Abstract
The pig represents an ideal large-animal model, intermediate between rodents and humans, for the preclinical assessment of emerging cell therapies. As no validated pig embryonic stem (pES) cell lines have been derived so far, pig induced pluripotent stem cells (piPSCs) should offer an alternative source of undifferentiated cells to advance regenerative medicine research from bench to clinical trial. We report here for the first time the derivation of piPSCs from adult fibroblast with only three transcription factors: Sox2 (sex determining region Y-box 2), Klf4 (Krüppel-like factor 4), and c-Myc (avian myelocytomatosis viral oncogene homolog). We have been able to demonstrate that exogenous Pou5f1 (POU domain class 5 transcription factor 1; abbreviated as Octamer-4: Oct4) is dispensable to achieve and maintain pluripotency in the generation of piPSCs. To the best of our knowledge, this is also the first report of somatic reprogramming in any species without the overexpression, either directly or indirectly, of Oct4. Moreover, we were able to generate piPSCs without the use of feeder cells, approaching thus xeno-free conditions. Our work paves the way for the derivation of clinical grade piPSCs for regenerative medicine.
Keywords
Introduction
Embryonic stem cells (ESCs), as well as induced pluripotent stem (iPS) cells, hold great promises for cell therapy, drug screening, and in vitro disease models. A major concern of ESCs and iPS technologies is their long-term safety, whose assessment requires extensive preclinical studies in an appropriate animal model. Mice, currently the nearly exclusive animal model used in embryonic stem cell research, are unreliable to adapt for use in human clinical applications, considering their physiological differences with humans and their reduced life span. On the contrary, pigs are an excellent model for human diseases, especially for transplantation medicine, immunology, and circulatory system surgery (10, 16). Therefore, as research in cell replacement therapies advances, the need for a source of pig embryonic stem cells (pESCs) becomes pressing. Unfortunately, currently no pESC lines respond to conventional criteria for pluripotency.
Recently, three independent groups showed that pig induced pluripotent stem cell (piPSC) lines derived from fetal or young fibroblasts displayed variable expression of pluripotency-related genes, and tendency to differentiate spontaneously (5,6,21). Furthermore, they showed that the overexpression of the four transcription factors Pou5f1 (POU domain class 5 transcription factor 1; abbreviated as Octamer-4: Oct4), Sox2 (sex determining region Y-box 2), Klf4 (Krüppel-like factor 4), and c-Myc (avian myelocytomatosis viral oncogene homolog), together with the presence of feeder cells, was indispensable to achieve reprogramming.
We describe here for the first time that piPSC lines could be obtained by the ectopic expression of only Sox2, Klf4, and c-Myc. In our system, Oct4 was not only completely dispensable, but its overexpression affected negatively the pluripotent potential of reprogrammed cells. Moreover, the derivation of iPSCs from adult pig ear fibroblasts on a coating of gelatin proved to be of the same efficiency and speed as in the presence of irradiated mouse embryonic fibroblasts (iMEFs). Derivation of iPS lines on gelatin has so far only been achieved using mesenchymal stem cells as starting population, and never from more differentiated cell types, in any species (19). Overall, the results of our studies set the bases for xenobiotic (xeno)-free pig iPSC (piPSC) generation and provide a tool to test the clinical applicability of piPSC technology in a large animal setting.
Materials and Methods
Fibroblast Isolation
Fibroblasts used in this study were derived from ear biopsies of 6-month-old White Landrace x Large White pig females. Cells were cultured in high-glucose Dulbecco's modified eagle medium (DMEM; Invitrogen), 10% fetal bovine serum (FBS; Invitrogen), GlutaMAX (200 mM; Invitrogen), penicillin-streptomycin (100 U ml−1 penicillin and 100 μg ml−1 streptomycin; Sigma). All the experiments were performed with passage 3 adult fibroblasts. Ear biopsies were kindly provided by M. Esteves (General Surgery Department, University Hospital Vall d'Hebron, Barcelona)
Plasmid Constructions and Retroviral Production
The construction of the pMX-Sox2-Klf4-Myc-mOrange (pMXs-SKMmOrange) plasmid involved amplifying the mouse Sox2 complementary deoxyribonucleic acid (cDNA), eliminating the Sox2 stop codon and adding a BspEI site and cloning into the plasmid pCRII (Invitrogen) to obtain pCRII-Sox2-Bsp (oriented NotI-5′cDNA3′-Acc65I). The mouse Klf4 cDNA was amplified using a forward primer containing an AgeI site followed by the P2A peptide sequence and a reverse primer eliminating the Klf4 stop codon and containing a BspEI site; this fragment was cloned into pCRII to produce pCRII-Age-Klf4-Bsp (oriented NotI-5′cDNA3′-Acc65I). pCRII-Age-Klf4-Bsp was cut by AgeI and Acc65I and cloned into pCRII-Sox2-Bsp cut at BspEI-Acc65I producing pCRII-Sox2-P2A-Klf4-BspEI. The same cloning strategy was repeated twice, to incorporate the Myc and mOrange (18) coding sequences (producing pCRII-SKMmOrange) into the polycistron; the final construct is therefore pCRII-Sox2-P2A-Klf4-P2A-myc-P2A-mOrange. The last open reading frame (ORF; mOrange) was cloned preserving the stop codon. pCRII-SKMmOrange was cut with EcoRI and the SKMmOrange insert was cloned into a unique EcoRI site located in the polylinker of the pMXs vector, giving rise to pMXs-SKMmOrange (hence referred to as SKM-Or) (Fig. 1a). The pMXs-Oct4-Sox2-Klf4-Myc-Green FluorescentProtein (hence referred to as OSKM-Gr) plasmid generation was described previously in Gonzalez et al. (7). Single retroviral plasmids consisting of pMSCV-based retroviral vectors expressing FLAG-tagged OCT4, SOX2, KLF4, and c-MYC are available from Addgene (20072, 20073, 20074, and 20075, respectively). Retroviruses were produced as previously reported (1,14).

Time line of adult pig fibroblast reprogramming onto irradiated mouse embryonic fibroblasts (iMEFs). (a) pMXs-SKM-mOrange (hence referred to as SKM-Or) construct expressing sex determining region Y-box 2 (Sox2; S), Krüppel-like factor 4 (Klf4; K), myelocytomatosis viral oncogene homolog (Myc; M), and mOrange (Or) linked by P2A peptide sequences from a polycistron was used for the generation of pig induced pluripotent stem cell (piPSC) lines. (b) One day before (T = time; (T = −2) starting viral transduction (T = 0) 80,000 cells were seeded in every six-well format well. Three days postinfection (T = 1) cells were transferred on feeders. Typical stem cell-like colonies are clearly visible after 4 more days (T = 5). Representative phase contrast and fluorescence images of SKM-Or piPSC lines after 8 more days (T = 9). Scale bar: 50 μm. Alkaline phosphatase (AP) staining of SKM-Or piPSC lines. Magnification 3.6x. (c) Flow cytometry analysis confirming that pig fibroblast acquired the pluripotency marker stage-specific embryonic antigen 4 (SSEA-4) during the reprogramming process. Note that at T = 5 around 30% of the cells express the pluripotency marker SSEA-4. (d) pMXs-OSKMGFP (hence referred to as OSKM-Gr) construct expressing Pou5f1 (POU domain class 5 transcription factor 1; abbreviated as Octamer-4: Oct4: O), Sox2, Klf4, Myc, and green fluorescent protein (GFP; Gr) linked by P2A peptide sequences from a polycistron was used for the generation of piPSC lines. Flow cytometry analysis shows that pig fibroblasts infected with OSKM-Gr construct acquired low expression of the pluripotency-related protein SSEA-4 during the reprogramming process. Note that at T = 5 only around 2.51% of the cells express the surface protein SSEA-4.
Transduction of Porcine Fibroblast Cells
Pools of 80,000 fibroblasts were infected three times at 12-h intervals with retroviruses carrying polycistronic vectors and centrifuged at 750 rcf (relative centrifugal force) at 32°C for 45 min. One day after the last infection (T = time; T= 1), cells were trypsinized and plated onto iMEFs or gelatin 0.1% (Millipore) in human embryonic stem media (hESM)/mouse embryonic stem media (mESM) (1:1). hES medium: Knockout (KO)-DMEM (Invitrogen), KO serum replacement (KOSR; 20%, Invitrogen), GlutaMAX (1 mM; Invitrogen), penicillin-streptomycin (100 U ml−1 penicillin and 100 μg ml−1 streptomycin; Sigma), nonessential amino acid solution (100 μM; Invitrogen), 20 %FBS (Invitrogen), 2-mercaptoethanol (100 μM; Invitrogen), recombinant human basic fibroblast growth factor (bFGF, 10 ng ml−1; PeproTech). mESM media: High-glucose DMEM (Invitrogen), leukemia inhibitory factor (LIF; 1000 U/ml; Chemicon), 15% FBS, GlutaMAX (1 mM), penicillin-streptomycin (100 U ml−1 penicillin and 100 μg ml−1 streptomycin), nonessential amino acids (100 μM), 2-mercaptoethanol (100 μM).
Fluorescence Activated Cell Sorting Analysis
For surface phenotyping and fluorescent protein production anti-stage-specific embryonic antigen 4 (SSEA-4) coupled to AlexaFluor (AF) 647 (MC813-70), anti-pig-integrin β1 chain (CD29) unconjugated (NaM160-1A3), and anti-mouse immunoglobulin (MsIgG) AF488 from Invitrogen were used. The specificity of the staining was verified by the use of control monoclonal antibodies of matched isotype. Characterization was performed in a total of 10,000 events. Hoechst 33258 (H258) was added at 1 μg/ml in the final wash to detect dead cells. For deoxyribonucleic acid (DNA) analysis, samples were processed using the FIX & PERM kit from Invitrogen and H258 dye was used at 10 μg/ml. For the cell cycle and proliferation analysis, Click-iT EdU AlexaFluor 647® kit and anti-GFP AF488 from Invitrogen were used. All analyses were performed on a MoFlo cell sorter (DakoCytomation) running three lasers [488 nm blue laser (30 mW), 350 nm UV laser (20 mW), and 633nm red laser (25 mW)], applying Summit software.
Molecular Analysis of piPSC Lines
Integration of the transgenes was checked by polymerase chain reaction (PCR) using three primer pairs spanning the OSKM-Gr sequence as previously described (7). One μg of ribonucleic acid (RNA) from SKM-Or piPSC and OSKM-Gr piPSC lines and fibroblasts were used for reverse transcription (RT)-PCR (SuperScript II RT kit, Invitrogen). The following PCR protocol was performed: 10 min denature step at 95°C followed by 40 cycles at the specific temperature recommended for each previously described primer pair (4—7,21) for 30 s, annealing step for 30 s, and elongation at 72°C for 45 s with a final extension at 72°C for 10 min. PCR products were resolved in a 1% agarose gel containing 1 μg/ml ethidium bromide. One μl RT reaction was used to quantify gene expression by real-time quantitative PCR (qPCR) using previously described primers (4—7,21). Quantitative PCR analysis was done in triplicate using Platinum SYBR Green qPCR Super Mix (Invitrogen) in an ABI Prism 7000 thermocycler (Applied Biosystems), and values were normalized to β-actin. The conditions for real-time RT-PCR were as follows: 50°C, 2 min; 95°C, 10 min; followed by 40 amplification cycles (95°C, 15 s; 60°C, 30 s). The reaction was terminated by an elongation and data acquisition step at 75°C for 1 min.
Staining and Immunohistochemistry
Cells were fixed in 4% paraformaldehyde. The following antibodies were used: red fluorescent protein (RFP; Abcam), green fluorescent protein (GFP; AVES), Nanog (Everest Biotech), tumor rejection antigen 1 (TRA-1) TRA-1-60 (MAB), TRA-1-81 (MAB), SSEA-4 (Hybridoma Bank), SSEA-3 (Hybridoma Bank), α1 fetoprotein (AFP; Dako), Forkhead box protein A2 (FOXA2; R&D System), neuron-specific class III β-tubulin (Tuj1; Covance), nestin (Hybridoma Bank), glial fibrillary acidic protein (GFAP; Dako), smooth muscle actin (SMA; Sigma), sex determining region Y-box 9 (Sox9; R&D Systems), vimentin (VIM; Sigma). Direct alkaline phosphatase (AP) activity was analyzed using AP blue/red membrane substrate solution kit (Sigma) according to the manufacturer's specification.
High-Resolution, Giemsa-Banded Karyotype
Giemsa-banded (G-banded) karyotype analysis was performed on 80% confluent cells growing on Matrigel (BD Systems Ltd). Cells were treated with colcemid at 20 ng/ml, followed by a 45-min incubation at 37°C. Then, after trypsinizing the cells into a single cell suspension, they were treated with Carnoy fixative at −20°C. Finally, the samples were analyzed with the software Cytovision (Applied Imaging).
In Vitro Differentiation
In vitro differentiation assay was performed by embryoid bodies (EBs) formation according to standard protocols (1,7). Colonies were detached from the iMEF feeder layer or gelatin-coated wells by manual dissection and transferred into differentiation medium (high-glucose DMEM, 20% FBS, 2 mM GlutaMAX) in low attachment plates (Corning). After 3 days of culture, EBs were transferred onto adherent, gelatin-coated tissue culture dishes and were cultured in differentiation medium supplemented with either 10 mM ascorbic acid (for induction of mesoderm differentiation), differentiation medium (endoderm differentiation), or 1 μM all-trans retinoic acid (ectoderm differentiation) and subsequently fixed for immunofluorescence analysis 15 days later.
In Vivo Differentiation
Severe combined immune deficient (SCID)-beige male mice, approximataely 8 weeks old, were injected with piPSCs (~1 million for each injection site) resuspended in phosphate-buffered saline (PBS) in the testicular parenchyma. All procedures involving animals were approved by the institutional Animal Ethical Board, and the protocols were approved by the Conselleria De Salut of Cataluña. The mice were briefly anesthetized with a gaseous mixture of halothane/oxygen/ambient air of 0.5:2:97.5. Injections were performed using a 1-ml syringe with a G25 needle. No more than 150 μl of cells were injected in each site. The animals were left to recover from anesthesia on a warm mat at 37°C, and caged individually after injections. The mice were sacrificed 12 weeks after the injections, or when a tumor was detected by palpation, whichever came first. Following this protocol a total of 10 mice were injected with 5 SKM-Or piPSC lines.
Analysis of the Tumors
Mice were euthanized by cervical dislocation. Teratomas were dissected and fixed in 4% paraformaldehyde in PBS overnight at 4°C with agitation. After fixation, tumor samples were embedded in paraffin. Subsequently, samples were sliced in 5—10-μm sections and mounted on glass slides. Slides were stained with hematoxylin and eosin following standard protocol and scored for the presence of derivatives of all three embryonic germ layers. Positive samples were selected and analyzed by immunohistochemistry. Briefly, after washing the slides in PBS containing 0.01% of Triton X-100 (TBS solution), unspecific signal was blocked with TBS 0.5% in the presence of 3% goat serum for 30 min. After blocking, slides were incubated with 1 ml of primary antibody in PBS containing 0.1% Triton X-100 and 3% goat serum overnight at 4°C. Antibodies listed in the Staining and Immunohistochemistry section were used to identify the three germ layers. Sections were washed in PBS containing 0.1% Triton X-100 and 3% goat serum solution. Next, 1 ml of the same buffer containing the specific secondary antibody was added for 1 h at 37°C. Finally, slides were washed in TBS solution. Before mounting, DAPI solution was added for 10 additional min.
Results
Generation of piPSC Lines
A polycistronic retroviral vector expressing three reprogramming factors Sox2, Klf4, c-Myc, and the mOrange protein was used to infect adult porcine fibroblast (Fig. 1a). One day after the last viral transduction (T = 1), cells were transferred onto iMEFs with hESM: mESM. After 8 days (T = 9) compact colonies appeared. The colonies were comprised of cells exhibiting high nucleus to cytoplasm ratio with prominent nucleoli, together with positive AP staining (Fig. 1b). Next, we evaluated if the acquisition of the surface protein SSEA-4 correlated positively with the appearance of the colonies, and found that the percentage of positive SSEA-4 cells increased as the reprogramming process progressed (Fig. 1c). On the contrary, when reprogramming was carried out with a polycistronic retroviral vector expressing the four reprogramming factors Oct4, Sox2, Klf4, c-Myc, and GFP (OSKM-Gr), we observed a smaller percentage of SSEA-4-positive cells, which appeared later during the reprogramming process (Fig. 1d). Thus, although both polycistronic constructs gave rise to pESC-like colonies, SSEA-4-positive cells appeared consistently sooner when using the SKM-Or transgene (Fig. 1 c, d). We therefore investigated whether this could be ascribed to Oct4 overexpression.
Overexpression of Oct4 Interferes Negatively with the Expression of Pluripotency-Related Genes in Pig Fibroblasts
In order to understand how reprogramming occurred in our system, we decided to run the same approach except that, at T = 1, transduced cells were seeded directly onto gelatine with hES/mES medium. Surprisingly, colonies appeared in the same time frame as in the presence of iMEFs (Fig. 2a). To assess if Oct4 overexpression was negatively interfering with the acquisition of a proper pluripotent phenotype in our system, we performed a comparative assay by infecting the cells with either SKM-Or or OSKM-Gr retroviral particles. At T = 1, retroviral transduced cells were sorted according to the respective fluorescent protein production (mOrange or GFP, respectively) (Fig. 2b) and cells were plated onto gelatin in a 1:1 ratio of each cell type (40,000 cells/type). We assayed the resulting population for specific fluorescence, SSEA-4 acquisition, and cell cycle pattern at different time points thereafter. We observed again that the SSEA-4-positive population appeared later during the reprogramming process in the OSKM-Gr population (Fig. 2c). However, the results indicate that the lack of exogenous Oct4 expression does not alter the cell cycle of fibroblasts, calculated by interpolating the cell cycle pattern with the absolute number of cells expressing each reporter, as well as the ratio between fluorochromes (Fig. 2c). While OSKM-Gr piPSCs were prone to spontaneous differentiation after a few passages, SKM-Or piPSCs could be grown undifferentiated for an extended period of time (upwards of 30 passages at the time of writing). We speculated that this difference might be due to the larger size of the OSKM-Gr vector, as larger plasmids display lower transcriptional efficiency in transfected cells. Therefore, we performed a series of reprogramming experiments using a stechiometric mix of monocystronic constructs (data not shown), each carrying one of the transcription factors. The results overlapped with those obtained with the polycistronic constructs, indicating that a difference in plasmid size between the two polycistronic constructs does not affect the efficiency of reprogramming or the level of stability of the piPSC lines obtained.
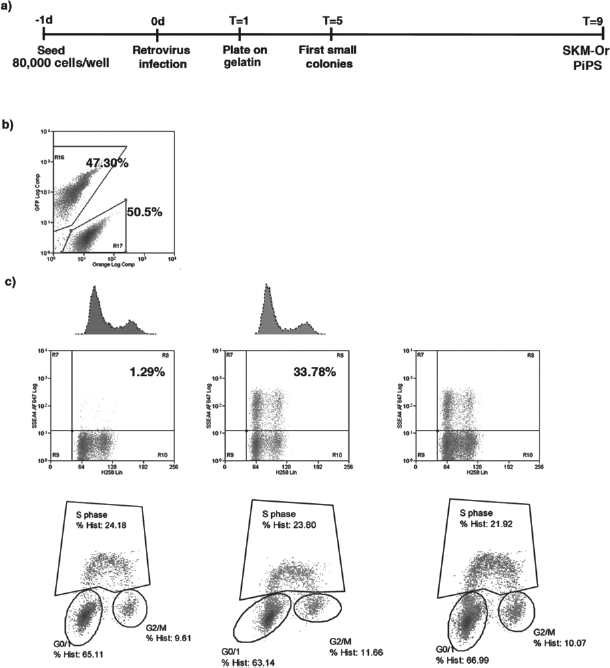
(a) Time line of adult pig fibroblast reprogramming onto gelatin. (b) Comparative cell cycle analysis of pMXs-OSKMGFP (hence referred to as OSKM-Gr) and pMXs-SKMmOrange (hence referred to as SKM-Or) expressing cells plated at a 1:1 ratio 2 days after infection. While the overall cell cycle profile and the total number of cells remain constant between the two compartments, SSEA-4-positive cells show a significant increase in the SKM-Or population, indicating more efficient reprogramming in the absence of exogenous Oct4.
Expression of Pluripotency-Related Genes in piPSC Lines
From day 9 in culture, tightly packed colonies with morphology similar to human ES cells appeared. These results were observed using both iMEFs and gelatin substrates. Established iPS cell lines were subjected to a PCR of genomic DNA by using two pair of primers (GFP, OS-K1/2 primers) designed to amplify specific regions of the pMXs-SKMmOrange vector, and we found that all clones analyzed had integrated the transgene in their DNA (Fig. 3a) as described previously (7). Several pluripotency associated genes have been described to be expressed in the inner cell mass (ICM) of pig embryos [e.g., Oct4, Sox2, Tdgf-1 (teratocarcinoma-derived growth factor 1; Cripto), and Tert (Telomerase), among others] (2,3,8). In our hands, quantitative PCR revealed that all the tested clones had reactivated the transcription of a set of endogenous pluripotency markers including Oct4, Sox2, and Nanog (Fig. 3b). Previous studies have reported the isolation of piPS cells based on both cell morphology and immunostaining with the embryonic surface proteins SSEA-3, SSEA4, TRA-1-60, TRA-1-81 (19). Indeed, immunofluorescence analysis showed that SKM-Or and OSKM-Gr piPSC lines expressed ESC-related proteins (Fig. 3c, d, respectively). In addition, we could confirm that under our culture conditions both SKM-Or and OSKM-Gr piPS display a normal karyotype (Fig. 3e).

(a) Genomic deoxyribonucleic acid (DNA) polymerase chain reaction (PCR) confirming the insertion of GFP (top) and Sox2-Klf4 transgenes (bottom) in OSKM-Gr#2 (# = number) and SKM-Or#1, 7, 8, 14, 15 piPSC lines. (b) Real-time quantitative PCR (qPCR) analysis for pluripotency-related genes, endogenous Oct4 (Endo Oct4), endogenous Sox2 (endo Sox2), Nanog, and Sox2-Klf4 transgenes (Trans Sox2-Klf4). OSKM-Gr#2 and SKM-Or#7 and SKM-Or#8 piPSC lines were analyzed together with pig fibroblast. (c, d) Immunofluorescence analysis of piPSC lines for pluripotency-related proteins. (c) SKM-Or piPSC lines express SSEA-4, tumor rejection antigen 1 (TRA-1) TRA-1-81, TRA-1-60, and the transcription factor Nanog. Notice that red fluorescent protein (RFP) expression indicates that the transgene is not silenced. 4′, 6-Diamidino-2-phenylindole (DAPI) is used for nuclear stain. Scale bar: 100 μm. (d) OSKM-Gr piPSC lines express SSEA-3 and the transcription factor NANOG. Notice that GFP expression indicates that the transgene is not silenced. Scale bar: 50 μm. (e) High-resolution, G-banded karyotype indicating a normal diploid female chromosomal content in SKM-Or#1 and OSKM-Gr#2 piPSC lines analyzed at passage 11.
Embryoid Bodies Formation: In Vitro Differentiation
We tested whether piPSC lines could form embryoid bodies (EBs) in the absence of bFGF and LIF on a non-adhesive substrate. All piPSC lines were examined, and all differentiated into EBs under such conditions. When placed on a gelatin-coated surface and cultured in the presence of mesoderm-, ectoderm-, or endoderm-inducing media, EBs readily attached to the substrate, began to spread, and achieved differentiation. In all cases, the SKM-Or and OSKM-Gr piPSC lines readily differentiated into derivatives of the three embryonic germ layers in vitro, as they acquired specific markers representing all three germ layers after each specific protocol already detailed in Materials and Methods (Fig. 4a, b).

(a) Representative images of in vitro differentiation of SKM-Or piPSC line into the three primary germ cell layers [endoderm-forkhead box protein A2 (FOXA2), ectoderm-nestin, and mesoderm-smooth muscle actin (SMA)]. Scale bar: 50 μm for endoderm and ectoderm differentiation images. Scale bar: 100 μm for mesoderm differentiation images. (b) Representative images of in vitro differentiation of OSKM-Gr piPSC line into the three primary germ cell layers [endoderm-FoxA2, ectoderm-neuron-specific class III beta-tubulin (Tuj-1), and mesoderm-sex determining region Y-box 9 (Sox9)]. Scale bar: 50 μm.
Teratoma Formation: In Vivo Differentiation
We tested the differentiation potential of five SKM-Or piPSCs lines in vivo by teratoma formation assay. When injected into the testicles of SCID-beige mice (two mice for every one of the five lines tested), two of the five SKM-Or piPSCs lines tested were able to form teratomas in both mice, demonstrating the potential of SKM-Or piPSCs to differentiate in vivo into the three germ layers of the embryo. The teratomas presented differentiation areas indicative of cells deriving from each of the three embryonic germ layers: endoderm, ectoderm, and mesoderm (Fig. 5a, b)

Teratoma formation assay by injection of SKM-Or piPSCs into the testis of severe combined immune deficient (SCID)-beige male mice. (a) After 12 weeks, teratomas were collected and analyzed by hematoxylin and eosin staining. piPSCs differentiated in vivo to endoderm (*), ectoderm (**), or mesoderm fates (***). Representative images of in vivo differentiation of SKM-Or piPSC line into the three primary germ cell layers by immunofluorescence. Endoderm-α1-fetoprotein (AFP). Ectoderm-glial fibrillary acidic protein (GFAP), neuron-specific class III β-tubulin (Tuj-1). Mesoderm-vimentin (VIM) and sex determining region Y-box 9 (Sox-9). Scale bar: 25 μm for endoderm differentiation images. Scale bar: 50 μm for ectoderm differentiation images. Scale bar: 75 μm for mesoderm differentiation images.
Discussion
The pig is a widely used model of human diseases and therapies, especially for the immune and circulatory systems (10,16), and for transplantation assays, because of its physiological similarity with the human organism and its extended lifespan compared to other animal models such as the mouse. Regenerative medicine research has received a strong impulse by the derivation of iPSCs in humans. In this context, performing human-scale cell replacement assays in pig becomes a necessity. Currently, the most frequent sources for cell replacement research are cell lines of embryonic origin. However, no validated pESC lines could be derived so far from ungulates, thus limiting the scope of regenerative medicine research in these species. The development of genetically and epigenetically stable piPSC lines would represent an ideal tool to study clinically relevant models in a large-animal species as well as to gain precious information concerning the characteristics of porcine pluripotent cells.
To mimic a therapeutic context, the piPSC lines would have to be free of contaminating cells and reagents from other species. We were able to generate piPSCs without the use of feeder cells, bringing our protocol for derivation one step closer to xeno-free conditions. Moreover, our use of adult fibroblast for reprogramming, instead of a cell type of embryonic or fetal origin, will enable us to test the functionality of self-iPSCs in a therapeutic scenario of autologous transplantation. Further research will be needed to identify a nonintegrative, transient expression technique to induce piPSCs and to develop a defined culture medium for pluripotent pig cells in order to fulfill therapeutic requirements.
To date, a few groups have generated piPSCs using integrating retroviral vectors expressing Oct4, Sox2, Klf4, and c-Myc (5,6,21). Oct4 is essential for the identity of the pluripotent founder cell population in the mammalian embryo (13,15,17), and its expression, either direct (20) or indirect (9), is an absolute requirement for the acquisition of pluripotency in all somatic cells studied so far. The piPSC lines that we generated without Oct4 overexpression, however, follow the standard criteria for pluripotency, differentiate in vitro along multiple tissue lineages, have a stable karyotype, and express endogenous pluripotency-related genes. Moreover, when piPSCs cells were injected into the testis of SCID beige mice, they were able to form complex-structured teratoma containing tissues from the three germ layers (ectoderm, mesoderm, endoderm). However, in our hands, only two among five SKM Or piPSC lines showed such potential. This result could be ascribed to the presence of an active transgene in the injected cells. Transgene expression might not be a strong inhibitory factor when cells are differentiated in presence of strong stimuli such as cytokines in specific mediums in vitro, but it might offer a sufficiently high barrier to differentiation when placed in a less differentiation-specific environment. These results confirm that teratoma assay should be carried out for complete characterization of iPS lines, especially when generated with integrative strategies, in never before tested species or donor cell types.
It is interesting to note that, in in vitro-cultured pig embryos at the stage of blastocysts, Oct4 has been detected in the trophectoderm as well as in the inner cell mass (ICM) (2,3,8), while Oct4 expression in monkey, human, and mouse blastocysts is restricted to the ICM (11,12), suggesting a less restricted role of Oct4 in pig development. On the other hand, it has been reported that pig fibroblast express low but detectable levels of endogenous Oct4 (4). This observation reinforces our hypothesis of a broader role for Oct4 in pig development and cell biology compared to other species. Furthermore, it offers a possible explanation for the easiness with which pig fibroblasts seem to reprogram, as piPSC colonies appear sooner and with greater efficiency than all other cells reprogrammed so far, and do not need feeder cells to reprogram. Because of the lack of reliable sequence data on the pig transcription factors, we have used mouse factors to reprogram pig fibroblast. This procedure is not uncommon, and mouse factors have been shown repeatedly to reprogram cells of other mammalian species. Although we cannot exclude formally that the presence of mouse protein during reprogramming of pig fibroblasts might have a role in the outcome of our experiments, this does not seem likely in the context of the broader scientific literature.
To the best of our knowledge, this is the first report of induced pluripotency of terminally differentiated cells in any species without overexpression of Oct4, either directly or indirectly, and without the use of feeder cells. Further studies will be needed to identify in detail the molecular mechanisms underlying porcine somatic reprogramming.
Footnotes
Acknowledgments
We are grateful to Meritxell Carrió, Laetitia Casano, and Vanesa Tobajas Fernandez for expert assistance with cell culture techniques. José Miguel Andrés Vaquero for assistance with flow cytometry. Lola Mulero Pérez, Cristina Morera, Cristina Pardo, and Mercé Martí Gaudes for bioimaging assistance, Cristina Gómez for expert assistance in molecular biology techniques. NM was partially supported by Juan de la Cierva Program, EG was partially supported by Sara Borrell Program. This work was partially supported by grants from MINECO, Fondo de Investigaciones Sanitarias (TERCEL, PI052847), Fundación Cellex and the G. Harold and Leila Y. Mathers Charitable Foundation. The authors declare no conflicts of interest.
