers and serum creatinine in both healthy subjects and subjects with renal disease is presented in Table 2. No significant correlation was observed between serum and urine levels for any of the three protein biomarkers, with correlation coefficients of r =
Pearson's correlation coefficients 1 between biomarkers in urine (u) and serum (s) from healthy subjects or renal disease subjects.

strong.
moderate.
weak.
As can be seen, in healthy subjects and in subjects with renal disease, serum FRA was moderately correlated with MSLN and MPF and, as expected, MSLN was strongly correlated with MPF since those two proteins derive from the same gene product through proteolytic processing. Creatinine only weakly correlated with FRA, MSLN, or MPF. Further, no significant correlation was noted for any of the three protein biomarkers relative to age or gender (data not shown).
Serum levels of FRA, MSLN, MPF, and creatinine are shown in Table 3 and demonstrate a highly significant (
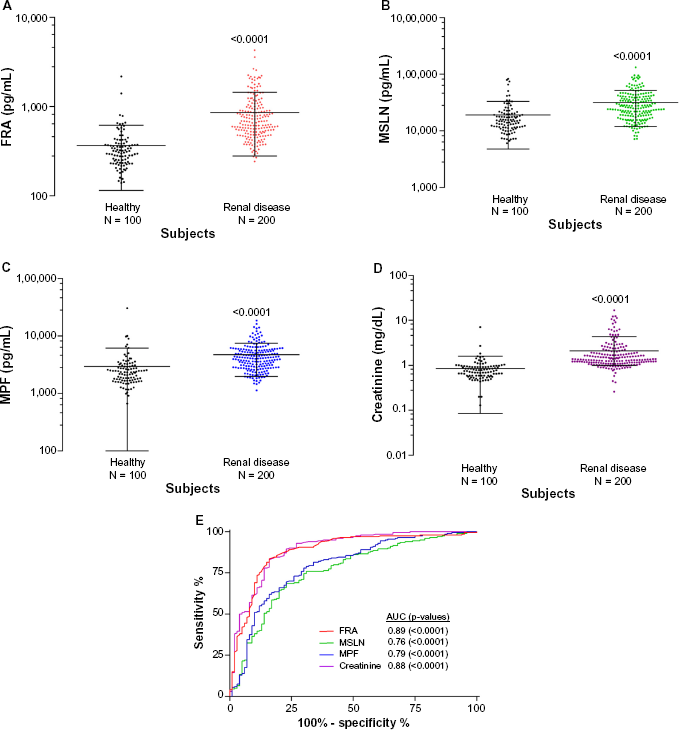
Scatter plots of the serum levels of: (A) FRA, (B) MSLN, (C) MPF, and (D) creatinine in healthy subjects and renal disease subjects. Data is plotted on a log scale. The line and error bars depict mean and standard deviation, respectively.

Scatter plots of the urine (spot collection) levels of: (A) FRA, (B) MSLN, and (C) MPF in healthy subjects and renal disease subjects. Data is plotted on a log scale. The line and error bars depict mean and standard deviation, respectively.
Serum biomarker levels for healthy subjects and renal disease subjects.

Urinary biomarker levels for healthy subjects and renal disease subjects.

ROC analysis (Fig. 1E) of serum levels of these markers resulted in AUCs of 0.89 (

Scatter plots of the serum levels of: (A) FRA, (C) MSLN, (E) MPF, and (G) creatinine in healthy subjects and renal disease subjects by stage of disease. Data is plotted on a log scale. The line and error bars depict mean and standard deviation, respectively.
These data suggest that additional studies aimed at combining a number of markers, including those described herein, may yield higher diagnostic accuracy and monitoring potential for renal disease.
Discussion
Creatinine remains the gold standard in the diagnosis and monitoring of impaired kidney function due to acute kidney injury (AKI), CKD, or subsequent to renal transplantation. Nevertheless, creatinine as a diagnostic marker has limitations, with levels being affected by many factors.3–9 Furthermore, studies have shown that significant renal disease can exist with minimal or no change in creatinine. 9 Owing to these limitations, substantial effort is directed toward the discovery and development of new markers of kidney function, or dysfunction. In the present work, we report preliminary data on three such protein biomarkers, FRA, MSLN, and MPF, measured in both serum and urine samples, to assess their potential value in the diagnosis or monitoring of progression of renal disease in comparison to the standard serum creatinine analysis.
Levels of FRA, MSLN, and MPF were significantly increased in both the serum and urine of subjects with renal disease compared to healthy subjects. Only a weak correlation was observed between serum and urine biomarker values.
The three biomarkers showed moderate to strong correlations with each other in serum, with Pearson's correlation coefficients ranging from 0.68 to 0.75. MSLN correlated well (Pearson's coefficient = 0.75) with MPF, but this is not surprising since these two proteins derive from the same gene product. There is no known biological connection between FRA and MSLN except for the fact that they are both GPI-anchored proteins. As such, it can be speculated that the relatively high correlation between the serum levels of these two proteins reflects a more generalized biologic process related to clearance of these two molecules. While MSLN has been shown to be a binding partner for Muc16 with unknown consequences on its circulatory half-life, FRA has been shown to bind to megalin in both the kidney and liver, and may be removed from circulation, impacting its serum half-life.
45
Creatinine was demonstrated to weakly correlate with all three protein biomarkers in serum (Pearson's coefficients = 0.310.56), which may lend itself to the development of multi-marker panels. Of note, only weak correlations were observed between the three protein biomarkers in urine. This may be a reflection of not only their glomerular clearance but of the sample and sampling
When measured in serum, each of the bio markers investigated showed significant associations with increasing stage of disease, although the level of significance varied. However, further work is required to validate these proteins as potential biomarkers of value for renal disease progression. Unfortunately, the present study was limited by the distribution of stages within the patient cohort and as such should be considered preliminary in nature. Since FRA is expressed at high levels in the proximal tubules, it is interesting to speculate that the increased levels of FRA observed in the serum of subjects with renal disease may be a direct reflection of tubule damage rather than a consequence of or in addition to decreased glomerular filtration
The three protein biomarkers described herein, namely FRA, MSLN, and MPF, have been evaluated as biomarkers in ovarian cancer,34–37,43 mesothelioma,34,37–39 pancreatic cancer, 37 and a subset of lung adenocarcinoma. 37 In the present work, serum levels of these three markers are described to be impacted by impaired renal function, thus suggesting their use as diagnostic biomarkers in cancer need to be evaluated cautiously. Further studies are required to elucidate the relationship between serum and urine concentrations of these markers relative to eGFR and disease state, including the possibility that renal disease of different pathologies, eg glomerular versus interstitial, might result in differential levels of one or more of these markers, and importantly, the impact of renal function on the clinical value of these markers in cancer.
Finally, renal disease is a complex family of diseases involving multiple pathophysiological processes. As a result, a number of biomarkers spanning the known causes of/processes related to kidney disease have been investigated to various degrees. These include markers related to impairments in renal function (Cys-C),46–50 oxidative stress [γ-Glutamyl transpeptidase (GGT)], 51 inflammation or fibrosis [Interleuken (IL)-18], 52 metabolic factors [Apolipoprotein A-IV, 53 fibroblast growth factor-2354,55], and damage to the kidney structure [Liver-type fatty acid-binding protein [L-FABP) 56 ; kidney injury molecule (KIM)-157,58; neutrophil gelatinaseassociated lipocalin (NGAL59,60)]. Similar to cancer therefore, kidney disease is not only a family of diseases but a disease of multiple etiologies. As such, it is likely that no single biomarker will have the required properties of sensitivity and specificity to be universally applicable to the diagnosis or monitoring of kidney function. Panels of biomarkers may be the only solution to this problem and the markers described herein should be considered potential candidates for inclusion in such a diagnostic panel.
Footnotes
Acknowledgments
The authors wish to acknowledge Lisa Marcucci for performing the assays described herein and Jennifer Venzie for assistance in the preparation of this manuscript.
Author Contributions
ES, DO and JV contributed to the writing of the MS. ES and DO conceived and designed the experiments. DO analyzed the data. DO wrote the first draft of the manuscript. ES and DO contributed to the writing of the manuscript. ES and DO agree with manuscript results and conclusions. ES and DO jointly developed the structure and arguments for the paper. ES and DO made critical revisions and approved final version. All authors reviewed and approved of the final manuscript.
