Abstract
Nowadays, cardiac regenerative medicine is facing many limitations because of the complexity to find the most suitable stem cell source and to understand the regenerative mechanisms involved. Mesenchymal stem cells (MSCs) have shown great regenerative potential due to their intrinsic properties and ability to restore cardiac functionality, directly by transdifferentiation and indirectly by paracrine effects. Yet, how MSCs could respond to definite cardiac-committing microenvironments, such as that created by resident cardiac progenitor cells in the form of cardiospheres (CSs), has never been addressed. Recently, a putative MSC pool has been described in the mediastinal fat (hmADMSCs), but both its biology and function remain hitherto unexplored. Accordingly, we investigated the potential of hmADMSCs to be committed toward a cardiovascular lineage after preconditioning with CS-conditioned media (CCM). Results indicated that CCM affects cell proliferation. Gene expression levels of multiple cardiovascular and stemness markers (MHC, KDR, Nkx2.5, Thy-1, c-kit, SMA) are significantly modulated, and the percentage of hmADMSCs preconditioned with CCM and positive for Nkx2.5, MHC, and KDR is significantly higher relative to FBS and explant-derived cell conditioned media (EDCM, the unselected stage before CS formation). Growth factor-specific and survival signaling pathways (i.e., Erk1/2, Akt, p38, mTOR, p53) present in CCM are all equally regulated. Nonetheless, earlier BAD phosphorylation (Ser112) occurs associated with the CS microenvironment (and to a lesser extent to EDCM), whereas faster phosphorylation of PRAS40 in FBS, and of Akt (Ser473) in EDCM and 5-azacytidine occurs compared to CCM. For the first time, we demonstrated that the MSC pool held in the mediastinal fat is adequately plastic to partially differentiate in vitro toward a cardiac-like lineage. Besides, we have provided novel evidence of the potent inductive niche-like microenvironment that the CS structure can reproduce in vitro. hmADMSCs can represent an interesting tool in order to exploit their possible role in cardiovascular diseases and treatment.
Keywords
Introduction
Cardiovascular diseases are among the leading causes of mortality and morbidity in industrialized countries. Several attempts have been made to repair tissues characterized by limited regenerative potential, such as the heart (3,21). Although heart transplantation still represents to date the gold standard therapy for patients with end-stage heart disease, over the years the use of adult stem cells able to transdifferentiate into cardiomyocytes, vascular smooth muscle, and endothelial cells has received great attention, representing the basic foundation of cardiac regenerative medicine.
In this regard, numerous cell sources have been tested in the attempt to regenerate the infarcted myocardium, creating great hope on the clinical exploitation of regenerative therapies (31). In particular, human adipose tissue-derived mesenchymal stem cells (hADMSCs) have become an attractive cell source. hADMSCs can be retrieved in high numbers, they are easily expanded in vitro, and they express the same surface markers as bone marrow-derived mesenchymal stem cells (BMMSCs) (4,10). Moreover, hADMSCs have been shown to reduce the infarcted area after injection in murine models of myocardial damage (36,51,57), exhibiting a strong and long-lasting angiogenic potential (5). Their differentiation toward a cardiogenic phenotype has been previously described (46,50), spontaneously (42), by using 5-azacytidine (44) or by exposing them to the extract of rat cardiomyocytes (15).
To date, hADMSCs are currently under clinical evaluation for the treatment of ischemic heart failure (www.clinicaltrials.gov identifiers: NCT01449032, NCT01709279).
Resident cardiac progenitor cells (CPCs) are a promising therapeutic population due to their unique intrinsic cardiac commitment among adult stem cells. CPCs in the form of cardiospheres (CSs) are widely characterized in an in vitro 3D model of spontaneous niche-like microtissue, made of CPCs and supporting mesenchymal-like cells (26), also representing a sort of reprogramming microenvironment (13). In particular, the microenvironment is considered critical for properly expanding and controlling the differentiation of resident or injected stem cells, both in physiological and pathological conditions (11).
Over the years, it has been clearly demonstrated that cardiac regeneration via stem cell delivery also occurs in a paracrine fashion, influencing the resident microenvironment, where different stem cell niches respond to the stimuli (20,35). It has also recently been suggested that a combined MSC/CPC therapy might have the advantage of increased therapeutic effects (56), despite the mechanisms involved being still mostly unknown. Consequently, the interplay between MSCs, CPCs, and the microenvironment has become highly interesting.
In 2012, a multipotent stem cell population from mediastinal depots was characterized, providing the evidence of a stem cell population in this site of the body (41). However, the biology and the potential of human mediastinal ADMSCs (hmADMSCs) still remain unexplored.
We have recently optimized the isolation and the expansion method of hmADMSCs (47). In this report, our aim has been to investigate the plasticity of hmADMSCs as well as the phenotypical modifications that an in vitro cardiac-specific microenvironment could induce on this novel stem cell population.
In light of this, we have evaluated the potential of CS-conditioned media (CCM) to commit hmADMSCs toward a cardiac-like lineage. Our data have shown that CCM drives hmADMSCs toward a cardiac/endothelial/muscular-like phenotype by modulating the expression levels of multiple cardiogenic markers and by inducing the activation of several intracellular survival signaling pathways.
Materials and Methods
Collection of Adipose and Cardiac Biopsies
Anterior mediastinal fat (four male patients with a mean age of 55 ± 11) and human auricola biopsies (four male patients with a mean age of 64.5 ± 10.2) were obtained from patients undergoing thoracic (thoracic benign diseases) and cardiac surgery (ischemic cardiomyopathy), respectively.
Written informed consent was obtained from patients before surgery. The protocol was conducted in compliance with the tenets of the Declaration of Helsinki for experiments involving human tissues and approved by the Ethical Committee of S. Andrea Hospital, Italy (Prot. CE No. 5195/2013). All data were deidentified and analyzed anonymously.
hmADMSC Cultures
hmADMSC cultures were obtained as we previously described with slight modifications (47). Briefly, biopsies were minced and digested by PBS (Cat. No. D8537; Sigma-Aldrich, St. Louis, MO, USA), trypsin-EDTA (Cat. No. AU-X0930-100; Aurogene, Rome, Italy), and 1 mg/ml collagenase (Cat. No. 17100; Gibco, Monza, Italy) for 45 min at 37°C. Cells were then counted, resuspended in complete medium (DMEM-low glucose/10% FBS/1% PenStrep/1% glutamine/1% nonessential amino acids; all Biowest, Nuaillé, France), and seeded at 4,000 cells/cm2. All nonadherent cells were removed after 72 h. The medium was changed every 3 days. Cultures were expanded until passage 3. Cell count and viability were measured by trypan blue (Cat. No. T8154; Sigma-Aldrich). All the experiments were performed at passage 3.
FACS Analysis, Colony-Forming Unit Fibroblasts Assay (CFU-F), and Transdifferentiation Tests
The phenotype, the clonogenic, and the transdifferentiation abilities of hmADMSCs were assessed as we previously described (47). Briefly, cells were stained for 30 min with the following panel of primary and secondary antibodies: 0.2 mg/ml CD105, 0.1 mg/ml CD44, 0.1 mg/ml CD90, 0.3 mg/ml CD45 (human mesenchymal stromal cell marker panel, Cat. No. Ab93758; Abcam, Cambridge, UK), 1:100 CD31 (Cat. No. P8590; Sigma-Aldrich), 0.25 mg/ml CD117 (Cat. No. Ab5506; Abcam), 1 mg/ml CD133 (Cat. No. Ab19898; Abcam), 1:10 CD34 (Cat. No. 130-081-001; Miltenyi Biotech, Calderara di Reno, Italy), anti-rabbit Alexa Fluor 488 (Molecular Probes OR, USA), mouse FITC-conjugated anti-mouse IgG (H+L) (Jackson ImmunoResearch, Suffolk, UK), and mouse IgG2a (Miltenyi Biotech, Calderara di Reno, Italy) were used as secondary antibodies. Cells were analyzed by cytofluorimetry (FACSAria II; analyzed by DiVa Software v6.1.1; BD Biosciences, San Jose, CA, USA).
For the CFU-F assay, cells were seeded at low density (20-10 cells/cm2/100 mm Petri dish). After 2 weeks of incubation, visible colonies were stained with Giemsa (Sigma-Aldrich). Clusters of cells >5 mm were considered positive and observed by microscope (47).
For transdifferentiation tests, hmADMSCs were subjected to osteogenic, adipogenic, and chondrogenic differentiation. Briefly, cells were cultured for 2-3 weeks using StemPro® Adipogenesis Differentiation kit (1 × 104 cells/cm2, Cat. No. A10070-01; Gibco), StemPro® Chondrogenic Differentiation kit (2.5 × 105, Cat. No. A10071-01; Gibco), and StemPro® Osteogenesis Differentiation kit (5 × 103 cells/cm2, Cat. No. A10072-01; Gibco), respectively. Accumulation of lipid droplets was tested by Oil red O staining (Cat. No. 01391-500ml; Sigma-Aldrich). The calcified matrix deposition and chondrospheres were stained with Alizarin red (Cat. No. A5533; Sigma-Aldrich) and Alcian blue (Cat. No. 05500; Sigma-Aldrich), respectively.
Cell Cultures for Conditioned Media Collection
CSs were isolated, expanded, and characterized as previously described (6). CSs were grown on poly-D-lysine (BD Biosciences)-coated plates in CS growth medium (CGM): 35% IMDM/65% DMEM/F-12 Mix (Gibco, Lonza, Italy), 3.5% FBS, 1% penicillin–streptomycin, 1% L-glutamine, 0.1 mM 2-mercaptoethanol, 1 unit/ml thrombin (Sigma-Aldrich), 1:50 B-27 (Invitrogen, Monza, Italy), 80 ng/ml bFGF, 25 ng/ml EGF (Peprotech, London UK), 4 ng/ml cardiotrophin-1 (Peprotech). Half of the CGM was partially replaced at day 4. CCM was collected 1 week after cell plating, centrifuged at 1,000 rcf for 5 min, and stored at −80°C.
Explant-derived cells (EDCs) were collected from primary cultures, according to the same protocol for CS isolation (6), but instead of plating them in CS-forming conditions, they were kept on fibronectin coating (BD, Milan, Italy) and IMDM medium (Gibco) supplemented with 1% penicillin-streptomycin, 1% L-glutamine, 0.1 mM 2-mercaptoethanol (Gibco), and 20% FBS for one passage. EDCs were then plated at the same cell density, same growth area/media volume ratio, and same FBS concentration (3.5%) as CSs for conditioned media collection after 1 week of culture.
Preconditioning of hmADMSCs
hmADMSC cultures were trypsinized, washed, then seeded in six-well plates (Cat. No. 353046; BD Falcon, Franklin Lakes, NJ, USA), cell density of 4 × 104 cells/well, with FBS (10%), CGM, CCM, EDC-conditioned medium (EDCM), or 5-azacytidine (10 μM; Sigma-Aldrich). All conditioned media were diluted 1:1 before use with the corresponding fresh basal media. After 4 days, half of the medium was removed, and fresh CGM, FBS, CCM, EDCM, or 5-azacytidine was replaced in culture. Cultures were incubated for 7 days. Cells were monitored by optical microscope every 3-4 days. Semiconfluent hmADMSC cultures were split at day 4, leaving half of the conditioned medium and adding fresh CCM, EDCM, or 5-azacytidine as described above.
Cell Size Analysis and MTS Proliferation Assay
Cell morphology analysis was performed on optical images by semiautomatic area tracing and roundness calculation on single cells (n=60 total) with ImageJ software (NIH, Bethesda, MD, USA) maintaining identical instrument settings for all conditions.
Proliferation rate was assessed by MTS assay as previously described (33,45). Briefly, 1 × 103 cells/well were seeded into 96-well plates (Cat. No. 353072; BD Falcon) in quintuplicate in complete medium and left adhering to the plate. The day after, complete medium was removed, and hmADMSCs were conditioned for 0, 3, 5, and 7 days with FBS, CCM, or EDCM as described above. At the end of the incubation period, the proliferation rate was measured by CellTiter 96®AQueousOne Solution Cell Proliferation Assay (Cat. No. G1112; Promega, Madison, WI, USA) according to manufacturer's instructions.
Real-Time PCR
Total RNA was extracted using Total RNA extraction kit (Norgen Biotek Corp., Thorold, Canada) according to manufacturer's instructions, then reverse-transcribed using high-capacity cDNA Reverse Transcription kit (Applied Biosystems, Monza, Italy). The evaluation of the expression of cardiac stem cell/differentiation genes (Table 1) was performed by quantitative real-time PCR (33,45) with SensiMix SYBR Hi-ROX kit (Bioline, London, UK) on a 7900HT Fast real-time PCR System equipped with SDS software (Applied Biosystems) for 40 thermal cycles (95°C for 15 s, 56°C/58°C for 15 s, 72°C for 15 s; see Table 1 for primer sequences and annealing temperatures). The ΔΔCt method was used for relative quantification, using GAPDH as the housekeeping gene (13,53) and CGM culture condition as the reference, unless specified.
Gene Primers for Real-Time PCR

Immunofluorescence
hmADMSCs were fixed with 4% paraformaldehyde (Cat. No. 158127; Sigma-Aldrich), permeabilized with 0.5% Triton X-100 (Sigma-Aldrich), incubated in blocking buffer (0.2% gelatin; Sigma-Aldrich) for 1 h at room temperature. The primary antibodies KDR (kinase insert domain receptor, 1:200, Upstate, Cat. No. 05-554; Merck Millipore, Vimodrone, Italy), MHC (1:200, Cat. No. Ab50967; Abcam), Nkx2.5 (1:200, Cat. No. Ab91196; Abcam), Cx43 (1:100, Cat. No. MAB3067; Millipore) were incubated overnight at +4°C, whereas the secondary antibodies (Alexa Fluor 594 or 488; Invitrogen) were added at room temperature for 45 min (1:400 MHC and 1:1,000 KDR and Nkx2.5). Images were acquired by fluorescent microscope (software IAS2000; Leica, Milan, Italy). Nuclei were stained by 4′-6′-diamidino-2-phenylindole (DAPI; 1:1,000, powder >98%; Cat. No. D9542; Sigma-Aldrich). Ten representative fields were randomly selected, and automated particle count was performed by ImageJ Software. Only the bright Nkx2.5+, MHC+, or KDR+ cells were considered.
Intracellular Signaling Analysis
hmADMSCs were starved overnight (0.2% FBS) and then exposed for 20, 60, or 180 min to the different media conditions described above. Cells were then lysed (Cell Lysis Buffer; Cell Signaling, Euroclone, Milan, Italy), and protein concentration was measured by standard Bradford assay (Cat. No. 500-0006; Bio-Rad Laboratories GMbH, Munchen, Germany). Time course screening of activated pathways was performed by PathScan Intracellular Signaling Array Kit (Cell Signaling), according to the manufacturer's instructions. Chemiluminescent readout of arrays was analyzed for densitometric quantification by ImageJ software.
Statistics
Statistical analysis was performed and the data plotted using GraphPad Prism 5 software (San Diego, CA, USA). The independent sample two-tailed t-test with associated 95% confidence intervals was used to compare the single data. For multiple comparisons, the one- or two-way analysis of variance (ANOVA) test and Bonferroni post hoc test were used. Values of p < 0.05 were considered to be significant. Data are expressed as means ± SEM unless specified.
Results
hmADMSC cultures were isolated from bioptic samples of anterior mediastinal fat (47). Cells were characterized according to the expression of MSC surface markers, to their ability to transdifferentiate into the three mesodermal lineages (osteogenic, chondrogenic, and adipogenic differentiation), and to form clones (47). By FACS analysis (Fig. 1A), cultures were highly positive for CD105, CD90, and CD44, displaying a negative phenotype for CD34, CD133, CD45, CD117, and CD31 as expected (47). In addition (Fig. 1B), hmADMSCs were able to function as stem cells, generating multiple colonies from single stem cells at passage 3 and thus meaning that human mediastinal adipose tissue holds functional stem cells, which were also shown to differentiate toward the three mesodermal lineages (osteogenic, adipogenic, and chondrogenic) (47).

(A) Representative forward versus side scatter dot plot (gate P1) and FACS analysis of hmADMSCs cultures at passage 3. The graph displays a negative phenotype for CD34, CD133, CD45, CD117, and CD31 and a positive phenotype for CD44, CD90, and CD105. Four independent experiments were averaged. (B) Representative image of CFU-F like showing the clonogenic capacity of hmADMSCs. Magnification: 5x. Isolated hmADMSCs show multipotent differentiation ability into the three mesodermal lineages. Specifically, osteogenic transdifferentiation: mineralization of the extracellular matrix was observed by alkaline phosphatase staining; adipogenic transdifferentiation: lipid droplets were visualized with Oil red O staining; chondrogenic transdifferentiation: chondrospheres were stained with Alcian blue. Magnification: 10x.
Proliferation and Phenotype Analysis of hmADMSCs After CCM Conditioning
In order to evaluate whether a cardiac-specific microenvironment is able to influence the plasticity of MSCs, we conditioned hmADMSCs derived from four subjects for 7 days with CCM derived from four CS lines of different patients (each CCM has been individually tested on single lines of ADMSCs), analyzing cell morphological changes, proliferation by MTS, and phenotype by realtime PCR according to a panel of cardiac/endothelial/muscular genes (Table 1).
After 7 days of preconditioning, hmADMSC morphology was not significantly affected, as confirmed by both cell area and roundness measurements, which were comparable among cultures in the three conditions (FBS, CCM, and CGM, p > 0.05) (Fig. 2A, B).

(A) Optical images of hmADMSCs preconditioned with FBS, CGM, and CCM at day 7. (B) The graph shows that both cell area and roundness measurements are comparable among cultures in the presence of FBS, CGM, or CCM. (C) MTS assay (0, 3, 5, 7 days). Cell proliferation of hmADMSCs significantly increases at day 7 after preconditioning with CCM (∞ symbol) or CGM (∴ symbol) compared with FBS. EDCM is able to significantly decrease cell proliferation compared with both FBS (day 5 ** and day 7 ⌂ symbol), CCM and CGM (both at days 5 and 7 #). Replicates were averaged, and the error bars represent the SEM of three independent experiments. **p< 0.01, #, ∞, ∴, ⌂ p< 0.001. O.D., optical density.
In addition, we detected (Fig. 2C) a statistically significant increase in cell proliferation of hmADMSCs at day 7 when cultured with CCM or CGM compared with FBS (both p < 0.001). Interestingly, the treatment with the conditioned media produced by EDCs (EDCM), that is the culture stage before the selective step of CS formation and thus representing our biological control, significantly decreased cell proliferation compared with FBS (day 5 p < 0.01, day 7 p < 0.001), CCM, and CGM (both at days 5 and 7 p < 0.001). No statistically significant difference in cell proliferation was found between CGM and CCM (p > 0.05).
Moreover, we observed a statistically significant upregulation of Nkx2.5, KDR, and MHC mRNA levels of hmADMSCs cultured in CCM compared to FBS (p = 0.03, p = 0.04, and p = 0.035, respectively) (Fig. 3), whereas Thy-1 and SMA mRNA levels decreased (p < 0.0001 and p = 0.005, respectively). A further significant downregulation of c-kit in hmADMSCs cultured in the presence of CCM was also found compared to FBS (p = 0.0001).

Gene expression profile of hmADMSCs after preconditioning with CCM, FBS, and EDCM. After 7 days in the presence of CCM, cells display significant upregulated mRNA levels of Nkx2.5, KDR, and MHC and lower levels of Thy-1, c-kit, and SMA compared with FBS. Moreover, the phenotypic trend of hmADMSCs in regard to Nkx2.5, KDR, MHC, and c-kit after preconditioning with EDCM resembled the FBS condition, but for Thy-1 and SMA, whose mRNA levels were similar to CCM. No statistically significant difference has been observed with regard to Cx43 and GATA-4 in all conditions. CGM was considered as reference. #p < 0.001, *p < 0.05, **p < 0.01.
To then investigate whether the CCM-mediated effect was specific to CSs, we also tested as biological control the conditioned media produced by the unselected population that is EDCs (EDCM) as for the proliferation assay. Similarly to the comparison with the FBS condition, we observed a significant upregulation of Nkx2.5, KDR, MHC, and c-kit mRNA levels after preconditioning with CCM with respect to EDCM (p = 0.04, p = 0.045, p = 0.033, p = 0.0008, respectively) (Fig. 3). No statistically significant difference was found between CCM and EDCM with regard to Thy-1 and SMA (p = 0.095, p = 0.08, respectively) and in the expression of Cx43 and GATA-4 in all conditions (p > 0.05).
Afterward, in order to quantify the percentage of hmADMSCs committed to a cardiac-like phenotype, we performed an immunofluorescence analysis on the upregulated genes (Nkx2.5, MHC, KDR). Results (Fig. 4A, B) indicated that the percentage of hmADMSCs positive for Nkx2.5, MHC, and KDR in CCM was significantly higher than in CGM (p < 0.05, p<0.01, p < 0.05, respectively), FBS (p < 0.01, p < 0.0001, and p < 0.001, respectively), or in EDCM (p < 0.001, p < 0.0001, and p < 0.0001, respectively). A statistically significant difference was also found between the CGM treatment and FBS and EDCM relative to KDR (both p < 0.01).

(A) Quantification of immunofluorescence. The percentage of hmADMSCs positive for Nkx2.5, MHC, and KDR is significantly higher after preconditioning with CCM compared with CGM, FBS, and EDCM. (B) Representative images of immunofluorescence staining with CCM, FBS, EDCM, and CGM. Nuclei were identified by DAPI staining (blue), MHC green, Nkx2.5, and KDR red. #p < 0.001, *p < 0.05, **p < 0.01. (C) Representative images of immunofluorescence for Cx43 of preconditioned hmADMSCs (FBS, CGM, EDCM, and the three CCM02, 10, 16) stained with secondary antibody alone and the specific antibody. DAPI staining (blue); Cx43 red. (D) Gene expression profile of hmADMSC cultures treated with 5-azacytidine for 7 days, displaying a huge interpatient variability (see text). #p < 0.001, *p < 0.05, **p < 0.01. (E) Representative images of immunostaining after treating hmADMSCs derived from different patients after treatment with 5-azacytidine. Nuclei were identified by DAPI staining (blue); MHC green, Nkx2.5, and KDR red. Pt, patient.
In addition, in order to validate the real-time PCR results, a nonmodulated gene marker such as Cx43 was also evaluated. Results showed a negative staining for hmADMSCs in all conditions (Fig. 4C).
In addition, we assessed whether hmADMSCs could respond to chemical transdifferentiation inducers, such as 5-azacytidine, a widely used cardiac differentiation agent (44). Surprisingly, results showed a dramatic interpatient variability, where only one (patient 5) out of five analyzed patients demonstrated a significant upregulation of MHC (patient 1 showed a moderate upregulation), whereas a positive modulation of SMA, KDR, and Cx43 varied considerably between patients (Fig. 4D). Interestingly, Nkx2.5, c-kit, Thy-1, and GATA-4 were consistently downregulated or unchanged in all patients but patients 2 (c-kit and Nkx2.5) and 5 (Thy-1). This result was also confirmed by immunofluorescence analysis, showing that 5-azaytidine induced a more pronounced cardiac-like phenotype evidenced by the expression of markers such as Nkx2.5, MHC, and KDR only in a subset of patients (Fig. 4E). Even a prolonged exposure to 5-azacytidine (up to 21 days of treatment) did not increase mRNA levels of the aforementioned gene markers in hmADMSC cultures (Fig. 5).
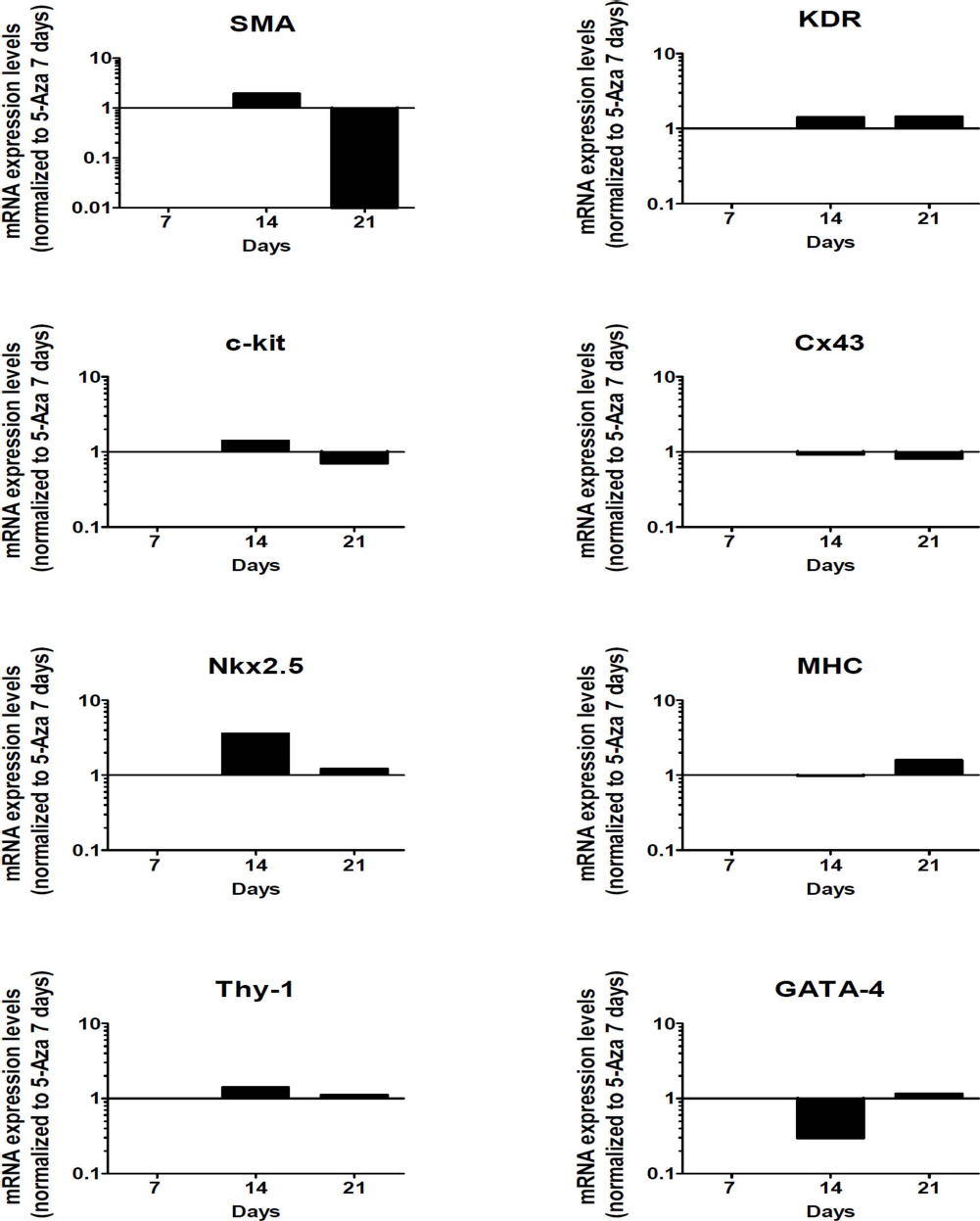
Time course (patient 5) of cardiac marker gene expression after treatment with 5-azacytidine up to 21 days, displaying no increase or even downregulation in almost all the gene markers analyzed. The condition 5-azacytidine 7 days has been considered as reference.
Intracellular Signaling
Finally, we investigated whether specific intracellular signaling nodes were involved when hmADMSCs were subjected to in vitro preconditioning with CCM by screening the phosphorylation and/or the cleavage of 17 proteins involved in key survival/growth pathways and their trend over time (time course 20, 60, and 180 min). Results showed that 20 min after conditioning with CCM, BAD was early phosphorylated relative to all conditions (p < 0.05), reaching a peak at 1 h (p < 0.001 vs. FBS and 5-azacytidine) and a plateau at 180 min where all stimulations converged (Fig. 6A, B). The same intracellular signal was mimicked by EDCM and CGM, however, displaying a delay (between 20 and 60 min) compared to CCM, although statistically significant compared to 5-azacytidine and FBS at 60 min as observed for CCM (p < 0.001). The delay in BAD phosphorylation was even more pronounced after treating hmADMSC cultures with 5-azacytidine and FBS, which reached a plateau only after 180 min of stimulation. Differently, EDCM conditioning significantly activated the early phosphorylation of Akt on Serin 473 after 20 min (p < 0.05) relative to CCM, CGM, and FBS, but similar to 5-azacytidine. This trend constantly increased concurrently to 5-azacytidine, and the difference with the remaining conditions was even increased over time (p < 0.01 and p < 0.001 EDCM vs. 5-azacytidine and CGM/FBS/CCM, respectively, at 60 min and p < 0.05 5-azacytidine vs. FBS/CCM/CGM).

(A) Representative images of the intracellular signaling arrays where hmADSMCs were serum starved and then conditioned with FBS, CGM, CCM, EDCM, and 5-azacytidine (5-AZA) for 60 min. (B) Graphs (time course 20, 60, 180 min) indicate that BAD phosphorylation is early induced (20 min) upon stimulation with CCM, whereas other intracellular signaling pathways such as PRAS40, S6 ribosomal protein, and Akt (Ser473) are preferentially activated in the presence of FBS, CGM, EDCM, and 5-azacytidine, respectively. Data were normalized on the array positive control. O.D., optical density. #p < 0.001, *p < 0.05, **p < 0.01.
PRAS40 and the S6 ribosomal protein were significantly activated by the use of FBS after 20 min (p < 0.01) or CGM after 1 h (p < 0.05) in culture, respectively.
No statistically significant difference has been observed with regard to other cellular proteins involved in different signaling pathways such as Stat-1, Stat-3, Akt (Thr308), AMPKα, mTOR, HSP27, p70 S6 kinase, p53, p38, SAPK/JNK, PARP cleaved, GSK-3β caspase 3, and Erk1/2 (all p > 0.05) (Fig. 7).

No difference was observed with regard to other intracellular survival pathways such as Stat-1, Stat-3, Akt (Thr308), AMPKα, mTOR, HSP27, p70 S6 kinase, p53, p38, SAPK/JNK, PARP cleaved, GSK-β, caspase 3, and Erk1/2. Data were normalized on the array positive control. O.D., optical density.
Discussion
Tissue microenvironment is essential to determine stem cell fate, where mutual interplay seems crucial for both physiological homeostatic maintenance and efficient tissue regeneration in case of injury (37). In myocardial response to damage, MSCs seem to play a pivotal role in the regeneration process because they are both able to exert a feeder-like support for CPCs and to directly participate through cell fusion with resident cardiomyocytes (40) or indirectly via paracrine action, releasing soluble factors (22).
Along with MSCs, CS-derived cells (CDCs) represent a further attractive cell source for cardiac regeneration, being able to improve cardiac function in the infarcted area when delivered into multiple animal models (28). CSs and CDCs promote cardiac regeneration by both direct and indirect mechanisms, that is, via cardiovascular differentiation and paracrine effects (7,29), these latter being also involved in MSC beneficial effects. Consequently, it is not inconceivable that the two cell populations (MSCs and CDCs) could cooperate if combined (56) or that one could influence the other, establishing an interplay mediated by the respective microenvironments, thus providing relevant biological advantages with regard to cardiac regeneration and function.
To date, the majority of studies are mainly focused on the paracrine action of MSCs on CPCs and tissue (19,30,38,54), indicating that MSCs can regulate cardiac stem cell niches (18,32). Nonetheless, nothing is reported about the opposite biological scenario (CPCs upon MSCs) and on the specific effects generated by CS-derived condition medium, particularly in view of the fact that cardiac tissue is mainly composed of mesenchyme, meaning that resident CPCs might undoubtedly act on the resident MSC pool in the heart. Only one recent study (43) supports the rationale that in vitro conditions could simulate the in vivo microenvironments, nevertheless employing human BMMSCs and not hmADMSCs and conditioned media derived from a non-stem cell cardiac population.
Novel tissue sources of MSCs are continuously discovered, such as the recently described mediastinal fat depots (41,47). Although authors suggest that the biological significance of hmADMSCs still remains unexplored, it is possible that due to its proximity to major organs such as the heart, mediastinal fat could play a role in some cardiac metabolic disorders (2,14,41,47).
Based on these assumptions, in this study we investigated the biology and cell plasticity of this novel and perhaps promising reservoir of MSCs, attempting to evaluate their potential commitment toward a cardiac-like phenotype and also to quantify the influence of the cardiac-committing microenvironment generated by CPCs with relation to phenotypical changes of MSCs. Cardiac differentiation of hADMSCs has been reported (8,24,34). Nonetheless, to the best of our knowledge, we have provided for the first time evidence of both an unexplored potential property of hmADMSCs and of the inductive efficacy of CSs via CCM on a “noncardiac” stem cell population. These experiments were performed without adding any differentiating agents in the medium, and specifically, we used primary CSs instead of CDCs.
We have investigated whether a short period of time (7 days) was sufficient to induce initial and/or partial cardiaclike differentiation phenomena. The preconditioning can be considered a sort of “priming,” aiming to enhance the proliferation and the differentiation of stem cell precursors within a hostile microenvironment as the one occurring during cardiac injury. In fact, over the years, it has been reported that pharmacological or genetic preconditioning prior to stem cell injection is able in some way to increase cell survival (12,16,20,39).
Our results show that CCM is able to significantly increase in vitro cell growth compared with FBS, thus confirming that CSs affect the mesenchyme through soluble factors, some of them displaying mitogenic properties. CSs as well as CDCs produce and release both in vitro and in vivo relevant amounts of many humoral factors, such as growth factors, cytokines and decoy/binding proteins (49). Particularly, primary CSs are able to release in vitro large amounts of vascular endothelial growth factor (VEGF), hepatocyte growth factor (HGF), and insulin-like growth factor-1 (IGF-1) (7), which mediate their prosurvival and proangiogenic properties, but they are also able to address cell differentiation toward the cardiomyogenic lineage (42).
Accordingly, our data suggest a link between CS paracrine action and their influence on MSC commitment, reflected in the molecular phenotypical changes that we observed (Fig. 3). Specifically, we found that the upregulated genes after preconditioning with CCM (vs. FBS) are the early nuclear cardiac markers Nkx2.5, essential for cardiac tissue differentiation and development (42), the cardiomyocyte-specific sarcomeric protein MHC (15) and KDR, required for the formation of the cardiovascular circuit (17). The downregulation of SMA, Thy-1, and c-kit could instead potentially indicate a propensity of the CCM, limited to hmADMSC population, to induce a non-smooth muscle differentiation (SMA) and a shift from an undifferentiated state (Thy-1) of hmADMSCs toward a more differentiated state, although sustaining lower mRNA levels of c-kit, whose positive expression has not been verified in hmADMSCs after isolation (41,47). Unmodified mRNA levels of GATA-4 and Cx43 have been observed in all conditions, indicating a nonspecific modulation of GATA-4 in our system and that a short period of conditioning does not probably allow upregulation of late cardiac markers, such as Cx43 (9). In addition, results suggest that none of the analyzed genes are modulated under cell culture conditions with FBS, or CGM basal medium, therefore indicating that biologically conditioned CCM could enhance preferential differentiation pathways.
To assess the specificity of the CCM effects observed, we used EDCs as a biological control. EDCs are collected from the primary explant culture and contain a subpopulation of CS-forming cells (6). Despite having been shown to possess some therapeutic potential (17), they represent an unselected mixed population, which has to go through CS formation and consequent functional maturation to obtain CPCs (1,13).
Interestingly, the treatment with EDCM resembles that with FBS, confirming that the two microenvironments generated by EDCs and CSs maintain distinct effects, consistent with the fact that they have been conditioned by two different cell populations.
A separate consideration has to be made for 5-azacytidine, a known cardiac transdifferention agent, which surprisingly seemed not to repetitively cause a cardiac transdifferentiation of hmADMSCs, at least in our system. The reason is certainly due to the huge interpatient variability of response to the treatment, and probably it may be associated with different methylation states of ADMSCs among patients. Our data confirm that 5-azacytidine is a controversial cardiac differentiation agent (27,44,55) and that it has not yet been proven to induce terminally differentiated cells (52). Adipose tissue-derived MSCs are able to differentiate into a cardiac phenotype even without 5-azacytidine induction (25). Besides, it is noteworthy to highlight that 5-azacytidine is considered a toxic chemical agent (although not toxic in our conditions as indicated by the MTS assay showing comparable proliferative rate to FBS; data not shown); thus, it cannot be considered an eligible candidate from a clinical standpoint. The remarkable difference between CCM and 5-azacytidine most likely relies on the consideration that 5-azacytidine is a single nonspecific chemical agent (thus further explaining the different responsiveness among patients), whereas CCM is the complex biological product of a primary human CPC population, which should mimic more faithfully the cardiac physiological microenvironment.
In accordance, interesting indications derive from intracellular signaling pathways. Each different treatment did not activate or induce definite protein phosphorylations or cleavage. In fact, growth factor-specific and survival signaling pathways such as Erk1/2, Akt (Ser308), p38, mTOR, and p53 are equally regulated in all treatments. Nevertheless, the timing and the modality by which some of them are early phosphorylated and/or sustained over time is more interesting. In particular, earlier phosphorylation of BAD on Ser112 occurs associated with the CS microenvironment (and to a lesser extent to that generated by EDC), whereas faster activation of PRAS40 in FBS and of Akt (Ser473) in EDCM and 5-azacytidine compared to CCM is found.
A variety of survival signals are reported to induce the phosphorylation of BAD at Ser112 by triggering its dissociation from Bcl-2/Bcl-X(L) (27) and thus protecting from apoptosis. Considering that the activation of BAD observed in our system follows a serum starvation, the soluble factors in the CCM allow a faster rescue from the serum deprivation-induced stress compared to other treatments.
So far, one of the main challenges of cardiac regenerative medicine still remains the optimization of protocols to induce putative “cardiogenic” gene expression programs in vitro. A considerable controversy exists on the type of reagents used in these protocols, ranging from cytokines (42), growth factors (48), to extracts of rat car-diomyocytes (15), nevertheless showing conflicting and marginal results (23,55). Moreover, regarding cardiaclike transdifferentiation of ADMSCs, the major limitation is the low yield.
In this regard, our report does not represent a new protocol to induce terminal cardiac-like transdifferentiation of ADMSCs nor does it show terminal cardiac differentiation (also given the short time of conditioning).
In conclusion, we have provided novel evidence of the potent inductive niche-like microenvironment that the CS structure can reproduce in vitro. Moreover, in our experimental conditions, the hmADMSCs themselves retain the distinctive plasticity of MSCs, such as the ability to partially commit/differentiate into a different lineage when subjected to a specific microenvironment.
Further in vitro and in vivo studies are certainly required to confirm not only our preliminary observations regarding the potential cardiac differentiation ability of hmADMSCs but moreover to possibly clarify whether other specific mechanisms are involved during this process. In addition, the interdonor variability observed in our system (data not shown) represents a further limitation of our study that needs to be addressed. Our data support the hypothesis that hmADMSCs might represent a useful tool to investigate the pluripotency of these cells and to be used for cardiac regenerative medicine research.
Footnotes
Acknowledgments
We acknowledge Maria Pia Docimo, Rosa Puca, and Fondazione Roma. We also thank Foto Ottica di Fabio Tonazzi for technical assistance. This work was supported by the grant University of Rome “Sapienza,” Ateneo 2010 prot. C26A10J5X4 to Prof. Giacomo Frati. C.S., I.C., C.N., G.M., G.S. performed the research; G.F., M.I., E.R. provided tissue samples; G.B.Z. performed statistical data analysis; A.C. contributed to reagents, tools, and interpretation of results; E.D.F. designed the research study; E.D.F. and G.F. wrote the article. The authors declare no conflicts of interest.
