Abstract
Direct reprogramming of an adult cell into another differentiated lineage—such as fibroblasts into neurons, cardiomyocytes, or blood cells—without passage through an undifferentiated pluripotent stage is a new area of research that has recently emerged alongside stem cell technology and induced pluripotent stem cell reprogramming; indeed, this avenue of investigation has begun to play a central role in basic biological research and regenerative medicine. Even though the field seems new, its origins go back to the 1980s when it was demonstrated that differentiated adult cells can be converted into another cell lineage through the overexpression of transcription factors, establishing mature cell plasticity. Here, we retrace transdifferentiation experiments from the discovery of master control genes to recent in vivo reprogramming of one somatic cell into another from the perspective of possible applications for the development of new therapeutic approaches for human diseases.
Introduction
Cell differentiation is a process by which cells become more specialized, acquiring new identities. Biologists often describe this process of development as proceeding from an undifferentiated (pluripotent) cell to a terminally differentiated cell that forms part of an adult tissue or organ. Different body parts arise in embryonic development because different, specific combinations of genes encoding transcription factors become activated (153). States of terminal cell differentiation are often considered to be fixed and irreversible, but in some cases, the balance of transcription factors can change and cells can inter-convert (151, 154). It is now known that the stable states of differentiated cells are controlled by dynamic mechanisms that can be perturbed under specific conditions, and an adult cell therefore can be reprogrammed, altering its pattern of gene expression—and thus its fate—to that typical of another cell type (136, 177).
Relevance of Reprogramming Adult Cells Into Another Phenotype
Understanding the stability of the differentiated state and the possibility of altering this state is interesting both from a basic biological perspective and for its potential to generate new research tools and cell sources for therapy. Final cell types of these reprogramming processes, progenitors and mature cells, may be directly used in applications such as cell replacement therapies.
Summary of Different Transdifferentiation Procedures Presented in the Literature
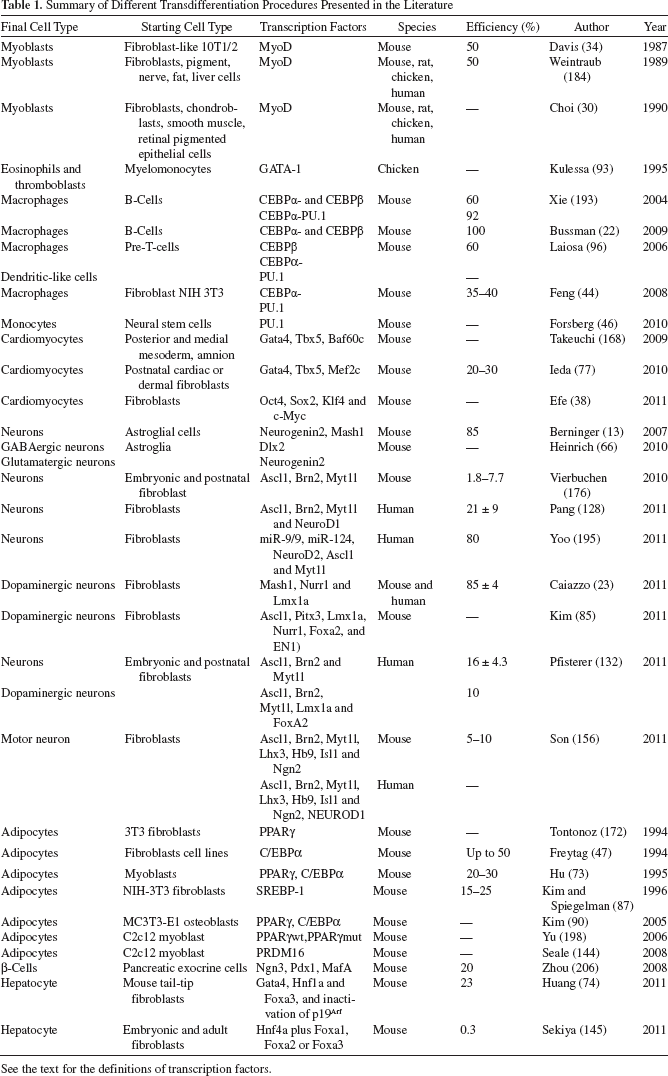
See the text for the definitions of transcription factors.
Reprogramming in Nature
Nature offers only rare examples in which cells of one type can be converted to another type in a process called cellular reprogramming (127). These events have been referred to as transdifferentiation or dedifferentiation (153).
Developmental biologist David Tosh has restricted the definition of transdifferentiation to irreversible switches of one differentiated cell type to another (12, 39, 120, 137, 146, 162). Metaplasia represents one example of natural transdifferentiation; it is the replacement in a tissue of one differentiated cell type with another or, more generally, transformation of one tissue type into another (149, 151, 152, 189). Metaplasias often occur in tissues that have been subjected to chronic trauma, damage, infection, or abnormal chronic stimulation (153).
Transdetermination is the name given to a form of metaplasia that involves transformations between different imaginal discs in Drosophila melanogaster. Occasionally, the disc will differentiate into a structure appropriate to another disc. It is a rare event involving a population of cells, and certain changes are more common than others, such as wing structures arising from the tissue of a leg disc (64).
Dedifferentiation is a cellular process in which a terminally differentiated and specialized cell reverts to an earlier, more embryonic developmental stage or to a simpler unspecialized form. It is often seen in more basal life forms such as worms and amphibians and also in plants (27, 57).
Important information about lineage reprogramming comes from dedifferentiation in regeneration, namely, the recreation of an adult structure by reactivation of embryonic genes that normally function during organ development (27, 152). Sometimes dedifferentiation and metaplasia result in an aberration of the normal development cycle and can have direct consequences. Indeed, they can predispose individuals to the development of cancer or be considered as the first step in a multistep progression to cancer (6, 43).
Reprogramming in the Laboratory
Scientists have long tried to reproduce natural reprogramming in laboratory experiments. This idea has been realized using three distinct experimental approaches to nuclear reprogramming: nuclear transfer, cell fusion, and transcription factor transduction. All of these experimental approaches have demonstrated that the differentiated state is not fixed but is instead actively maintained and reversible (17, 60, 167, 194).
Pioneering cloning studies in frogs in 1952 revealed that the transfer of nuclei from early blastocysts into enucleated oocytes resulted in cloned organisms (18). This experiment provided the first conclusive evidence that genes are not lost or permanently silenced during cell specialization; however, this finding was difficult to reproduce with donor cells from more specialized tissues (194).
In the 1960s, somatic cell nuclear transfer was achieved in which nuclei from differentiated adult cells were reprogrammed to a totipotent state after injection into enucleated eggs (61, 70). Using this approach, Wilmut and colleagues fused a mammary cell with an enucleated oocyte, and in 1997, successfully cloned a mammal for the first time, Dolly the sheep (190).
Cell fusion involves melding two or more cell types into one cell called a heterokaryon or hybrid (194). This fusion allows study of the effect of one genome on another, and in this way in the late 1960s, the existence of transacting repressors and tumor-suppressor proteins was discovered. Several decades ago, it was shown that silenced genes can be expressed again in differentiated cells by the fusion of different cell types (14–17). This approach provides the tools needed to clarify the regulatory mechanisms, such as DNA demethylation, that are required for nuclear reprogramming (194).
In 2006, Takahashi and Yamanaka described a groundbreaking work, reproduced and expanded by many other groups, showing that expression of key pluripotent transcription factors reprogram adult skin cells to a pluripotent state that resembles the embryonic stem cell (ESC) (113, 129, 166, 167, 186, 197). The authors identified four key factors that sufficed to induce pluripotency in fibroblasts—octamer-binding transcription factor 4 (OCT4), sex determining region Y box 2 (SOX2), Krüppel-like factor 4 (KLF4), and avian myelocytomatosis viral oncogene homolog (c-MYC), or as an alternative group, OCT4, SOX2, Homeobox protein NANOG, and cell line age 28 protein (LIN 28)—and designated these cells “induced pluripotent stem cells” (iPSCs) (36, 167). The iPSCs meet the original defining criteria for ESCs: they give rise to all three germ layers, form teratomas, and can form adult chimaeras and generate functional germ cells. Moreover, growth at clonal density demonstrated their self-renewal capacity (121, 129, 186). This finding opened the path to the possibility of generating patient-specific pluripotent cells for disease mechanism studies and therapy development.
In addition to pluripotent reprogramming, other reprogramming approaches more directly convert mature cells to progenitors or other mature cells without reversion to pluripotent cells. Below, we describe the transdifferentiation process by retracing experiments from the discovery of master control genes to in vivo reprogramming of one somatic cell into another and the underlying clinical relevance.
Master Control Genes
Twenty years ago, Lewis introduced the term “master control genes” for the homeotic genes of the Drosophila bithorax complex (103) that switches the fate of segments between alternative differentiation pathways. This observation revealed that the activation of only one or a few specific genes is responsible for determining differentiation in a specific cell type. Master control genes are either the first genes activated in a hierarchy or a transcription factor that activates a specific pathway. The expression of this group of factors is both necessary and sufficient to trigger activation of many other genes, leading to the development of a specific tissue or organ. Master control genes specify the body plan of animals and humans.
It was demonstrated first in D. melanogaster (50, 142) that the overexpression of a single master gene in somatic cells activates cohorts of genes typical of other somatic cell types, leading to important alterations in cell fate (194). Indeed, Gehring and colleagues reported that ectopic overexpression of the homeotic Antennapedia gene, under the control of a heat-shock gene promoter, results in a change in body plan in D. melanogaster larvae, inducing an additional set of legs in the antennal discs (142). In the years following this discovery, homeotic mutations in Drosophila led to the identification of several master control genes that specify the body plan, organizing anterior– posterior polarity, segmental identity, organogenesis, and identity of individual cells.
Subsequently, master genes have also been identified in mammals (34). In particular, in mice, Weintraub and colleagues induced a phenotypic conversion to the myogenic lineage by transfecting into fibroblast or adipoblast cell lines a single muscle gene, myogenic differentiation antigen (MyoD) (98, 124, 140, 169, 184). MyoD (169), a member of the basic helix–loop–helix family of myogenic regulatory, sequence-specific DNA-binding proteins, is a master regulatory gene for skeletal myogenesis and is expressed in skeletal muscle (40, 125, 183). This classical experiment is considered one of the most important landmarks in this field, clearly providing for the first time the idea that a single gene can induce cell fate conversion, even between distantly related somatic cell lineages.
Another outstanding example of a master control gene is paired box gene 6 (Pax6), one of the most analyzed among the Pax genes. Considerable evidence has defined Pax6 as a “master control” gene that plays a key role in the development of eyes and other sensory organs, certain neural and epidermal tissues during neurogenesis, and other homologous structures, usually derived from ectodermal tissues.
Pax6 encodes a transcription factor that contains a paired domain and a homeodomain. It was first cloned in mouse (69, 178) and in humans (171) and subsequently shown to be affected in the mouse mutant Small eye and in people with aniridia (50). Pax6 genes from various animal phyla can induce ectopic eye development on the legs, wings, and antennae of D. melanogaster, indicating that Pax6 is a master control gene for eye morphogenesis and evolution (50). Sometimes a single master control gene is not sufficient for reprogramming, but as shown by recent work with iPSCs, a group of transcription factors could be required to trigger the reprogramming of fibroblasts into pluripotent stem cells (84, 114, 129, 166, 167, 186, 197).
Transdifferentiation
Transdifferentiation is the reprogramming of one specialized cell type into another, without reversion to pluripotent cells. A primary goal of regenerative medicine is to convert existing adult cells from one cell type to another to provide a source of patient-specific cells. The first step is to identify key genes for transdifferentiation. Achieving this goal requires gene expression profiling of the target cell types for selection of signature factors that play a key role in the cells' determination and/or specification. Selected factors will be introduced in vitro or in vivo into other cell types, which then will be reprogrammed into the target cells for assessment of the genes' function in direct reprogramming. In vitro cultures allow testing of a large number of factors in parallel; on the other hand, in vivo systems are more demanding and provide context for the cells in their native environment.
Transdifferentiation in Mesodermal Derivatives
Transdifferentiation in Skeletal Muscle
In 1987, Davis and colleagues first identified a myogenic master control gene (34). It was known that myogenic, adipogenic, and chondrogenic clones can be derived from embryonic mouse fibroblasts after treatment with the DNA demethylating agent 5-azacytidine (170). The relative high efficiency of this conversion suggested that few key genomic loci are involved. At present, this reprogramming is attributed to demethylation of cytosine-phosphate-guanine (CpG) island promoters. Based on this evidence, Davis' group screened a myocyte cDNA library, isolating and testing a number of cDNA clones specific for myogenic cells and muscle tissue (34). They found that one of these cDNAs transfected into fibroblast-like 10T1/2 cells was sufficient to convert them to stable myoblasts (Fig. 1). Davis and colleagues cloned cDNAs into an expression vector under the control of the Moloney sarcoma virus long terminal repeat (LTR) with a simian virus 40 (SV40) poly(A) addition signal (11). Transfected cells were selected for G418 (antibiotic) resistance and fixed and scored for myosin heavy chain (MHC) expression with an alkaline phosphatase-linked immunostaining. About 50% of selected colonies were myosin-positive, with multinucleated syncytia, and morphologically indistinguishable from myogenic colonies derived by 5-azacytidine treatment of 10T1/2 cells. Furthermore, the team also confirmed myogenic conversion in other mouse fibroblast cell lines and in adipoblasts (34). In 1988, Tapscott et al. recognized that this gene is MyoD (169).
Reprogramming of fibroblasts. Fibroblasts can be converted into several adult cell types based on overexpression of different transcription factors. For an explanation of the panels and abbreviations, see the text.
A few years later, using the same viral vector, the same group reproduced the experiments with different cell lines and concluded that forced expression of MyoD can activate markers indicative of muscle differentiation in cell lines from each of the three germ layers (pigment, nerve, fat, liver, and fibroblasts) and of several species (mouse, rat, chicken, and human) (184). However, because chicken cells do not yield stable clones, a MyoD encoding an amphotropic retrovirus (LMDSN) was used (171), yielding multinucleated myotubes and 50% of cells intensely positive for MHC and desmin (184).
Based on the same transfection method and with the construct donated by Weintraub, in 1990 Choi confirmed the conversion by MyoD of dermal fibroblasts, chondroblasts, gizzard smooth muscle and retinal pigmented epithelial cells (30). The obtained myotubes presented the same characteristics of normal control day-10 embryonic breast muscles in terms of elongated morphology, number of nuclei, density of myofibrils, and the sarcomeric localization of antibodies to α-actinin, titin, nebulin, α-actin, troponin-I, tropomyosin, major histocompatibility complex, myosin light chain, myomesin, and desmin (30). Intriguingly, these myotubes contracted spontaneously. There is, however, an important difference with respect to the capacity for replication of myogenic cells in primary control cultures and of myogenic cells generated in MyoD-converted cultures (30). It is likely that the overexpression of MyoD limited proliferation capacity (34, 105), as Davis et al. previously reported (34).
Muscular dystrophies, including Duchenne muscular dystrophy (DMD) and Becker's muscular dystrophy (BMD), are genetic, progressive degenerative disorders of skeletal muscles, without any effective cure. Cell transplantation of donor muscle satellite cells as a therapeutic strategy both in DMD patients and in BMD patients has been unsuccessful, mainly because of immune rejection, poor survival, and limited diffusion after local intramuscular injection (114, 134). The use of genetically modified autologous myogenic cells should avoid most of the immune rejection problems, even if immunogenicity remains a critical open question. In theory, because iPSCs can be generated from a patient's own somatic cells, they will be autologous. However, Zhao et al. recently reported that mouse iPSCs were rejected with massive cluster of differentiation 4 positive (CD4+) T-cell infiltration even when the transplantation was performed in a syngeneic mouse (204). This important finding suggested that further investigation is needed to confirm and understand the immunogenicity. In fact, the reaction could be a response against undifferentiated cells that form a teratoma, while the immunogenicity of iPSC-derived differentiated cells has not yet been assessed (122). Subtle epigenetic changes, genomic mutation, and/or other modification occurring during transdifferentiation and prolonged culture of reprogrammed cells could have important consequences for immunity response and must be experimentally addressed. Both ESCs and iPSCs would be expected to develop mutations in their genomes during the cultivation process, but if the immunogenicity of such cells is weak, we should be able to control the immune responses of the recipient through a proportionally reduced immunosuppressant treatment. However, the use of ex vivo genetically corrected myogenic cells is seriously limited by the number of primary myogenic cells that can be isolated and transplanted into the patient's muscles. Therefore, fibroblasts converted to myoblasts would represent an alternative, abundant source of autologous myogenic cells, avoiding immune rejection. In fact, fibroblasts are not affected in their growth capacity in dystrophies and can be easily obtained from a single biopsy and expanded in vitro without significant limitation. The converted myogenic cells could then, in theory, be corrected in their defective gene ex vivo and reimplanted into the patients.
Several groups have explored the possibility of using the MyoD gene to induce myogenic conversion of nonmuscle, primary cells in a quantitatively relevant way (175). In one study, primary human and murine fibroblasts from skin, muscle, or bone marrow were infected by an E1-deleted adenoviral vector carrying a retroviral LTR-promoted MyoD cDNA. MyoD-converted cultures were injected into regenerating muscle of immunodeficient mice, where they gave rise to new muscle fibers, supporting the feasibility of this approach (99). To modulate more precisely the fibroblast–myogenic conversion, this group previously described the feasibility of using an inducible promoter to switch on MyoD on demand (35). The strategy of performing transplantation with myogenic cells converted from fibroblasts is hampered by the comparatively lower engraftment efficiency relative to that achieved using primary myogenic cell-derived skeletal muscle cells.
Modulating the sequential expression of one or more muscle master gene transcription factors like Pax3/7, gene myogenic factor 5 (Myf-5), and MyoD or others would be of interest to achieve a myogenic cell phenotype, like mesoangioblasts (165) or CD133+ cells (173), that has a more favorable behavior in a cell transplantation setting; these cell types may also allow for the possibility of systemic endovascular administration. One futuristic idea is to achieve the direct in vivo conversion of fibro-adipose tissue that infiltrates dystrophic muscles with a regulatable expression of specific myogenic and nonmyogenic transcription factors with or without administration of the defective gene (like wild-type full-length dystrophin or mini-dystrophin) by gene therapy or contemporaneous gene correction with other strategies like exon skipping.
Transdifferentiation in Heart Tissue
Despite a robust understanding of the transcriptional regulation of the developing heart (126, 160), no single master control gene like MyoD for skeletal muscle (34) was identified as the activator of the cardiac gene program de novo in mammalian cells or tissues until 2009. In fact, in that year, Takeuchi showed that a group of master control genes, two cardiac transcription factors, guanine-adenine-thymine-adenine (GATA) binding protein 4 (Gata4) and t-box transcription factor 5 (Tbx5), and a cardiac-specific subunit of the polymorphic SWItch/Sucrose NonFermentable (Swi/Snf)-like brahma-related gene 1 [Brg1]/brahma homologue [Brm]-associated factor (BAF) chromatin remodeling complexes, Baf60c, can directly reprogram mouse mesoderm into beating cardiomyocytes (168). Baf60c, a cardiac-enriched BAF complex subunit, has important roles in cell-type specification and differentiation (97, 102, 104, 150, 192), linking DNA-binding transcription factors to BAF complexes. Gata4, a transcription factor gene expressed in the developing embryonic heart, is essential for normal cardiac patterning and vascularization (33, 71, 94, 118, 135, 200, 206), whereas Tbx5 has been implicated in cardiac morphogenesis (21, 28). Induction of cardiac differentiation was assessed by expression of the early cardiac marker gene actin, a cardiac muscle 1 (known as Actc1), gene myosin, light chain 2, regulatory (known as Myl2) messenger RNA, tropomyosin, and cardiac troponin T (cTnT) proteins, specific markers of the embryonic heart. Cotransfection was performed also in the posterior and medial mesoderm, which are normally not fated to cardiac lineages, and in the amnion, resulting in ectopic cTnT and beating tissue induction. Gata4 with Baf60c initiated ectopic cardiac gene expression; in particular, Baf60c is required to allow Gata4 to bind cardiac genes in noncardiac mesoderm. Addition of Tbx5 allowed differentiation into contracting cardiomyocytes and repression of noncardiac mesodermal genes (168).
Recently, Ieda and colleagues examined a total of 14 key cardiac regulators, and after a systematic process of progressive elimination, they defined a minimal combination of three developmental transcription factors, Gata4, myocyte enhancer factor 2c (Mef2c), and Tbx5, that rapidly and efficiently reprogrammed postnatal cardiac or dermal fibroblasts directly into differentiated cardiomyocyte-like cells (77) (Fig. 1). To select candidate reprogramming factors, they focused on genes that are expressed at high levels in cardiomyocytes with respect to cardiac fibroblasts and that caused developmental defects in cardiac tissue when mutated in mice (77). Cells were transduced by retroviruses or lentiviruses. The induced cardiomyocytes were similar, even identical, to neonatal cardiomyocytes in global gene expression profile and markers of cardiac fibroblasts, suggesting that global transcriptional and protein profile reprogramming occurred. The authors also demonstrated that fibroblasts were epigenetically reprogrammed into cardiomyocytes by comparing active and repressive histone modifications and DNA methylation in the promoter region of a small set of cardiac-specific genes. Gata4 might open chromatin structure in cardiac loci (31), thus allowing binding of Mef2c and Tbx5 to their specific target sites and leading to full activation of the cardiac program (77).
Intriguingly, trimethylated H3 on lysine (K)-27 (H3K27) levels were higher in tail-tip fibroblasts than in cardiac fibroblasts, suggesting that fibroblasts from different tissues might have different propensity to specific lineage reprogramming. Indeed, cells were electrophysiologically active and could contract spontaneously, with a phenotype similar to ventricular cells. Further investigations are needed to determine whether other types of cardiomyocytes, such as cells of the atrium or sinus node, can be generated directly from fibroblasts, for instance adding different reprogramming factors. The authors provided also some insights on the behavior of the reprogrammed cardiomyocytes in vivo. They transplanted cardiac fibroblasts 1 day after viral transduction and injected them into nonobese diabetic severe combined immunodeficient (NOD-SCID) mouse hearts, demonstrating that grafted cells completed their cardiac differentiation in vivo. These findings warrant further investigation for the development of cell repair strategies in cardiac diseases (77).
A completely different approach was described by Efe et al., who obtained from fibroblasts spontaneously contracting patches of differentiated cardiomyocytes over a period of 11–12 days using four transcription factors (Oct4, Sox2, Klf4, and c-Myc) (38). After transfection, fibroblasts were cultured in leukemia inhibitory factor-free medium containing 5% fetal bovine serum, small molecules that might enhance cardiogenesis, and the small-molecule inhibitor of the Janus kinase/signal transducer and activator of transcription (JAK-STAT) pathway. Given the fact that four iPSC reprogramming factors were used to directly reprogram cardiomyocytes from mouse iPSCs, this method likely took advantage of the rapid mouse iPSC conversion and fast cardiomyocyte differentiation from mouse iPSCs. A detailed mechanism underlying this method must be carefully dissected and confirmed with human fibroblasts (38).
Overall, these experiments provided evidence that the fibroblast-to-cardiac conversion occurred directly, without passing through a cardiac progenitor-like state or a pluripotent state. This reprogramming potential has implications for human health. Heart disease is the leading cause of adult and childhood mortality and morbidity in the western world. These diseases consist of heart failure, typically the loss of cardiomyocytes, or congenital heart malformations, an improper development of cardiomyocytes during embryogenesis. The heart has little or no regenerative capacity after damage; current therapeutic approaches are limited due to the lack of a renewable source of cardiomyocytes. The human heart is composed of cardiomyocytes, vascular cells, and cardiac fibroblasts. The latter comprises over 50% of the whole cells in the heart (10, 24, 155), representing a potential source of beating cardiomyocytes for regenerative therapy if it was possible to reprogram them directly. Such a reprogramming would have many therapeutic implications; in fact, large amounts of cardiomyocytes can be obtained from a single biopsy and could eventually be used for transplantation in damaged hearts. A major goal is to introduce defined factors into the heart, directly converting the endogenous fibroblasts in vivo.
Transdifferentiation in Immunosystem and Hematopoietic Cells
Hematopoietic differentiation initiates when a multipotent hematopoietic progenitor differentiates to form a common lymphoid and a common myeloid progenitor, followed by a process of binary decisions, resulting in committed cells of various lineages (1, 2, 42, 115).
Each lineage exhibits a distinct gene expression pattern regulated by a set of several lineage-restricted transcription factors that establish new gene expression programs during differentiation while suppressing old ones (25, 41, 54, 125, 147, 148). Among these transcription factors, GATA-1 has emerged as a candidate regulator of hematopoietic cell differentiation. GATA-1 was originally identified as an erythroid-specific factor involved in globin gene regulation (93, 95) and expressed in megakaryocytes, mast cells, and eosinophils (108, 109, 138, 159, 208). Using this evidence as a starting point, together with the finding that GATA-1 is highly expressed in thrombocytic and erythroid progenitors and completely downregulated during myelomonocytic differentiation (56), Kulessa and colleagues investigated the role of GATA-1 in lineage determination by forcing its expression in v-myb myeloblastosis viral oncogene homolog-, v-ets erythroblastosis virus E26 oncogene homolog (Myb-Ets)-, and v-Myc-transformed chicken myelomonocytic cell lines (Fig. 2). Cells electroporated with the retroviral expression vector pneomicin(NEO)-GATA-1 were converted into eosinophils and thromboblasts, as indicated by the expression of markers specific to eosinophilic cells and thrombocytic cells, such as eosinophil-specific cell surface antigen 47 (EOS47) or multipotent hematopoietic precursors 21 (MEP21), and by the change in morphology and mRNA expression (93). A correlation between the GATA-1 expression level and the induced phenotype was demonstrated as the suppression of myelomonocytic markers in myeloblasts caused by GATA-1 activation (93). GATA-1, therefore, resembles the MyoD family of transcription factors (34). However, Kulessa surmised that other transcription factors such as the cytidine-cytidine-adenosine-adenosine-thymidine (CCAAT)/enhancer binding protein (C/EBP) may have a role in hematopoietic differentiation (93).
Transdifferentiation in the immune system. (A) Reprogramming of monocytes into eosinophils through guanine-adenine-thymine-adenine (GATA) binding protein 1 (Gata-1) overexpression (93); (B) forced expression of cytidine-cytidine-adenosine-adenosine-thymidine (CCAAT)/enhancer binding protein (C/EBPα) and purine-rich (PU) box binding protein 1 (PU.1) in fibroblasts induces the formation of macrophages (44, 193), whereas the overexpression of C/EBPα alone in B-cells and T-cells induces the conversion into macrophages (22, 193); (C) transdifferentiation of pre-T-cells into dendritic cells through the overexpression of PU.1 (96); (D) acquisition of monocytes through overexpression of PU.1 in neural stem cells (46).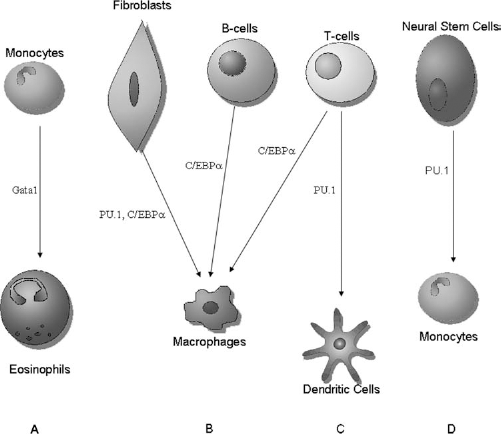
Xie and colleagues confirmed this hypothesis in 2004 (193). In particular, their study reported that enforced expression of C/EBPα and C/EBPβ in differentiated B-cells leads to their rapid and efficient reprogramming into macrophages (193) (Fig. 2). C/EBPα is a basic region-leucine zipper (bZip) family transcription factor, strictly required for the development of granulocyte and macrophage precursors (201–203). In addition, the overexpression of this factor induces myelomonocytic differentiation in human multipotent progenitors (78) and in committed B-cell precursors (72, 193). The related family member C/EBPβ is also expressed in myelomonocytic cells and eosinophils and can rescue granulocyte formation in C/EBPα-defective mice (81). Xie's group used a retroviral approach to express the myeloid transcription factors C/EBPα and C/EBPβ in primary B-cell precursors derived from C57Bl/6J mice bone marrow. Infected cells downregulated CD19 and upregulated Mac-1 (CD11b), with ~60% of the cells becoming CD19- Mac-1+ after 4 days (191). Their similarity to macrophages was demonstrated by morphological changes, as the reprogrammed cells were large and contained granules, and by their phagocytic capacity (54).
Bussmann et al. repeated this result with higher efficiency (100%) within 2–3 days using an estradiol-inducible form of C/EBPα in the absence of stroma (22). In the Xie et al. experiments, gene profiling indicated that C/EBPα and β reprogram B-cells into macrophages through a series of sequential changes that require endogenous purine-rich(PU) box binding protein 1 (PU.1) (193). This transcription factor of the Ets family is required for the development of myelomonocytic cells (183, 185, 202), dendritic cells, B-cells, and T-cells (4, 58, 112, 143, 157) and repre-sents another candidate gene with a key role for lymphoid and myeloid commitment.
Xie's group suggested that C/EBP nonresponder cells may express low levels of endogenous PU.1 and subsequently coinfected these cells with both C/EBPα and PU.1 viruses, reaching 92% efficiency (193). Despite the fact that each lineage, lymphoid and myeloid, exhibits a distinct gene expression, the Xie study reported for the first time the transdifferentiation from mature lymphoid cells to myeloid cells.
Laiosa et al. also demonstrated this specific conversion, finding that C/EBPα and C/EBPβ induce the formation of inflammatory macrophages from pre-T-cells whereas PU.1 reprograms them into myeloid dendritic-like cells (95) (Fig. 2). Pre-T-cells from lck ancestry mice were infected with retroviruses containing genes encoding C/EBPα, C/EBPβ, and PU.1. The obtained cells were analyzed by fluorescence-activated cell sorting: cells induced by C/EBPα and C/EBPβ were Mac-1+Gr-1+CD62L+CD11c– MHCII–F4/80–, a phenotype characteristic of inflammatory macrophages (51); cells generated by PU.1 infection were Mac-1+Gr-1–CD62L–CD11c+MHCII+F4/80lo, resembling myeloid dendritic cells (147). Cell morphology also confirmed these findings. Approximately 60% of the C/EBPα- and C/EBPβ-infected pre-T-cells were reprogrammed into macrophages that were functional and able to internalize opsonized sheep red blood cells. Semiquantitative RT-PCR showed that at day 9, expression of all myeloid genes tested was evident in both C/EBP-infected samples, whereas expression of all T-cell genes tested had become undetectable. Furthermore, Notch was identified as an important gene, providing a direct link between the transcription factor network and extracellular signals (96).
All of the studies described so far concerned the conversion of committed hematopoietic cells into another blood cell type that was closely developmentally related, probably sharing transcriptional networks and chromatin configurations. However, in 2008, Feng and colleagues obtained macrophage-like cells from a cell type distantly related to blood cells, fibroblast cells (44) (Fig. 2). Based on the unexpected finding that PU.1 and C/EBPα upregulate Mac-1 expression in fibroblasts (193), Feng's group investigated whether their expression is sufficient to activate a myeloid program in a fibroblast cell line as well as in primary embryonic and adult fibroblasts (44) (Fig. 1).
NIH 3T3 cells were coinfected with PU.1 and C/EBPα viruses tagged with green fluorescent protein (GFP) and human CD4, respectively. PU.1 or C/EBPα alone induced the upregulation of the myelomonocytic marker Mac-1 and the pan-hematopoietic marker CD45 in 35–40% of the infected cells after 5 days, whereas their coexpression increased the number of Mac-1+ cells to 90%. The result was also confirmed in induced primary fibroblasts and in cultures prepared from mouse embryos, as well as in a skin biopsy of an adult mouse, although with lower efficiency. Converted cells more closely resembled a macro phage cell line than the original fibroblast cells in terms of morphology, phagocytic capacity, and partial inflammatory response. The gene expression profile validated that the converted cell lines were macrophage-like cells, exhibiting upregulated macrophage-restricted genes and downregulated fibroblast-associated genes. However, the reprogrammed fibroblasts of Feng et al. represented intermediates that required the continuous and exogenous expression of PU.1 and C/EBPα to maintain a macrophage phenotype (44).
After all of these studies in the hematopoietic field, the specific transcription factor PU.1 emerged as crucial. Indeed, recently, Forsberg's group reported that adult neural stem cells can be reprogrammed to monocytes at very high efficiency by ectopic expression of the Ets transcription factor PU.1 (46) (Fig. 2). Neural stem cells were derived from the adult mouse forebrain, cultured as neurospheres, and transduced with a lentiviral control vector containing only GFP or a vector bicistronically expressing hemagglutinin (HA)-tagged PU.1 and GFP. High-efficiency reprogrammed neural stem cells resembled monocytes in marker profile, morphology, and chemotactic response. The reprogramming capacity appears to be unique to neural stem cells; indeed, PU.1 is insufficient to efficiently reprogram fibroblasts, which are more distantly related mesodermal cells (46). Although neural stem cells are even more distantly related to the hematopoietic lineage, they are more amenable to reprogramming thanks to their plasticity; they express some of the reprogramming factors, such as Sox2 and c-Myc (88), at high basal levels, which may enhance their ability to modify their fate in response to extracellular signals.
The speed and efficiency with which hematopoietic cells can be reprogrammed by C/EBP and PU.1 led to the question of whether this plasticity can be relevant in pathological conditions, such as leukemia. Some myeloid leukemias show immunoglobulin gene rearrangements (116), and Hodgkin's lymphoma cells express some genes specific to monocytic and dendritic cells and contain immunoglobulin rearrangements and somatic hypermutations (95).
A recent interesting study from Szabo et al. described the direct conversion of human dermal fibroblasts into multilineage blood progenitors (164). The latter were able to differentiate into cells expressing markers of granulocytic, monocytic, megakaryocytic, and erythroid lineages. The induction of hematopoietic lineage phenotype was promoted by the expression of the pluripotency key gene OCT4 in combination with specific culture conditions permissive for hematopoiesis. This result was unexpected because OCT4 is a transcription factor regulating pluripotency not expressed in the hematopoietic system. Indeed, in these specific experimental conditions, OCT4 did not induce pluripotency but rather promoted a specific lineage commitment. Future studies may clarify whether Oct4 mimics other hematopoietic Pou domain-containing transcription factors, such as Oct1 or Oct2, or may induce some partially reprogrammed cells that can subsequently be differentiated into blood lineages with appropriate environmental signals. Indeed, the authors provided a remarkable example of a reasonable basis for autologous cell replacement therapies. In fact, they demonstrated the in vivo engraftment capacity of obtained multipotent blood progenitors. Reprogrammed cells engrafted in all transplanted recipients up to a level of 20% (164). Knowledge about master genes involved in hematopoietic differentiation may be useful for converting fibroblasts or other specific patient cells into hematopoietic cells that may be a source for autologous transplantation.
Transdifferentiation in Adipocytes
Adipocyte differentiation is a complex process that involves dramatic changes in cell morphology and gene expression regulated mainly by a cascade of transcription factors (20). In recent years, scientists have identified potential regulators of this process. In adipocytes, differentiation appears to be controlled by three major factors or groups of factors: C/EBPα, peroxisome proliferator-activated receptor (PPAR)γ, and sterol regulatory element-binding protein-1 (SREBP-1). As noted, C/EBPs are members of the basic leucine zipper transcription factor family (111). Three C/EBP isoforms, C/EBPα, C/EBPβ, and C/EBPδ, are expressed at high levels during adipocyte differentiation (26); the best characterized, C/EBPα, plays an important role in terminal adipocyte differentiation (106, 202, 203). PPARγ proteins are members of the nuclear hormone receptor superfamily that are expressed primarily in adipocytes and play a crucial role in the activation of genes involved in very early adipocyte differentiation (7, 9, 106, 158, 200). SREBP-1 has been identified as a proadipogenic basic helix–loop–helix transcription factor, expressed in particular in the early phase of mouse adipocyte differentiation (71, 87).
Tontonoz et al. (172) and Freytag et al. (47) provided the first evidence of the C/EBPα and PPARγ key role in 1994. Tontonoz et al. expressed PPARγ ectopically in fibroblasts 3T3 using the Moloney murine leukemia retrovirus (MMLV) (119). Transfected cells, cultured under permissive conditions for adipogenesis, differentiated into lipid-containing cells that morphologically resembled cultured adipocytes and expressed typical adipocyte-linked differentiation genes (Fig. 1). PPARγ exerts its adipogenic effects probably through the transcriptional activation of genes required for the adipocyte differentiation program or through interaction with other proteins like retinoid X receptor (172).
Freytag's group transduced the C/EBPα gene into mouse fibroblastic cell lines by retroviruses, generating adipocyte colonies at variable frequencies (47) (Fig. 1). The most efficient result was obtained with NIH 3T3 cells (>50%). One year later, in 1995, Hu and colleagues showed that adipocytes can be obtained from a related but highly specialized cell type, myoblasts, by transfection with the same two transcription factors used by Tontonoz et al. and Freytag et al. (47, 73, 172).
In 1996, Kim and Spiegelman demonstrated adipocyte reprogramming using another factor, SREBP-1 (86) (Fig. 1). Recent studies have reported obtaining adipocytes from osteoblasts and myocytes (198). In fact, PPARγ acts also as a molecular switch between osteogenic and adipogenic pathways (3, 80, 101). In differentiation medium, ectopic expression of this transcription factor, together with C/EBPα, has led to differentiation of osteoblasts into adipocytes (89). To generate osteoblastic cells (MC3T3-E1) stably overexpressing PPARc2 and/or C/EBPα, these cells were transduced with a respective retrovirus. Foci of adipocytes arose from day 4 after treatment both with PPARγ and C/EBPα-PPARγ; however, the lipid droplet size remained smaller in cells transfected with PPARγ compared with the lipid droplets obtained with C/EBPα-PPARγ transfection (89).
As reported above in 1995 (73), myoblasts can be converted into adipocytes with retroviral transfection; in 2006, another group confirmed the result, overexpressing only a plasmid carrying wild-type PPARγ or mutated PPARγ (serine 112 was mutated to alanine) in mouse myoblast cells (7). Transfected cells expressed adipogenic marker genes such as adipocyte fatty acid binding protein, lipoprotein lipase, and glycerol-3 phosphate dehydrogenase and showed increased adipogenic gene expression. The ability to transdifferentiate was greater in cells with mutated PPARγ than in cells expressing wild-type PPARγ (194).
The central role of PPARγ was confirmed by Seale and colleagues (144). They showed that the C2C12 myoblast cell line and myoblasts isolated from postnatal skeletal muscle and transduced with gene PR domain containing 16 (PRDM16)-expressing retroviruses then transdifferentiated into lipid-filled adipocytes using adipogenic inducers. In the obtained adipocytes, the myotube-specific genes such as myoD, myogenin, myogenic regulatory factor 4 (mrf4), MHC, and myosin light chain kinase were blocked, whereas adipocyte-specific genes, including PPARγ and adipocyte protein 2/fatty acid binding protein 4 (aP2/FABP4), were upregulated. Moreover, PPARγ was found in the PRDM16 complex and was demonstrated to interact with and be coactivated by PRDM16. PRDM16 was unable to promote adipogenesis in PPARγ-deficient fibroblasts, indicating that PPARγ activation is essential (144). All of these studies together demonstrated that PPARγ is the master control gene of adipogenic differentiation (8, 139, 172).
Human lipodystrophies represent a heterogeneous group of diseases characterized by generalized or partial adipose tissue deficiency (49). They result from either the failure of adipocyte development or the immune-mediated premature destruction of adipocytes; moreover, the remaining mature adipocytes appear to be dysfunctional. The immediate consequences of this disorder include abnormalities in energy storage, postprandial lipid buffering, and adipokine production. Adipocytes obtained through fibroblast transdifferentiation can be useful as a source of fat cells that can be grafted into lipodystrophic areas. Currently, the fat used for fat transfer is extracted from part of the patient's body and injected into another area that requires it. However, it can be difficult to detect fat areas in patients, and reprogrammed fibroblasts can represent a promising alternative.
Transdifferentiation in Ectodermal Derivatives: Neurons
Several neuronal-specific transcription genes are involved in neuronal commitment from stem/precursor cells during development and thus can be potential candidate genes in transdifferentiation experimental approaches.
As we mentioned above, Pax6 is one of the master genes responsible for neuronal differentiation; indeed, it is expressed in radial glia of the developing cortex (50), and it is indispensable for cortical neurogenesis from these cells (68, 182). Moreover, Pax6 plays a crucial role in generation of neurons from astroglial stem cells in the adult subependymal zone (53, 62). Postnatal astroglia can be redirected toward neurogenesis by forced expression of Pax6 in vitro (68).
In recent years, many efforts have been made in the field of neuronal reprogramming. Berninger and colleagues showed that the proneural genes neurogenin-2 (Ngn2) and mammalian achaete scute homolog 1 (Mash1) can reprogram astroglial cells from early postnatal cerebral cortex into neurons (13). These factors were chosen because they can drive a glutamatergic versus γ-aminobutyric acid (GABA)ergic fate in precursors of the developing forebrain (59) as well as in neural stem cells from the adult subependymal zone in vitro (13). Postnatal astroglia were transduced using vesicular stomatitis virus G protein pseudotyped retroviruses containing a Pax6–internal ribosomal entry site (IRES)–GFP (63), Ngn2–IRES– GFP, or achaete-scute complex homolog 1 (Mash1)– IRES–GFP (13, 65). The efficiency of the method was ~85%. After 12 days, converted cells expressed proteins characteristic of immature neurons, such as class III β-tubulin (68), and differentiated into excitable cells as demonstrated by electrophysiological characterization. However, astroglia-derived neurons showed no spontaneous or evoked synaptic activity, a severe limitation toward functional neurogenesis, probably because of a delay in the maturation process or an incomplete reprogramming. Moreover, given the lack of functional presynaptic output, the neuronal subtype generated could not be defined (13).
A few years after this work, Heinrich et al. tried for a stronger and more persistent expression of neurogenic fate determinants for a more complete reprogramming of astroglia toward synapse-forming neurons (66). The LTR-driven MMLV-derived retroviral constructs that Berninger's group had used were expressed only about two- to threefold compared to endogenous expression (67, 68) and were prone to silencing (48). Thus, Ngn2 or gene distal-less homeobox 2 (Dlx2) were expressed under the control of an internal chicken β-actin promoter with a cytomegalovirus enhancer optimized for long-term expression over months in the adult mouse brain together with DsRed located behind an IRES (201). Postnatal cortical astroglia transduced with Ngn2, which directs the generation of exclusively excitatory glutamatergic neurons in the dorsal telencephalon, differentiated into class III β-tubulin-positive, glial fibrillary acidic proteinnegative synapse-forming glutamatergic neurons after 10 days in culture with high efficiency. The group also tested another transcription factor, the homeobox Dlx2, which is involved in GABAergic neuron specification in the developing ventral telencephalon (131) and in adult neurogenesis (19). Forced expression of Dlx2 in postnatal cortical astroglia gave rise to cells with a neuronal identity as revealed by class III β-tubulin or microtubule-associated protein 2 (MAP2) expression; however, the efficacy of neurogenesis was significantly lower than that obtained with Ngn2. Patch-clamp recordings revealed that Dlx2 induced a GABAergic identity in the reprogrammed astroglia, even if Dlx2-induced reprogramming remained partial in most of the cells. Coexpression of Dlx2 and Mash1, a transcription factor located upstream of Dlx2 in the interneuron fate specification (133), produced a higher proportion of transfected neurons exhibiting specific interneuronal firing patterns compared to Dlx2 alone, indicating that these two factors indeed act synergistically. Moreover, the efficiency of reprogramming toward a GABAergic neuronal subtype was increased, expanding postnatal cortical astroglia under neurosphere conditions. Finally, it was demonstrated that Ngn2 or Dlx2 overexpression can also be reprogrammed into functional neurons from reactive astroglia from adult cortex following injury after prior expansion as neurospheres (66).
The direct relationship between the reprogramming factors used and the neurotransmitter phenotype is very close to the physiological events occurring during the development that are likely recapitulated in this process. Indeed, the reprogramming occurred without cell division as in the vast majority of direct differentiation experiments.
These studies can open the path to the direct in vivo conversion of astrocytes into the selected desired neuronal populations for the treatment of neurological diseases.
In the same year (2010), Vierbuchen and colleagues obtained ectodermal neurons from mesodermal murine fibroblasts, a distantly related cell type (176) (Fig. 1). Starting from a pool of 19 candidate genes, by progressive elimination, they identified a combination of three factors, achaete-scute homolog 1 (Ascl1), brain-specific homeobox/POU domain protein 2 (Brn2), and gene myelin transcription factor 1-like (Myt1l) (BAM), that suffice to rapidly and efficiently convert mouse embryonic and postnatal fibroblasts into functional neurons in vitro. Ascl1 is a master gene in neural development that regulates neuronal versus glial specification and promotes neuronal phenotype. Mouse embryonic fibroblasts (MEFs) from TauEGFP knock-in mice, which express GFP specifically in neurons (174, 186), were infected with lentiviruses containing selected genes under the control of the tetracycline operator (185). Fresh MEF media containing doxycycline was added after 16–20 h in media containing lentivirus to activate expression of the transduced genes. These induced neuronal cells (iN) expressed multiple neuron-specific proteins, like microtubule-associated protein 2 (MAP2) and β-tubulin, generated action potentials, and formed functional synapses, thus showing authentic neuronal functions. An interesting aspect was that the majority of obtained neuronal cells was excitatory (glutamatergic) and expressed markers of cortical identity. The conversion was rapid and efficient; after only 5 days, neuronal cells with long, branching processes were readily detected, and the efficiency ranged from 1.8% to 7.7%. The single factor Ascl1 was sufficient for immature neuronal feature induction, and in fact, the omission of Ascl1 had a dramatic effect on efficiency. To generate mature neuronal cells, the additional expression of Brn2 and Myt1l is needed (176); Brn2 is expressed in cortical development and might play a role in the predominant glutamatergic phenotype observed.
Recently, the same group (128) showed that human fetal fibroblasts also could be directly converted into neurons through an infection with the same transcription factors: Brn2, Ascl1, and Myt1l (BAM). However, the BAM factors induced neuronal features but did not generate functional neurons. Only with the addition of neural differentiation 1 (NeuroD1) (BAMN), another basic helix–loop–helix transcription factor, was the efficiency improved. Indeed, they obtained cells with neuronal morphologies, labeled positive for pan-neuronal markers [class III β-tubulin, neuronal nuclei (NeuN), polysialic acid-neural cell adhesion molecule 1 (PSA-NCAM), and MAP2] and with active membrane properties (128).
In addition to the use of transcription factors, the possibility of using microRNAs (miRNAs) has recently been suggested for promoting the reprogramming and transdifferentiation process without the use of viral vectors. Using miRNAs makes the generation of iPSCs more efficient (5), suggesting that downregulation in addition to upregulation of specific genes can modulate the reprogramming.
Based on their previous studies showing that miR-9 and miR-124 critically regulated neuronal differentiation, Yoo et al. recently demonstrated that their forced expression in human fibroblasts induced reprogramming into neurons (195). The obtained cells showed a reduction in proliferation and neuron-like morphologies, but only a low percentage (less than 5%) was MAP2 positive. To increase the percentage of neuronal cells, they infected fibroblasts with miR-9/9*-124 together with NeuroD2, Ascl1, and Myt1l and found approximately 10% of the initially plated cells positive for MAP2 with an extensive neurite outgrowth as shown by β-III tubulin staining. Moreover, the obtained cells could fire repetitive specific action potentials and presented glutamatergic and GABAergic phenotypes (195). The exploration of miR-NAs as a tool for other transdifferentiation applications warrants further investigation.
In a complementary way, transient induction of the four iPSC reprogramming factors (Oct4, Sox2, Klf4, and c-Myc) can directly differentiate murine fibroblasts into functional neural progenitor cells (NPCs), in association with specific NPC culture conditions (90). Compared with induced neurons, transdifferentiated NPCs present the advantage of being expandable in vitro and retaining the ability to give rise to multiple neuronal subtypes and glial cells. These results point out the potential utility of iPSC-based reprogramming factors as a tool not only for full stem reprogramming but also for transdifferentiation.
Of interest, the transdifferentiation strategy also can be successfully applied to achieve specific neuronal populations with clinical meaning, such as the dopaminergic phenotype. In fact, a combination of three transcription factors—Mash1, nuclear receptor related 1 protein (Nurr1), and LIM homeobox transcription factor 1α (Lmx1a)—is sufficient to directly produce functional dopaminergic neurons from mouse and human fibroblasts from healthy donors and, of greater interest, from Parkinson's disease patients (23). Fibroblast-derived dopaminergic neurons produce dopamine and exhibit appropriate neurophysiological features (23).
In a parallel and independent experiment, another group has obtained similar results showing that the overexpression of the transcription factors Ascl1, Brn2, and Myt1l (BAM combination) with Lmx1a and forkhead box A2 (FoxA2), two proteins involved in dopamine neuron generation, can differentiate human fibroblasts into dopaminergic neurons (132). These dopaminergic neurons might be useful for in vitro disease modeling as well as for cell replacement therapies for the treatment of Parkinson's disease.
In the last three decades, dopaminergic cell transplantation has emerged as a potentially reparative therapy for Parkinson's disease (PD), given the relatively circumscribed nature of the neurodegenerative process involving predominately the nigrostriatal areas. Sources for such cells are varied and include the developing ventral mesencephalon, several autologous somatic cell types, as well as ESCs and iPSCs (37, 187). Recently successful engraftment of human ESC-derived dopamine neurons was demonstrated in animal models (using three host species: mice, rats, and monkeys) of PD by Kriks et al. (91). The authors described a novel floor-plate-based strategy for the derivation of human dopaminergic neurons that efficiently engrafted in vivo, suggesting that past failures were due to incomplete specification rather than to a specific vulnerability of the ESCs derived cells. However, while pluripotent derived neurons integrated very well into animal models of PD after grafting, in some experiments with iPSCs, they led to the formation of neural overgrowth because of the pluripotentiality of the starting cells (29, 188). The use of direct induction of somatic cells into dopaminergic neurons (iDAs) can circumvent this obstacle. In fact, the in vivo differentiation potential of iDA cells was assessed by orthotopic transplantations into neonatal mouse brains (23). Two and 6 weeks after transplantations, donor cells were found integrated in the host tissue showing an extremely elaborated morphology and an appropriate dopaminergic phenotype. Indeed, it has been recently described that ectopic expression of defined transcription factors (11 factors selected based on their known functions in the development and survival of midbrain DA neurons) in mouse fibroblasts is sufficient to induce neurons with midbrain dopaminergic features. In addition, transplantation of these iDA cells ameliorated symptoms in a mouse model of PD (37). Thus, iDA neurons generated from abundant somatic fibroblasts by direct lineage reprogramming hold promise for cell-based therapies of PD. Further studies in PD models are needed to evaluate the therapeutic effect of these cells.
The direct induction of other specific neuronal subtypes like motor neuron cells (iMNs) has recently been demonstrated (156). The authors investigated different combinations of the BAM factors with known motor neuron specification factors, employing embryonic murine fibroblasts carrying a fluorescent reporter under the motor neuron-specific promoter Hb9 to track the conversion to desired phenotype. They determined an optimal minimal combination of seven factors that generated a 5–10% conversion of mouse embryonic fibroblasts into functional active iMNs. These cells had an established transcriptional appropriate program, exhibited the electrophysiological characteristics of motor neurons, and formed functional synapses with muscle in vitro. Indeed, they displayed even in vivo engraftment capacity in chicken fetal spinal cord similar to ES-derived motor neurons. In addition, these iMNs presented both cell-autonomous and nonautonomous sensitivities to degenerative stimuli, strongly suggesting that they can be used as in vitro model of motor neuron disorders, like amyotrophic lateral sclerosis. Finally, the authors also successfully added NeuroD1 to their seven-factor reprogramming combination to reprogram human embryonic fibroblasts into functional, cholinergic hiMNs (156). Although requiring further investigation, other relevant neuroectodermal populations like astrocytes or oligodendrocytes can be directly obtained from somatic cells for basic pathogenetic studies (e.g., of nonautonomous events) or as cell sources for cell therapy.
Comparisons are lacking in the literature between neurons derived from direct conversion and those derived from pluripotent cells. We know that the process of neuron differentiation from pluripotent stem cells recapitulates all the steps of the developing embryonic nervous system and gives rise to all subtypes of neurons: spinal motor neurons, midbrain dopaminergic neurons, and cortical neurons [for review, see (130)]. However, the ability to derive specific neural subtypes from induced neurons remains elusive, although several groups have generated dopaminergic induced neurons (23, 85, 132) and motor neuron induced neurons (156). Thus, it is crucial to assess the bona fide phenotype of transdifferentiated neurons, characterizing them both from expression profiles and functionality. In fact, these aspects of induced neurons might have potential clinical relevance when a specific target population is needed for cell transplantation purposes.
In spite of these considerations, induced neurons might have potential clinical relevance. At present, there are no effective treatments for diseases and injuries of the nervous system. The replacement of damaged cells and the restoration of function are considered the ideal therapeutic strategy that can be addressed with reprogrammed neurons. Indeed, the use of differentiated neurons avoids teratoma formation, which can occur with embryonic stem cell transplantation. Moreover, terminally differentiated cells can potentially be used as an in vitro model to discover and test novel drugs and to study disease mechanisms. Furthermore, it seems that the direct conversion of fibroblasts into neurons can be more time effective—occurring, for example, as early as 8 days (128)—in vitro than the generation of iPSCs for basic studies and cell therapy purposes.
Transdifferentiation in Endodermal Derivatives: Hepatocytes
Under physiological conditions, liver hepatocytes originate from endogenous hepatic progenitor cells(199). However, previous experiments have demonstrated that the hepatocytes can also be obtained from nonhepatic cells after exposure to particular signals or fusion with hepatocytes (141, 179, 181). These data suggest that specific hepatocytic master genes can be activated in these situations. In fact, the expression of three specific combinations of only two transcription factors, comprising hepatocyte nuclear factor 4α (Hnf4α) plus Foxa1, Foxa2, or Foxa3, related to hepatocyte differentiation during liver development, is sufficient to directly differentiate mouse embryonic and adult fibroblasts into bona fide hepatocytes in vitro (145).
In a parallel experiment, the expression of another set of genes—Gata4, Hnf1α, and Foxa3—and the contemporaneous inactivation of p19 (alternative reading frame; Arf) produce the direct induction of functional hepatocyte-like (iHep) cells from mouse tail-tip fibroblasts (74). In both cases, the authors planned to evaluate the reprogramming effect of a larger pool of genes for 12 and 14 transcription factors, respectively, progressively eliminating one factor at a time until the key combination was found. This type of progressive approach has been used by Yamanaka with iPSCs (starting with 24 factors) (167) and also in other direct transdifferentiation approaches (176). Of note, in both cases, the fibroblast-derived hepatocytes can reconstitute hepatic tissues in vivo in the livers of fumarylacetoacetate hydrolase (Fah)-deficient (Fah-/-) recipient mice after transplantation, even if they could only partially compensate liver function (74, 145).
These studies offer new insights into the molecular mechanisms of hepatocyte differentiation and open the path to novel potential therapeutic approaches for liver diseases (145).
Transdifferentiation In Vivo
Because promising results have been obtained in vitro, one goal of medicine may be to understand reprogramming mechanisms and take advantage of them for tissue repair and regeneration. To achieve this goal, an important step is to carry out transdifferentiation in vivo. In vivo cells would reside in their native environment, which might promote their survival and/or maturation and influence the reprogramming process.
A few examples are available in the literature of in vivo conversion. The first is a study by Xie et al., who reprogrammed B-cells into macrophages in vitro and then determined whether this result could also be observed in vivo (193). Sublethally irradiated Recombination Activating Gene 2 interleukin 2 γC receptor knockout (RAG2–/–γc–/–) mice were transplanted intravenously with B-cell precursors from a bone marrow pool of CD19 ancestry mice infected with a C/EBPα-carrying virus. In the three animals treated, 6 days after transplantation, 51% of the C/EBPα-expressing cells were CD19– Mac-1+ in the bone marrow and 32% in the spleen, markers specific to macrophages. Most of the reprogrammed cells also exhibited relatively low side scatter values, suggesting that the in vivo reprogrammed cells resembled monocytes (193).
The second in vivo study was by Zhou and colleagues (206). Based on previous attempts to convert adult liver cells into β-cells in vivo by expressing pancreatic transcription factors (45, 83, 117, 179), starting from 20 transcription factors and after progressive elimination, they identified a specific minimal combination of three transcription factors, Ngn3, pancreatic and duodenal homeobox 1 (Pdx1), and pancreatic β-cell-specific transcriptional activator (MafA), that reprograms differentiated pancreatic exocrine cells (in adult mice) into cells that closely resemble β-cells. The three selected adenoviruses were injected as a mixture into the splenic lobe of the dorsal pancreas of 2-month-old adult Rag1–/–mice. Twenty percent of infected cells were converted to insulin+ cells, in which, by day 10, the level of insulin was comparable to that of endogenous β-cells (Fig. 3). Thus, the conversion process was rapid, requiring only few days, without cell division, and stable for months. Consistent with previous reports (180), most infected cells (95%) were found to be mature amylase+ exocrine cells. Most of the insulin+ cells coexpressed genes required for β-cell endocrine function, such as glucose transporter 2, glucokinase, prohormone convertase, and the key β-cell transcription factors NeuroD, gene NK2 homeobox 2 (Nkx2.2), and Nkx6.1. However, the induced β-cells lacked organization into islet structures and remained as single cells or small clusters, probably impairing their function (206). Interestingly, the induced β-cells secreted insulin and rescued streptozotocin-induced hyperglycemia in adult animals, demonstrating that the reprogrammed cells possessed functional properties and that this strategy can have therapeutic meaning.
Direct in vivo reprogramming. Schematic experimental design of in vivo transdifferentiation: injection of the transcription factors neurogenin 3 (Ngn3), pancreatic and duodenal homeobox 1 (Pdx1), and pancreatic β-cell-specific transcriptional activator (MafA) intravenously induces pancreatic exocrine cells to convert into β-cells (206).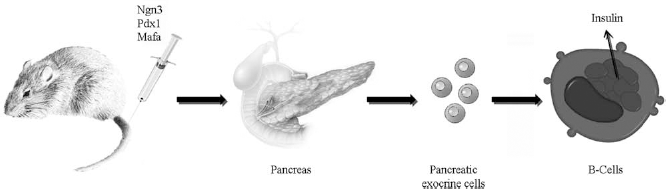
β-Cell destruction by immune system or their dysfunction causes diabetes mellitus type 1. A promising strategy of regenerative medicine is to obtain β-cells directly from patients' pancreatic exocrine cells. In general, although limited by the obvious complications associated with in vivo gene delivery, in vivo reprogramming approach can theoretically allow an endogenous and autologous tissue repair without any invasive transplantation, even if this goal seems to be a long way off.
Conclusion
The lack of effective therapies for most neurodegenerative diseases has raised significant interest in cellmediated approaches as potential strategies. Cells can exert therapeutic action through several mechanisms, including cell substitution and neuroprotection. Some of the common degenerative diseases are potentially amenable to cell therapy to replace damaged cells, tissues, or organs. Indeed, a primary goal of regenerative medicine is to produce new cells to repair or replace diseased and damaged tissues. Understanding the stability and the possibility of altering the differentiated state of various cell types is interesting both from a basic biological perspective and for its potential to generate cells for therapy. Cell types obtained from somatic cells can be pluripotent stem cells or other differentiated somatic cells (via transdifferentiation). These differentiated cells, such as neurons or cardiomyocytes, obtained from a patient's own somatic cells (from pluripotent stem cells or through transdifferentiation processes), would be useful in the regenerative medicine field because they are autologous and could thus theoretically preclude immunological rejection in cell transplantation approaches.
The direct reprogramming of adult somatic cells into the desired target cells holds many possible translational applications. With this technology, it is now achievable to generate disease-specific cells that harbor the genome of the donor and can mimic human disease. This approach might allow obtaining cell lines from many different patients to study the nature and complexity of disease, comparing how the same disease varies among subjects. Often, translation of drug discovery from levels of cellular and animal models to human therapeutics failed, because of the huge differences between the two systems. High-throughput drug screening using human transdifferentiated cells based on disease models could be useful in filling the gap between animal models and clinical trials. Drug screening platforms can be developed to test compounds that are able to revert disease-related phenotype to that of the control. It would be possible to perform a personalized toxicological risk assessment as well as to evaluate the response to specific drugs in cells with identical genetic blueprint of the individuals from which they are generated.
Moreover, transdifferentiation could be an innovative way of achieving the goal of regenerative medicine, considering that existing adult cells may be converted directly into other medically important cell types to repair diseased or damaged tissues. Indeed, the final product of transdifferentiation would be readily useful for direct clinical application because of their full commitment, whereas ESCs and iPSCs must be converted into mature or progenitor cells through various steps of differentiation. One major weakness of pluripotent stem cells is that they can generate tumorigenic cells in vivo, while transdifferentiated somatic cells would overcome this issue. The generation of iPSCs is still a lengthy and inefficient process, as is the generation of specific differentiated phenotypes from iPSCs, which is a multiphasic procedure requiring several chronologically regulated molecular signals and often producing mixed populations.
In theory, eliminating the pluripotent stem cell stage probably avoids the risk of tumorigenesis when the cells are transplanted. On the other hand, the absolute absence of replication, as in the case of direct differentiation of somatic cells into neurons, might generate a problem in terms of the number of cells available for transplantation. It is likely that for clinical application, the amount of donor cells requested for transplantation will be huge and not easily satisfied by the direct generation procedure. In fact, the efficiency of direct reprogramming is generally not very high—less than ~5% on average among the studies describing fibroblast-to-neuron conversion. Yoo et al. reported that the efficiency was even further diminished when starting from adult human cells, while neuronal maturation was delayed as well (195). Indeed, the age of the donor and the number of cell passages for starting cells like fibroblasts can influence the success of the transdifferentiation.
Another consideration is how long the cells originating from transdifferentiation can be maintained in culture and if they are too fragile or susceptible to stress to be cultured in the long term and then used for cell transplantation. Thus, it is necessary that reprogramming techniques be further developed to allow for the robust and efficient generation of target desired cells for practical application. In fact, although the number of cells obtained up to now with direct differentiation can be sufficient for in vitro basic studies, it might not represent a sufficient amount of cells for in vivo cell transplantation purposes. At present, it is not clear if the potential for beneficial results from the transplantations would arise more from partially differentiated or fully differentiated cells and have added relevant text. In the field of neurodegenerative diseases, the use of neural stem cells as an alternative to already differentiated neurons can apply for different purposes. For example, it might be possible to replace lost neurons or glial cells by transplantation of stem cell-derived cells that have been predifferentiated in vitro to various stages of maturation, for example, into neuroblasts (i.e., immature neurons). On the other hand, transplanted neural stem cells can be more effective in producing functional improvement by releasing therapeutic molecules that are neuroprotective.
Direct converted cell transplantation has yet to be experimentally tested in most organ systems, even if some experimental demonstration has been provided. For example, transplanted fibroblast-derived hepatocytes repopulated the livers of fumarylacetoacetate-hydrolase-deficient (Fah–/–) mice and rescued half of recipients from death by restoring liver function (74). Regarding the cardiovascular system, it has been described that fibroblasts induced into cardiac cells further completed their cardiac differentiation in vivo in immunosuppressed NOD-SCID mouse hearts, with interesting implication for the development of repair strategy for cardiac diseases (77). A remarkable example of autologous cell replacement therapies was provided by using multipotent blood progenitors obtained from fibroblasts through the forced overexpression of OCT4 (164). Reprogrammed hematopoietic cells engrafted and repopulated bone marrow in all transplanted recipient animals at 20% level (164). This represents a proof of principle that converting fibroblasts or other patient specific cells into hematopoietic cells may be a source for autologous transplantation. Indeed, it has been recently demonstrated the possibility of using fibroblast-derived neuron cells as a source of transplantable cells with therapeutic regenerative potential in PD model (37). However, only one functional test was used to evaluate the disease improvement, too limited with respect to the complete investigation of PD phenotype that is required in preclinical trials. Indeed, the number of effective induced cells required is greater than that required with neuronal precursors derived from CNS. Overall, it seems that directly differentiated neurons, generated with the available methods, are only partially therapeutic effective with respect to those generated from pluripotent stem cells or derived from tissues-specific precursors. A direct comparison is still lacking. Further studies are needed for the effective development of this strategy. Regardless, transdifferentiated cells must be widely characterized before use in the clinic. The first step to determine the identity of the cell type produced and to ensure that they do not express markers of the initial cell type is evaluation of marker genes by immunocytochemistry and gene expression analysis. Analysis of cellular morphology can provide further information about the successful generation of a particular cell type. Furthermore, physiological behaviors can be tested to establish whether the cells display functional properties that are similar to those of the same type in vivo. It can also be important to evaluate the transcription profile, DNA methylation, and histone modifications of the differentiated cells relative to their in vivo correlated tissue (32). Indeed, it is necessary to determine the fidelity of reprogramming and define a clinically acceptable level of genomic integrity for transdifferentiated cells. Multiple kinds of genomic changes have been observed in human iPSCs (109), and chromosomal aneuploidy and translocations, megabase-scale duplications/deletions, and point mutations have all been described (52, 76, 100, 109, 110). Moreover, the reprogramming process may cause epigenomic reorganization, DNA methylation alteration, and histone modifications. As in the case of iPSCs, these events may occur during the reprogramming process for the cell transdifferentiation. Up to now, no specific studies have analyzed this aspect in transdifferentiated cells.
Most current methods of producing transdifferentiated cells involve viral gene delivery that could cause abnormalities in the induced cells. Random integration of the reprogramming genes may result in insertional mutagenesis that causes malignant transformation of a clonal cell population. Research should focus on producing induced cells without using viral vectors as with iPSCs, such as applying nonintegrating viral (161), nonviral episomal (123, 196), and excisional (82, 191) techniques for reprogramming, or treatment with small molecules (75, 205), cell signaling peptides [e.g., wingless-type MMTV integration site family (Wnt)] (107), or microRNAs (163). Moreover, coculture with animal cells or animal media could lead to immune reactions against animal proteins in the cells and infection caused by animal microbes. Using animal substance-free culture media, feeder cells, or a feeder-free matrix in derivation, passaging, expansion, and cryopreservation procedures can avoid these problems and produce clinical-grade cells with Good Manufacturing Practice (GMP) quality.
Distinct cell types are defined by their unique combination of epigenetic marks that are accumulated progressively during numerous developmental steps. Thus, other crucial points to deal with are the epigenetic features of the obtained cells. Changes in transcriptional activity could lead to genome-wide modifications such as DNA methylation, histone adjustments, and alteration of chromatin remodeling complexes (79, 207). In light of this information, a potential benefit of transdifferentiation approaches is that only some of the numerous epigenetic markers established during development need to be rearranged, so this process requires fewer proliferation steps and may reduce mutation events. Meanwhile, iPSC generation removes several epigenetic markers that must be reestablished during the in vitro differentiation process. These changes could result in unstable phenotypes when cells are transplanted in vivo (207).
Another promising advantage of the transdifferentiation process is the possibility of inducing the desired cell types directly in vivo for in situ regeneration and repair (55). Indeed, the in vivo model can provide an appropriate three-dimensional setting, as it has been shown that differentiated human cells mature more properly if transplanted into an adult mouse than when cultured only in vitro (92); therefore, it may be advantageous to induce transdifferentiation directly in the native tissue. Transdifferentiation thus represents an attractive approach that may become an important feature of regenerative medicine. However, other studies regarding safe delivery of defined factors and development of current protocols in preclinical model systems are required to improve this technology for potential application in preclinical and clinical studies.
Footnotes
Acknowledgments
S.C. was supported by Grant MIUR, RBFR08RV86, “Development of a stem cell approach for motor neuron diseases,” and G.P.C. by the ARISLA grant “iPS derived neural stem cells for amyotrophic lateral sclerosis.” We wish to thank the Associazione Amici del Centro Dino Ferrari for their support. We wish to thank Dr. Serena Ghezzi for her help with the graphics. The authors declare no conflict of interest.
