Abstract
Somatic cells can be directly reprogrammed to alternative differentiated fates without first becoming stem/progenitor cells. Nevertheless, the initial need for viral-mediated gene delivery renders this strategy unsafe in humans. Here, we provide evidence that exposure of human skin fibroblasts to a Radio Electric Asymmetric Conveyer (REAC), an innovative device delivering radio electric conveyed fields at a radiofrequency of 2.4 GHz, afforded remarkable commitment toward cardiac, neuronal, and skeletal muscle lineages. REAC induced the transcription of tissue-restricted genes, including Mef2c, Tbx5, GATA4, Nkx2.5, and prodynorphin for cardiac reprogramming, as well as myoD, and neurogenin 1 for skeletal myogenesis and neurogenesis, respectively. Conversely, REAC treatment elicited a biphasic effect on a number of stemness-related genes, leading to early transcriptional increase of Oct4, Sox2, cMyc, Nanog, and Klf4 within 6–20 h, followed by a downregulation at later times. The REAC action bypassed a persistent reprogramming toward an induced pluripotent stem cell-like state and involved the transcriptional induction of the NADPH oxidase subunit Nox4. Our results show for the first time the feasibility of using a physical stimulus to afford the expression of pluripotentiality in human adult somatic cells up to the attainment of three major target lineages for regenerative medicine.
Introduction
A major obstacle in future cell-based therapies is the immunological rejection of mismatched cellular grafts. One solution to this problem would be to generate a source of autologous pluripotent stem cells that can be committed toward specific phenotypes by previous epigenetic reprogramming of somatic cells to a pluripotent embryonic stem (ES) cell-like state. In this regard, cellular reprogramming of somatic cells into induced pluripotent stem (iPS) cells and subsequent differentiation to mature cells have opened up new possibilities for tissue regeneration and repair (4,24,25,32).
More recently, different authors have shown that somatic cells can also be directly converted into different mature cell types by expression of genes orchestrating cell commitment toward a specific phenotype (6,8,26). In particular, a group of researchers (8) identified a cocktail of specific factors that could reprogram mouse fibroblasts into cardiac muscle, without having to revert to a stem cell state throughout the process. This strategy could be an advantage over using iPS or ES cells to regenerate complex mature cells. In fact, it is now evident that stray cells that have not fully differentiated might have the ability to turn into an unwanted cell type, like a tumor or a cell that just does not fulfill the desired requirement(s) for a targeted tissue repair.
We have previously shown that exposure of mouse ES cells to radiofrequency (RF) energy through a Radio Electric Asymmetric Conveyer (REAC), an innovative device delivering radio electric conveyed fields (RECF) of 2.4 GHz with its conveyer electrodes immersed into the culture medium, remarkably enhanced the commitment toward different lineages, including cardiogenic, neurogenic, and skeletal myogenic, without the aid of chemical compounds or viral vector-mediated gene delivery (11).
In the current study, we investigated whether REAC-RECF may directly reprogram human skin-derived fibroblasts (hSFs) toward complex lineages. Gene and protein expression analysis and immunohistochemical characterization revealed that this new strategy was able to afford a rapid, high-throughput, and stable commitment of the exposed cells toward cardiogenesis, skeletal myogenesis, and neurogenesis, leading to the appearance of specific marker proteins for terminal differentiation. We also provided evidence that the REAC effect encompassed the transcriptional upregulation of the nicotinamide adenine dinucleotide phosphate (NADPH) oxidase subunit Nox4, a major signaling component in the intracellular generation of reactive oxygen species (ROS).
Materials and Methods
Description of Radio Electric Asymmetric Conveyer (REAC)
The REAC technology was developed to convey in an asymmetric way, the radio electric currents generated by the interaction between the weak electromagnetic field produced by the REAC device (19,20), with a radiated power of about 2 mW, and the electromagnetic field generated by the human body, with a radiated power of about 54 mW (28). The REAC device was placed into a CO2 incubator and was set at a frequency of 2.4 GHz. Its conveyer electrodes were immersed into the culture medium of hSFs, as previously described (11). The electromagnetic quantities have been measured with the spectrum analyzer Tektronix model 2754p (TekNet Electronics, Inc., Alpharetta, GA, USA), orienting the receiving antenna for maximum signal. With cell cultures at a distance of 35 cm from the emitter, we measured a radiated power of approximately 400 μW/m2. Electric field E = 0.4 V/m, magnetic field = 1 mA/m, specific absorption rate (SAR) = 0.128 μW/g; given s = 1 A/Vm, and r = 1,000 kg/m3, the density of radio electric current flowing in the culture medium during a single radiofrequency burst by the REAC is J = 30 μA/cm2. The instrument that we used is registered under the trademark (B.E.N.E.) Bio-Enhancer Neuro-Enhancer®, produced by ASMED srl, Firenze, Italy.
Culture of Human Skin Fibroblasts
Human foreskin fibroblasts were purchased from LGC Standards (Sesto San Giovanni, Milan, Italy) and cultured in minimum essential medium Eagle-alpha modification (Alpha MEM) containing 10% fetal bovine serum (FBS), 2% l-glutamine, and 2% penicillin/streptomycin (all Gibco/Invitrogen, Monza, Italy). For the experiments, cells were seeded at a concentration of 5×103 cells/well in a multi-well tissue culture plate (BD Biosciences, San Jose, CA, USA) and treated in the absence or presence of REAC. After 72 h, REAC-treated cells were cultured for an additional 4 or 7 days without REAC stimulation and then processed for transcriptional analysis and immunostaining. The culture medium was not changed during the 10 days of culturing.
Assessment of Cell Proliferation and Apoptosis
Cell viability was determined by the trypan blue dye exclusion test (Gibco). Both attached and floating cells were harvested and counted by using the Countess automated cell counter (Invitrogen). To assess apoptosis, caspase 3 activity was detected using a commercial kit (Fluorimetric Caspase 3 Assay Kit, Sigma-Aldrich Corp., St. Louis, MO, USA) according to the manufacturer's instructions. Briefly, the cell pellet was dissolved in 35 μl of lysis buffer, and cell lysates were mixed with the peptide substrate acetyl-Asp-Glu-Val-Asp-7-amido-4-methylcoumarin (Ac-DEVD-AMC). Release of the fluorescent 7-amino-4-methylcoumarin (AMC) moiety was assessed as a measure of substrate hydrolysis by caspase 3. Lactate dehydrogenase (LDH) release by dead cells was detected with an LDH cytotoxicity assay kit II (MBL International Corporation, Woburn, MA, USA).
Flow Cytometry Analysis
Cells were harvested by treatment with 0.08% trypsin, EDTA (Invitrogen). After a fixation/permeabilization step, cells were incubated with a primary antibody directed against β-3-tubulin (BD Biosciences), myogenic differentiation 1 (myoD; Santa Cruz, Dallas, TX, USA), or α-sarcomeric actinin (Sigma) (1 mg/106 cells) for 1 h at 4°C and with 1 mg of fluorescein isothiocyanate-conjugated secondary antibody (BD Biosciences) for 1 h at 4°C in the dark. After washing, cells were analyzed on a flow cytometer (FACSAria, BD Biosciences) by collecting 10,000 events, and the data were analyzed using the FACS Diva software (BD Biosciences).
Gene Expression
Total RNA was isolated using Trizol reagent according to the manufacturer's instructions (Invitrogen). Total RNA was dissolved in RNAase-free water, and for RT-PCR, cDNA was synthesized in a 50-μl reaction volume with 1 μg of total RNA and Moloney Murine Leukemia Virus reverse transcriptase (MMLV RT) according to the manufacturer's instructions (Invitrogen). Quantitative real-time PCR was performed using an iCycler Thermal Cycler (Bio-Rad, Hercules, CA, USA). Two microliters of cDNA were amplified in 50-μl reactions using Platinum Supermix UDG (Invitrogen), 200 nM of each primer, 10 nM fluorescein (BioRad), and Sybr Green. After an initial denaturation step at 94°C for 10 min, temperature cycling was initiated. Each cycle consisted of 94°C for 15 s, 55-59°C for 30 s, and 60°C for 30 s. The fluorescence was read at the end of this step. All primers used in this study were from Invitrogen: the primers for human NK2 homeobox 5 (Nkx-2.5), guanine-adenine-thymine-adenine-binding protein 4 (GATA4), glyceraldehyde 3-phosphate dehydrogenase (GAPDH), neurogenin 1, and MyoD were previously described (21,29). The other specific primers used were for human Krüppel-like factor 4 (Klf4) forward 5′-CCC TCCGACGGCTCCCTTCA-3′ and reverse 5′-AACTTGC CCATCAGCCCGCC-3′; for human sex-determining region Y box 2 (Sox2) forward 5′-CACATGAACGGCTGGA GCA-3′ and reverse 5′-TGCTGCGAGTAGGACATGC TG-3′; for human octamer binding transcription factor 4 (Oct4) forward 5′-CTCACCCTGGGGGTTCTAT-3′ and reverse 5′-CTCCAGGTTGCCTCTCACTG-3′; for human Nanog forward 5′-CATGAGTGTGGATCCAGCT-3′ and reverse 5′-CCTGAATAAGCAGATCCAT-3′; for human v-myc myelocytomatosis viral oncogene homolog (cmyc) forward 5′-GGCCGTTTTAGGGTTTGTTGG-3′ and reverse 5′-ACAAGTTTCCAGCCACCTCC-3′; for human Nox1 forward 5′-AATCCTTGGGTTTTGCAGCC-3′ and reverse 5′-GTCGTGTTTCGGGACTGGAT-3′; for human Nox4 forward 5′- AATGCACCAACAAATGGGGC-3′ and reverse 5′-AGTGTGTGGAGTTGACCCAG-3′; for human T-box protein 5 (Tbx5) forward 5′-CAGAGTCGGCACAGCG GCAA-3′ and reverse 5′-GTGGGGAGCCATGGTTGG CC-3′; for human myocyte enhancer factor-2c (Mef2c) forward 5′-GCCCTGAGTCTGAGGACAAG-3′ and reverse 5′-AGTGAGCTGACAGGGTTGCT-3′; and for human prodynorphin forward 5′-TGGCCAAGCTCTCTGGGTCA-3′ and reverse 5′-TCATGGCCCATGCTATCCCC-3′. To evaluate the quality of product of real-time PCR assays, melting curve analysis was performed after each assay. Relative expression was determined using the “delta-CT method” with glyceraldehyde 3-phosphate dehydrogenase (GAPDH) as reference gene (29).
Immunostaining
Cells were cultured for 3 days with or without REAC exposure and then for an additional 4 days without REAC treatment. In another set of experiments, cells were preincubated for 2 h prior to REAC treatment with the free radical scavenger N-(2-mercapto-propionyl)-glycine (NMPG) (100 mM; Sigma). After 7 days, cells were treated with trypsin, and the resulting suspension was cultured at low density to permit visualization of individual cells. The cultures were fixed with 4% paraformaldehyde (Invitrogen). Cells were exposed for 1 h at 37°C to mouse monoclonal antibodies against α-sarcomeric actinin (Sigma), β-3-tubulin, MyoD or myogenin, or with rabbit polyclonal antibodies against myosin heavy chain and stained at 37°C for 1 h with fluorescein-conjugated goat IgG (all from Santa Cruz).
All microscopy was performed with a Leica confocal microscope (LEICA TCSSP5, Milan, Italy). DNA was visualized with 1 mg/ml 4′,6-diamidino-2-phenylindole (DAPI, Sigma).
Data Analysis
The statistical analysis of the data was performed by using the Statistical Package for Social Science (SPSS), version 13 (IBM, Armonk, NY, USA). For this study, we used nonparametric statistical tests: Kruskal–Wallis and Wilcoxon signed rank test. The first test was to evaluate the distribution and homogeneity of variance of each group at different times of observation, while the second test was used to evaluate, in the same group, the differences (delta CT) between the data collected over a period of observation and the reference value at baseline. Cell viability necrosis and apoptosis were analyzed by two-tailed, unpaired Student's t test, or one-way analysis of variance with subsequent Bonferroni test. All results with p < 0.05 have been considered statistically significant.
Results and Discussion
Effects of REAC Exposure on Cellular Proliferation and Apoptosis
REAC exposure had no toxic effect on hSFs. In fact, there was no significant difference in cell growth between exposed and unexposed cells (Fig. 1A). Moreover, REAC did not significantly affect the amount of apoptotic (Fig. 1B) or necrotic cells (Fig. 1C), as compared with untreated hSFs. Trypan blue exclusion also revealed no significant difference between exposed and unexposed cells (not shown).
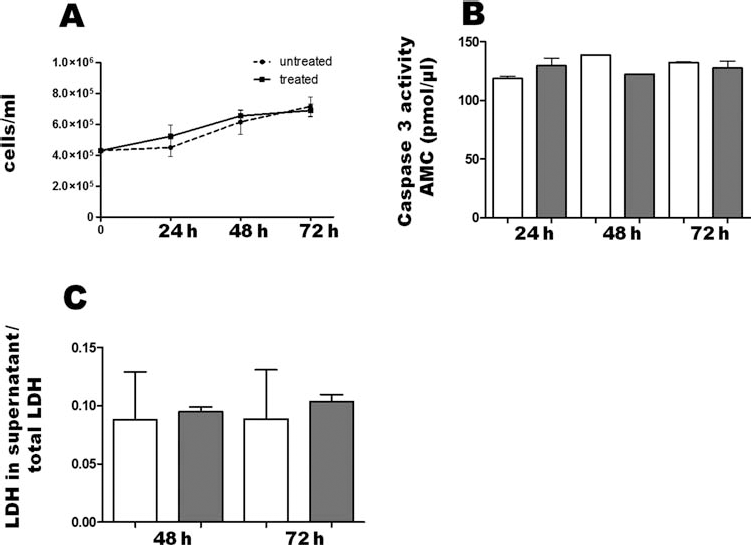
Effect of REAC on cell viability, apoptosis, and necrosis. After plating (2 × 105/well), cells were left untreated (white bars) or treated (gray bars) with a Radio Electric Asymmetric Conveyer (REAC) for the indicated times and counted to estimate cell growth (A) or processed to assess the amount of apoptosis (B) or necrosis (C) (mean ± SE; n = 6). In (B), release of the 7-amido-4-methylcoumarin (AMC) moiety (pmol/ml) after cleavage of the fluorogenic acetyl-Asp-Glu-Val-Asp-7-amido-4-methylcoumarin (Ac-DEVD-AMC) caspase substrate by cell lysates was calculated from a standard curve determined with defined AMC solutions. (C) Cell supernatants and pellets were collected, and total proteins extracted from both fractions. Lactate dehydrogenase (LDH) activity in each supernatant was normalized to total LDH activity of their own pellet + supernatant.
REAC-Delivered RECF Induces the Expression of Cardiogenic Genes
The zinc-finger transcription factor GATA4 Tbx5, a member of the T-box transcription factor family, and Mef2c, belonging to the myocyte enhancer factor-2 (MEF2) family of MADS (MCM1, agamous, deficiens, serum response factor) box, have an essential role in the establishment of the cardiac lineage. In this regard, it has been previously described that overexpression of these factors allows the conversion of mouse skin-derived fibroblasts into cardiomyocytes, expressing sarcomeric-like structures (8). In a previous study in mouse ES cells, we demonstrated the ability of REAC stimulation to increase the expression of the cardiogenic genes GATA4, Nkx2.5, and prodynorphin, leading to a high yield of spontaneously beating myocardial cells (11). We also provided evidence for the ability of REAC treatment in promoting the repair of surgical wounds (2) and injury (5).
In the current study, we sought evidence on whether REAC-mediated exposure may be able to orchestrate a myocardial lineage transition in hSFs. For this purpose, a REAC apparatus (11), working into a CO2 incubator, was set at a transmission frequency of 2.4 GHz and its conveyer electrodes were immersed into the culture medium. Radiated power was about 2 mW. Electromagnetic field E = 0.4 V/m, with culture medium of hSFs 35 cm away. Duration of single radiofrequency burst was 200 ms with an off interval of 2.5 s.
To determine whether there was activation of cardiogenic signaling, we performed quantitative real-time PCR analyses of Mef2c, Tbx5, and GATA4 in hSFs treated in the absence or presence of REAC for 24, 48, and 72 h. Interestingly, we observed a 40-fold increase in Mef2c expression, peaking after 72 h of treatment (Fig. 2A). REAC exposure significantly increased Tbx5 gene expression after 48 h, resulting in a 50-fold enhancement of Tbx5 mRNA levels at 72 h (Fig. 2B). The expression of GATA4 was increased by 15-fold after 48 h of REAC treatment (Fig. 2C). Moreover, cells treated with REAC up to 72 h and then cultured in the absence of REAC stimulation for an additional period of 4 or 7 days, exhibited a further increase in the expression of all the cardiogenic genes (Fig. 2). These findings indicate that upon REAC exposure a cardiogenic program was induced. Further support to this view arises from the observation that REAC exposure also remarkably increased the transcription of prodynorphin and Nkx2.5 (Fig. 3A, B), two genes deeply involved in stem cell cardiogenesis (10,12,30,31). In particular, Nkx2.5 mRNA levels were early increased after 24 h of treatment, reaching an 80-fold increase after 72 h, as compared with unexposed cells. Prodynorphin mRNA levels shown a significant increase during the first 72 h of REAC treatment, followed by a further increase after 7 days in culture in the absence of REAC stimulation (Fig. 3).

REAC exposure induces the expression of genes involved in cardiac reprogramming. Cells were exposed for 24, 48, or 72 h in the absence or presence of REAC or were treated with REAC for 72 h and then left unexposed for additional 4 or 7 days (day 7 or day 10 from time zero, respectively). The amounts of myocyte enhancer factor 2C (Mef2c) (A), T-box protein 5 (Tbx5) (B), or guanine-adenine-thymine-adenine-binding protein 4 (GATA-4) (C) mRNA from REAC-treated or untreated cells were normalized to glyceraldehyde 3-phopshate dehydrogenase (GAPDH), and the mRNA expression of REAC-exposed cells was plotted at each time point as fold change relative to the expression in control untreated cells, defined as 1 (mean ± SE; n = 6). ∗Significantly different from untreated cells.

Effect of REAC stimulation on the expression of genes orchestrating cell commitment toward cardiogenic, skeletal myogenic, and neurogenic lineages. Cells were exposed for 24 or 48 or 72 h in the absence or presence of REAC or were treated with REAC for 72 h and then left unexposed for additional 4 or 7 days. The amounts of prodynorphin (A), NK2 homeobox 5 (Nkx2.5) (B), neurogenin 1 (C), and myogenic differentiation 1 (MyoD) (D) mRNA from REAC-treated or untreated cells were normalized to GAPDH, and the mRNA expression of REAC-exposed cells was plotted at each time point as fold change relative to the expression in control untreated cells, defined as 1 (mean ± SE; n = 6). ∗Significantly different from untreated cells.
REAC Induces the Expression Genes Controlling Multiple Lineage Commitment
We have previously provided evidence that REAC exposure resulted in the induction of targeted genes controlling neurogenic and myogenic commitment, followed by the appearance of the corresponding phenotypes, in mouse ES cells (11). Here, we evaluated the effect of REAC treatment on the transcription of neurogenin 1 for neurogenesis, and myoD, for skeletal myogenesis. Compared to the gene expression of the cardiogenic transcription factor Nkx2.5, neurogenin 1 transcription was enhanced at a later time by REAC treatment, being significantly increased after 72 h of exposure (Fig. 3C). Worthy to note, after a 4-day additional culture in the absence of REAC, neurogenin 1 transcription continued to increase, the specific mRNA levels being 200-fold more expressed than in the control cells that did not receive any previous REAC treatment (Fig. 3C). Comparative time-course analyses also revealed that in REAC-treated cells the overexpression of Nkx2.5 declined earlier than that of neurogenin 1 (Fig. 3B, C). The mRNA levels of myoD were 40-fold increased after 72 h of treatment and when cultured up to 4 days without REAC exceeded by 60-fold the levels detected in unexposed controls (Fig. 3D). On the whole, these observations indicate a modularity in the timely expression of tissue-restricted orchestrators of complex differentiating patterning in REAC-reprogrammed hSFs. So far, a remarkable complexity has been encountered in the expression profiles of these transcription factors, with unexpected multidirectional actions, as shown by the ability of Nkx2.5 itself to drive noncardiogenic decisions, including the commitment to neuronal differentiation in both skeletal muscle and ES cells (18,27). Deciphering the molecular mechanism that interconnect with the transcriptional units intervening in REAC-mediated commitment of hSFs may provide a window into the cellular circuitry that specifies the attainment of multiple fates from these cells.
Immunocytochemical Analysis of Cardiogenic, Neurogenic, and Myogenic Differentiation
To further confirm cardiogenic, neurogenic, and myogenic differentiation of hSFs, cells cultured in the absence or presence of REAC were subjected to confocal microscopy analysis in the presence of selected antibodies targeted against α-sarcomeric actinin and α-myosin heavy chain for cardiogenesis, MyoD for skeletal myogenesis, and β-3-tubulin for neurogenesis. As shown in Figure 4, expression of each selected marker protein could be observed only in REAC exposed cells. Flow cytometry analysis performed in hSFs treated with REAC for 72 h and then left unexposed for additional 4 days revealed that 16.30 ± 1.8% of cells expressed β-3-tubulin, whereas the percentage of myoD- and α-sarcomeric actinin-positive elements was 21.60 ± 2.20% or 32.00 ± 2.00%, respectively (mean ± SE; n = 6).

REAC induces fibroblast commitment toward cardiac, neuronal, and skeletal muscle lineages. The expression of α-myosin heavy chain (MHC), α-sarcomeric actinin (α-actinin), MyoD, and β-3-tubulin (tubulin) was assessed by confocal microscopy in fibroblasts cultured for 72 h in the absence or presence of REAC and then left unexposed for additional 4 days. In separate experiments, cells were preincubated for 2 h prior to REAC treatment with the free radical scavenger N-(2-mercapto-propionyl)-glycine (NMPG) (100 μM). Scale bars: 40 μm. Nuclei are labeled with DAPI (blue). Representative of five separate experiments.
Fibroblasts Treated with REAC Are Directly Reprogrammed Without a Persistent iPSC State
Oct4, Sox2, and Nanog play a major role in maintaining stem cell identity. These transcription factors are master regulators, silencing genes that are waiting to create the next generation of cells, and upon their disabling stem cells rapidly begin to differentiate (9,13,14,16). Ablation of the Oct4 gene in mouse embryos prevented proliferation of inner cell mass (ICM) cells and promoted differentiation into trophectoderm (15). Once expressed, Nanog blocks differentiation (15). Thus, negative regulation of Nanog is required to sustain differentiation during ES cell commitment (3). Similarly, early phases of ES cell differentiation involved a downregulation of Sox2 expression (11). Viral vector-mediated transduction of hSFs with Sox2, Oct4, Klf4, and c-Myc can directly reprogram somatic cells into a pluripotent state (24), opening new routes for regenerative therapies. Nevertheless, persistent activation of these genes in engineered cells is associated with very low differentiation efficiencies (24), also raising cautionary issues concerning the fate of the large proportion of cells that remain undifferentiated after commitment to defined lineages. We have previously shown the effectiveness of REAC treatment in downregulating the expression of the genes controlling pluripotency in mouse ES cells after 24 h of exposure. Here, time-course analysis at shorter intervals of Oct4, Sox2, cMyc, Nanog, and Klf4 transcription, in exposed and unexposed hSFs, revealed that REAC elicited a biphasic response encompassing an early increase in the gene expression during the first 6–20 h, followed by a downregulation after 24 h (Fig. 5A–E). Such a decline persisted after 72 h of exposure, being still clearly evident even when cells were maintained in culture for an additional 4 or 7 days in the absence of REAC treatment (Fig. 5A–E). These findings indicate that REAC may have initially switched hSFs to pluripotency without freezing cells into an iPSC-like intermediate state, thus avoiding a decrease in differentiation efficiency from a long-term overexpression of stemness-related genes.

REAC treatment time-dependently modulates the expression of genes involved in fibroblast to induced pluripotent stem cell (iPSC) reprogramming. Cells were exposed in the absence or presence of REAC for different time periods up to 72 h or were treated for 72 h and then left unexposed for additional 4 or 7 days (day 7 or day 10 from time zero, respectively). The amounts of octamer-binding transcription factor 4 (Oct4) (A), sex-determining region Y box 2 (Sox2) (B), v-myc myelocytomatosis viral oncogene homolog (cMyc) (C), Nanog (D), and Krüppel-like factor 4 (Klf4) (E) mRNA from REAC-treated or untreated cells were normalized to GAPDH, and the mRNA expression of REAC-exposed cells was plotted at each time point as fold change relative to the expression in control untreated cells, defined as 1 (mean ± SE; n = 6).
REAC-Mediated Commitment Involves Selective Induction of Nox4 Subunit of NADPH Oxidase
Previous studies revealed a role of reactive oxygen species (ROS) in transducing mechanically or electrically driven cardiovascular differentiation in mouse ES cells (22,23). Cardiac differentiation was accompanied by an upregulation of the NADPH oxidase isoforms Nox2 and Nox4 (1). In the current study, REAC-exposed hSFs exhibited a time-dependent increase in the gene expression of Nox4, which reached a maximum at 7–10 days in cells that were left unexposed after the first 72 h of treatment (Fig. 6A). Different from Nox4, Nox1 transcription was downregulated during the first 72 h of REAC treatment, while increasing after an additional 7 days in culture without further treatment (Fig. 6B). These observations prompt one to verify whether ROS may be involved in the differentiating responses elicited by REAC in hSFs. Such a possibility is indeed inferred from the finding that all the differentiating responses elicited by REAC were abolished when cells where cultured in the presence of NMPG, a free oxygen radical scavenger (Fig. 4). Studies are in progress to dissect the molecular patterning and cell signaling network involved in the ROS action.
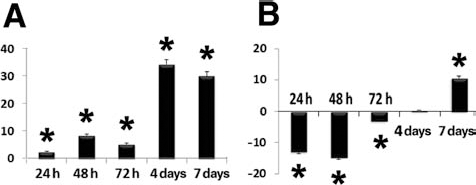
REAC exposure selectively activates the transcription of the Nox4 subunit of NADPH oxidase. Cells were exposed for 24 or 48 or 72 h in the absence or presence of REAC or were treated with REAC for 72 h and then left unexposed for additional 4 or 7 days. The amounts of nicotinamide adenine dinucleotide phosphate (NADPH) oxidase subunit 4 (Nox4) (A) and Nox1 (B) mRNA from REAC-treated or untreated cells were normalized to GAPDH, and the mRNA expression of REAC-exposed cells was plotted at each time point as fold change relative to the expression in control untreated cells, defined as 1 (mean ± SE; n = 6). ∗Significantly different from untreated cells.
On the whole, our findings indicated that RECF energy may be delivered with an ad hoc designed device to afford expression of pluripotentiality from human somatic cells. hSFs may be reprogrammed by a physical stimulus to acquire three differentiating outcomes (i.e., cardiogenic, neurogenic, and skeletal myogenic) that have long been pursued as major target lineages holding future promise for regenerative medicine in most morbidity-associated diseases, such as cardiovascular, neurodegenerative, and (neuro)-muscular disorders. So far, the acquirement of a pluripotent state in somatic cells, including fibroblast reprogramming to iPSCs or direct commitment to cardiac or neuronal fates without an iPSC intermediate, has only been achieved by targeted genetic engineering through viral vector-mediated technologies or cumbersome and expensive protein transduction methods or by combinatorial approaches encompassing both viral-mediated gene delivery and chemical stimulation of the target cells (7,8,17).
This is the first report demonstrating the feasibility to afford a rapid, long-lived, and inexpensive commitment of human fibroblasts to multiple complex lineages, without the use of intricate chemistry or viral transduction. Studies are on the way to dissect the electrophysiological properties of REAC-committed cells and to assess whether these cells may provide effective structural and functional rescue in defined animal models of heart failure, neurodegeneration, and skeletal muscle dystrophy. In the affirmative, the strategy proposed in the current study may pave the way to novel unprecedented approaches of regenerative medicine in humans.
Footnotes
Acknowledgments
We thank Dr. Eng. Matteo Lotti Margotti for data analysis, Lucia Aravagli M.D., and Stefania Bini M.D. of Rinaldi Fontani Institute, Florence, Italy, for their helpful support. This research was supported by Ministero della Salute, Italy, Programma per la Ricerca Sanitaria: attività di Ricerca sulle Cellule Staminali; Fondazione Fornasini, Poggio Renatico, Italy; Fondazione Cardinale Giacomo Lercaro, Bologna, Italy; Fondazione Luisa Fanti-Melloni, Bologna, Italy; Tavola Valdese, Rome, Italy. Salvatore Rinaldi and Vania Fontani are the inventors of the REAC. The other authors report no conflicts of interest.
