Abstract
BACKGROUND:
ErbB/PI3K signaling is widely recognized as a critical modulator of malignancy and miRNAs have been found to play a crucial role in the regulation of this pathway.
OBJECTIVE:
This study aimed to identify novel miRNAs related to the ErbBs loci and investigate the functional effects of these miRNAs on ErbB/PI3K signaling in cancer progression.
MATERIALS and METHODS:
Bioinformatics tools and RNA-seq data were used to discover novel miRNAs in breast and colon cancer cells. Gene expression levels were determined using RT-qPCR. Western blotting and dual-luciferase assays were used to identify the regulatory mechanism between ErbB4-miR1/2 and related genes. The effects of ErbB4-miR1/2 on cell proliferation, viability, ROS production, and migration were assessed by PI-flow cytometry, colony formation, MTT, ROS, scratch, and transwell assays in SKBR3 and SW480 cells.
RESULTS:
MicroRNA prediction tools, RNA-seq data, RT-qPCR, and sequencing results identified ErbB4-miR1 and ErbB4-miR2 (ErbB4-miR1/2) as novel miRNAs encoded by
CONCLUSIONS:
ErbB4-miR1/2 are novel tumor suppressor miRNAs which attenuate ErbB/PI3K signaling in breast and colon cancer cells.
Abbreviations
microRNA Untranslated Region Phosphoinositide 3-Kinase Polymerase Chain Reaction Quantitative Reverse Transcription PCR Dulbecco’s Modified Eagle Medium Fetal Bovine Serum Bovine Serum Albumin Polyvinylidene Difluoride Propidium Iodide Enzyme-Linked Immunosorbent Assay 3-(4, 5-Dimethylthiazol-2-yl)-2, 5-Diphenyltetrazolium Bromide Epithelial Mesenchymal Transition Fluorescence-Activated Cell Sorting Standard Error of Mean Sequence Read Archive University of California, Santa Cruz The National Center for Biotechnology Information
Introduction
One of the most important challenges in the diagnosis and treatment of cancer is the genetic changes and epigenetic events in cell signaling pathways accompanied by considerable cellular and molecular heterogeneity. Despite tremendous advances in cancer research, the molecular details and mechanisms underlying cancer progression are not well understood. Hence, to develop novel therapeutic approaches for the prevention of cancer progression and advances in its diagnosis, prognosis, and treatment, it is important to recognize the genes and molecular pathways involved in tumor growth and carcinogenesis.
Homo- and heterodimerization of ErbB family receptors activate downstream PI3K/AKT signaling cascade [1, 2]. ErbB2, in combination with ErbB3, activate PI3K/AKT, which is the most important downstream pathway in cancer triggered by the ErbB family [3, 4]. The hyperactivation of ErbB/PI3K signaling has been reported in many types of human cancers [3, 5–7]. Thus, targeting the regulators of this pathway has the appealing therapeutic potential [8, 9]. Hence, ErbB receptors and key components of the PI3K pathway are promising targets for cancer diagnosis and treatment. Theoretically, a deeper molecular understanding of the regulatory elements related to the ErbB/PI3K signaling pathway may lay grounds for strategies to prevent cancer progression.
MicroRNAs (miRNAs) are a group of small non-coding RNAs that post-transcriptionally regulate target gene mRNAs by binding to their 3’-UTRs, leading to translational suppression or mRNA degradation [10]. Aberrant miRNA expression is thought to lead to carcinogenesis by regulating critical cancer-related genes and signaling pathways [11–13].
In recent years, technological developments such as bioinformatics have enabled the discovery of many miRNAs. To date, more than 38,589 hairpin precursors and 48,860 mature microRNAs have been recorded in miRBase (release22, 2018; www.mirbase.org) [14]. However, the human genome is thought to contain approximately 55,000 miRNAs [15]. Genetic approaches for identifying novel miRNAs are challenging because of their small size and tolerance for mutations that might alter their seed sequences [16]. Hence, several bioinformatics tools, such as RNAfold [17] and Mfold [18], have been developed to scan genomic sequences for miRNA prediction. Since the first discovery of miRNA, various computational and experimental methods have been used to find putative miRNAs, and the advent of RNA-Seq technology has substantially facilitated the identification of new miRNAs [16, 20].
Several studies have demonstrated the regulatory effects of miRNAs on ErbB/PI3K signaling [21–24]. These miRNAs’ multi-layered regulation of the signaling pathway illustrates their potential for diagnosis and cancer treatment.
In the present study, we aimed to identify novel miRNAs located in the
Materials and methods
In silico analysis
The workflow for the present study was as follows: (1) The genomic sequence of the ErbB4 gene (distance and orientation of this gene located in Chr2q34 were surveyed by the Ensembl database (release 97 based on hg38, https://www.ensembl.org)) was scanned and pairing characteristics of the predicted hairpins and RNA secondary and stem-loop structures were analyzed using RNAfold web server (https://rna.tbi.univie.ac.at/cgi-bin/RNAWebSuite/RNAfold.cgi) based on the minimum free energy algorithm. (2) Conservation status of the predicted miRNAs was analyzed by the University of California Santa Cruz (UCSC) Genome Browser (2013, GRCh38/hg38, https://genome.ucsc.edu/) among 100 vertebrates. (3) Using small RNA-seq data from ENCODE/Cold Spring Harbor Lab track in the UCSC Genome Browser and NCBI BLAST search against several SRA miR-Seq data (Sequence Read Archive; https://www.ncbi.nlm.nih.gov/sra; SRX2154261, SRX2069669, SRX2069668, SRX130232, ERX1217535, SRX2069616, SRX2069617, SRX2154245), the existence, sequence, actual length, and expression of the novel miRNAs were obtained. (4) There were no identical or similar miRNA sequences to any of the recorded miRNAs to ErbB4-miR1 in the miRBase database. ErbB4-miR2 showed high sequence similarity with hsa-miR-744-5p. (5) The sequence of the miRNAs (cloned in TA vector) was confirmed using Sanger sequencing. (6) Potential target genes for ErbB4-miR1 and ErbB4-miR2 were predicted using Diana-MR-microT (based on Ensembl v84, https://mrmicrot.imsi.athenarc.gr/) and miRTar online analysis tools (https://mirtar.mbc.nctu.edu.tw/) and validated by RNAhybrid software (version 2.2, https://bibiserv.cebitec.uni-bielefeld.de/rnahybrid).
Cell culture and tissue samples
Breast cancer cell lines (SKBR3, MDA-MB-231, and MCF7) and colorectal cell lines (Caco2, HT29, HCT116, and SW480) were grown in HDMEM (Gibco, USA) and human embryonic kidney (HEK) 293T cells were cultured in DMEM-F12 (Gibco, USA) supplemented with 10% FBS (Gibco, USA), 100 U/mL penicillin, and 100 U/mL streptomycin (Sigma, UK), and then incubated at 37°C with 5% CO2. Normal breast cell line (MCF10A) was cultured in DMEM-F12 (Gibco, USA) containing 5% horse serum (Gibco, USA), 10μg/ml insulin (Gibco, USA), 20 ng/ml epidermal growth factor (EGF) (Invitrogen, USA), 100 ng/ml cholera toxin (CT) (Calbiochem, USA), and 0.5μg/ml hydrocortisone (HC) (Sigma, USA). All cell lines were obtained from Pasteur Institute, Iran. All experiments were performed using passage numbers below 30. All transfections were performed using TurboFect Reagent (Invitrogen, USA) according to the manufacturer’s protocol.
Breast cancer tissues and adjacent normal breast tissues were obtained from Breast Cancer Research Center, Motamed Cancer Institute, ACECR, Tehran, Iran. Colorectal cancer tissues and normal ones were obtained from the Imam Khomeini Hospital, Iran. All tissue samples were collected, immediately snap-frozen in liquid nitrogen, and stored at –80°C for later experiments. Informed consent forms were obtained from all patients.
RNA extraction and quantitative real-time PCR
Total RNA from tissues and cultured cells was isolated using RiboEx reagent (Invitrogen, USA) following the manufacturer’s protocol. The integrity and purity of the extracted total RNA were evaluated using agarose gel electrophoresis and NanoDrop (Thermo Scientific, USA). The first-strand cDNA was synthesized from total RNA (1μg) using reverse transcriptase (Takara, China). For miRNA detection, polyA tail was added to the 3′ ends of RNAs before cDNA synthesis. RT-qPCR was performed using the Step Onetrademark Real-Time PCR System (Applied Biosystems, USA) and Real-Time PCR Master Mix (Biofact, USA). The expression levels and relative expression levels were calculated using the 2-ΔCt and 2-ΔΔCt methods, respectively. The gene-specific primers are provided in Table 1. The mRNAs and miRNAs expression levels were normalized against GAPDH or U48 small RNA as the internal controls, respectively.
The primers list for RT-qPCR
The primers list for RT-qPCR
PsiCHECK-2 dual-luciferase vector (Promega, USA) was used to clone the 3’-UTRs of all predicted ErbB4-miR-1/2 target genes. To study the direct interaction, HEK293T cells were co-transfected with wild-type (control) reporter plasmid (psiCHECK-2 construct) and ErbB4-miR-1/2 (mock vector) using TurboFect Reagent (Thermo Scientific). A fragment without any predicted microRNA recognition element (MRE) was used as a control for targets of the novel miRNAs. Luciferase activity was measured 48 h after transfection using a dual-luciferase reporter assay system (Promega, USA) according to the manufacturer’s protocol.
Western blot assay
The cells used for protein extraction were lysed on ice in RIPA Buffer (Cell Signaling Technology, USA) and the Bradford assay measured protein concentrations. Quantified proteins were separated on a 10% polyacrylamide gel and electro-transferred to polyvinylidene fluoride (PVDF, Thermo Scientific, USA) membranes. The membranes were blocked with 5% BSA (Roche, Germany) for 2 h and then incubated with primary antibodies against phospho-AKT (Santa Cruz, USA, #sc-271966), AKT (Elabscience, USA, #E-AB-30471), and β-actin (Santa Cruz, USA, #sc-47778) overnight at 4°C. After washing, the membranes were incubated at room temperature for 1 h with secondary HRP-conjugated antibodies (BIORAD, USA, #1721011) or mouse anti-rabbit (Santa Cruz, USA, #sc-2357). The bands were detected using an enhanced chemiluminescence (ECL) kit (Tanon, China) and analyzed with ImageJ software (version 1.52a). Relative expression levels of the indicated proteins were compared with β-actin expression.
MTT assay
SKBR3 and SW480 cells were seeded in a 96-well plate and cultured at 37°C with 5% CO2. 0, 24, 48, and 72 h after transfection with ErbB4-miR1/2 or mock control vectors, 20 μL MTT (5 mg/ml) was added. Following incubation at 37°C for 4 h, formazan crystals were dissolved in 200μL dimethyl sulfoxide (DMSO, Sigma). Following incubation at room temperature for 10 min, cell viability was measured using a microplate ELISA reader (ThermoFisher, USA) at 570 nm.
Cell cycle assay
The cells (SKBR3 and SW480) were transfected with ErbB4-miR1/2 or mock control vectors in 24 well plates using TurboFect Reagent (Invitrogen, USA) according to the manufacturer’s instructions. 48 h after transfection, the cells were harvested, washed with cold PBS, fixed in 70% ice-cold ethanol (Bidestan, Iran), incubated with propidium iodide (PI, Sigma, USA) and RNase A (Thermo Scientific, USA), and analyzed using a FACS flow cytometer (BD Biosciences, USA).
Colony formation assay
SKBR3 and SW480 cells were transfected with ErbB4-miR1/2 or mock control vectors in six-well plates and cultured at 37°C for 14 days in neomycin (Invitrogen, USA) supplemented medium. The cell colonies were counted by staining with 1% crystal violet and were photographed using a Nikon microscope. The colonies were quantified using ImageJ software (version 1.52a).
ROS assay
48 h after transfection with ErbB4-miR1/2 or mock control vectors, SKBR3 cells were mixed with dihydroethidium (DHE; Thermo Scientific, USA) solution and incubated at 37 °C for 1 h. The intensity of fluorescence was measured using FACS (BD Biosciences, USA) flow cytometry.
Wound-healing assay
SKBR3 and SW480 cells were seeded in a 24-well plate, and four hours after transfection, straight wound lines were created by scraping with a yellow tip. Cell movements toward the wound were photographed at 0, 24, 48, and 72 h after transfection using an Olympus inverted microscope (Japan). The distance between the edges was measured using ImageJ software (version 1.52a).
Transwell assay
SKBR3 and SW480 cells were seeded in 24-well plates and were transiently transfected with ErbB4-miR1/2 overexpressing or mock control vectors. After 12 h, the transfected cells were collected and 50,000 cells were added to the upper well of the transwell chamber. After 24 h of incubation, cells that had migrated to the lower surface were stained with 0.5% crystal violet solution, washed with PBS, and photographed.
Statistical analysis
All experiments were performed in triplicate, and all data were expressed as mean±SEM. Statistical significance analyses were evaluated using GraphPad Prism software (version 6.0) and determined by nonparametric
Results
Bioinformatics prediction of novel miRNAs encoded within the ErbB4 gene locus
The genomic sequence of the ErbB4 gene based on Ensembl (GRCh38), which is located on the 2nd human chromosome (chr2 : 211,375,717-212,538,802, minus strand), was searched for possible stem-loop structure formation. The pairing characteristics of the predicted hairpins were analyzed using an RNAfold web server. Several stem-loop structures have been predicted within this gene, which could generate putative mature miRNAs. These putative miRNAs appeared to have a typical conservation pattern. Among the predicted stem-loops, two (named ErbB4-miR1/2) located in the 4th intron of the

Bioinformatics prediction of novel miRNAs located in the
The mature ErbB4-miR1/2 expression levels were measured in both breast and colon cancer tissues and cell lines using RT-qPCR. The results indicated that the expression levels of ErbB4-miR1/2 in breast cancer specimens (10 samples) were strongly reduced compared to the matched adjacent non-tumor tissues. Also, the results showed that the expression levels of ErbB4-miR1/2 were significantly reduced in the nine colon cancer tissues compared to those in the normal tissues (Fig. 2A). Furthermore, ErbB4-miR1/2 expression has been investigated in several human breast and colon cancer cell lines. The expression levels of ErbB4-miR1/2 were lower in breast and colon cancer cell lines, including SKBR3, MDA-MB231, HCT116, and SW480 cells, than in the normal breast (MCF10A) and colon (Caco2) cell lines (Fig. 2B).
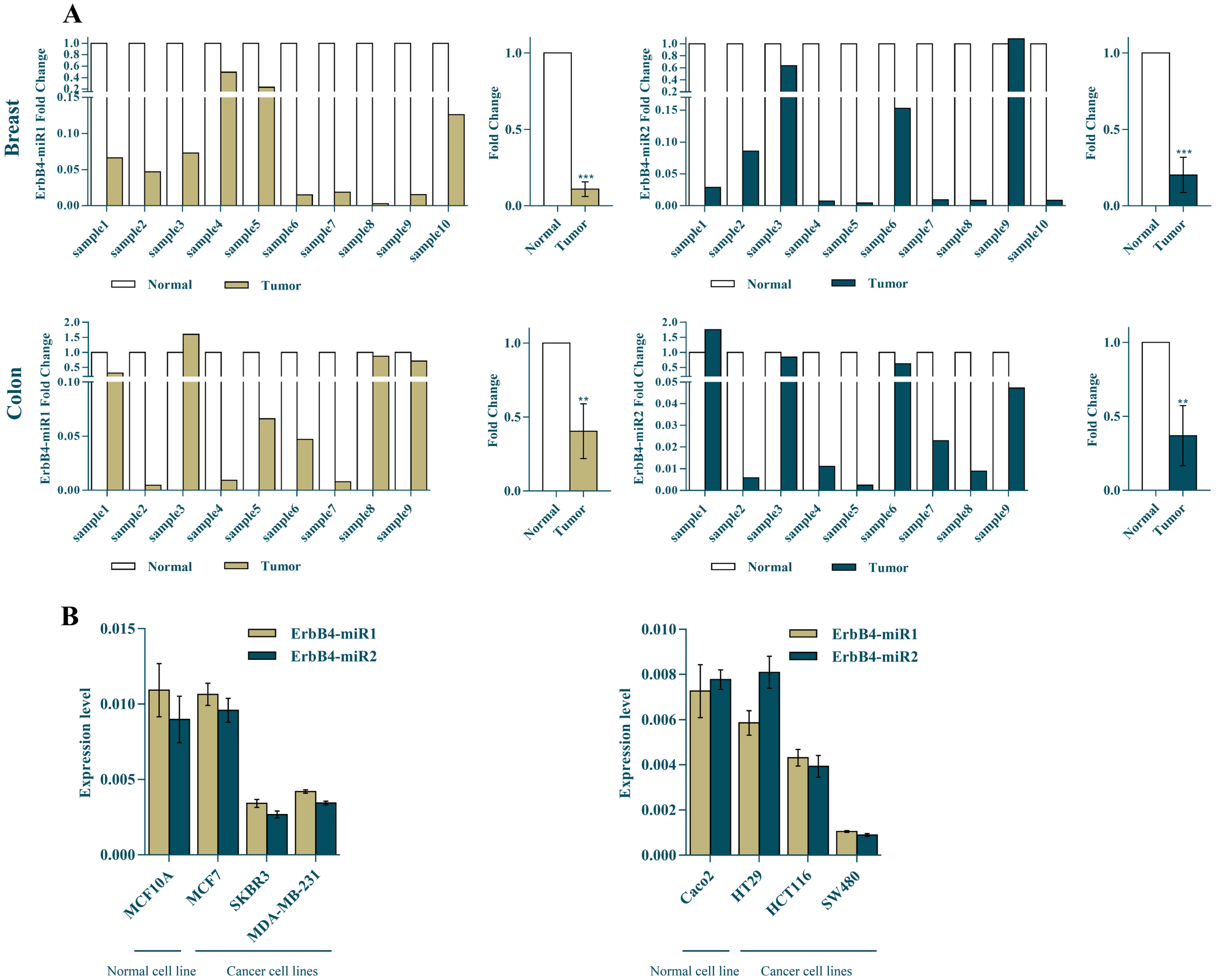
ErbB4-miR1/2 expression status in breast cancer and CRC tissues and cell lines. (A) Shows the ErbB4-miR1/2 expression fold change in human breast (
Overall, these results demonstrated that the ErbB4-miR1/2 expression was decreased in cancerous breast and colon tissues compared to the normal ones, and their expression was lower in more aggressive cell lines. These findings suggest that ErbB4-miR1/2 can act as tumor suppressors in both tested breast and colon cancers.
Potential target genes of ErbB4-miR1/2 were predicted using miRNA target prediction tools including DIANA-MR-microT, miRTar, and RNAhybrid. Using these prediction algorithms, several candidates were identified as common target genes for ErbB4-miR1 and ErbB4-miR2. Interestingly, strong complementary patterns were found between the 3’-UTR sequences of the ErbB2 and ErbB3 mRNAs and ErbB4-miR1/2 (Fig. 3A).

ErbB4-miR1/2 regulate the ErbB/PI3K signaling pathway. (A) Shows the predicted binding sites of ErbB4-miR1/2 in the 3’-UTRs of
The constructs containing the precursor of ErbB4-miR1/2 were transfected into breast and colon cancer cell lines and the relative expression levels of mature ErbB4-miR1 and ErbB4-miR2 were detected 48 h after transfection (Fig. 3B). Authenticity of the amplified products was confirmed by Sanger sequencing. These sequences were submitted to the NCBI database by NCBI accession numbers ON980707 and ON980708 for ErbB4-miR1 and ErbB4-miR2, respectively.
To investigate the effect of ErbB4-miR1/2 on ErbB/PI3K signaling, alteration in the expression of the genes involved in the pathway were examined in SKBR3 and SW480 cells 48 h after transfection with ErbB4-miR1/2 overexpressing vectors. RT-qPCR results indicated that ErbB2, ErbB3, AKT1, and AKT3 expression levels were significantly reduced in SKBR3 and SW480 cells following the ErbB4-miR1/2 overexpression, compared to mock-transfected cells (Fig. 3C).
Furthermore, a dual-luciferase reporter system was used to verify the direct interaction between the target genes and ErbB4-miR1/2. To this aim, the 3’-UTR sequence of each target gene was cloned downstream of the Renilla luciferase gene. Luciferase activity of ErbB2 and ErbB3 in HEK293 cells decreased after co-transfection of the 3’-UTR with ErbB4-miR1/2 relative to the mock and no-MRE control (a plasmid lacking any binding site for these miRNAs) vectors (Fig. 3D). To assess the effect of ErbB4-miR1/2 on the PI3K/AKT pathway, western blot analysis was performed to determine the p-AKT and total AKT protein levels. The results showed that the p-AKT/AKT ratio was reduced in SKBR3 cells overexpressing ErbB4-miR1/2, compared to the mock control vector (Fig. 3E). Overall, these findings suggested that ErbB4-miR1/2 regulate the ErbB/PI3K pathway by targeting the genes involved in this pathway.
To further validate the antitumor role of ErbB4-miR1/2 in breast and colon cancer, its effect on SKBR3 and SW480 cell viability was investigated using MTT assay. The results demonstrated that ErbB4-miR1/2 overexpression was followed by a significant reduction in SKBR3 and SW480 cell viability 24, 48, and 72 h after transfection (Fig. 4A).

Effects of ErbB4-miR1/2 overexpression on cell cycle, proliferation, and ROS production of breast and colon cancer cells. (A) MTT assay results of SKBR3 and SW480 cells at 0, 24, 48, and 72 h after transfection with ErbB4-miR1/2 or mock vectors. (B) Cell cycle histogram and bar plot analysis 48 h after transfection of SKBR3 and SW480 cells with ErbB4-miR1/2 or mock control vectors. (C) Colony formation assay showing the effect of ErbB4-miR1/2 versus mock vectors on SKBR3 and SW480 cell proliferation. (D) RT-qPCR analysis showing the effect of ErbB4-miR1/2 versus mock vectors on PCNA mRNA expression 48 h after transfection of SKBR3 and SW480 cells. (E) ROS production analysis in SKBR3 cells 48 h after transfection with ErbB4-miR1/2 or mock vectors. Assays were performed in triplicate. Means±SEM was shown. Statistical analysis was performed using Student’s
The inhibitory effect of ErbB4-miR1/2 on SKBR3 and SW480 cancer cell proliferation was also investigated. PI-flow cytometry cell cycle analysis revealed that ErbB4-miR1/2 overexpression compared to mock-transfected cells, resulted in a considerable increase in the sub-G1 cell population and a decrease in the G1 cell population (Fig. 4B). Consistently, ErbB4-miR1/2 overexpression significantly reduced the colony numbers of SKBR3 and SW480 cells (Fig. 4C) and PCNA (a cell cycle marker) gene expression (Fig. 4D) in the ErbB4-miR1/2 transfected cells compared to mock-transfected cells. Moreover, the intracellular levels of ROS product in the SKBR3 and SW480 cells were detected 48 h post-transfection with ErbB4-miR1/2 or mock control vectors. Flow cytometry analysis showed a significant increase in ROS formation in the cells overexpressing ErbB4-miR1/2, compared to the control (Fig. 4E). Taken as a whole, these data indicate that ErbB4-miR1/2 suppress cell proliferation and viability of SKBR3 and SW480 cells.
To investigate the effect of ErbB4-miR1/2 on SKBR3 and SW480 cell migration and the EMT process, wound-healing, transwell assays, and expression of epithelial to mesenchymal transition (EMT) markers were applied. The cell migration was observed under a microscope and photographed at different time points (0, 24, 48, and 72 h) after scratching. SKBR3 and SW480 mock-transfected cells showed accelerated wound healing whereas ErbB4-miR1/2-overexpressing cells exhibited reduced wound healing 48 and 72 h after transfection (Fig. 5A). Furthermore, to assess the effect of ErbB4-miR1/2 on EMT (which has been proposed as a key process in cancer progression),

Effects of ErbB4-miR1/2 overexpression on cell migration and EMT process. (A) Wound healing images and quantitative analysis of scratch wound closure in SKBR3 and SW480 cells transfected with ErbB4-miR1/2 or mock vectors at 0, 24, 48, and 72 h post scratching (40×magnification). (B) RT-qPCR analysis of epithelial and mesenchymal markers in SKBR3 and SW480 cell lines upon ErbB4-miR1/2 transfection compared to mock vector-transfected cells. (C) Representative images of transwell migration assay of SKBR3 and SW480 cells transfected with ErbB4-miR1/2 or mock vectors 48 h after transfection. Assays were performed in triplicate. Means±SEM was shown. Statistical analysis was performed using Student’s
The ErbB receptor family of RTKs belonging to the ErbB/PI3K pathway has emerged as an attractive candidate for anticancer therapy [25, 26]. Recent studies have demonstrated that miRNAs can regulate ErbB/PI3K pathway by targeting components of the pathway and play key roles in cancer initiation and progression [27–32]. Considering the current application of small molecules that block ErbB receptors to reduce PI3K signaling in clinical and preclinical cancer models [33–35], miRNAs could be considered as potential endogenous small molecules for this purpose.
Specific bioinformatics tools such as RNAfold [17], Mfold [18], and SSCprofiler [36] are used to predict stem-loop structures capable of encoding novel miRNAs, as their detection by forward genetics is reasonably inefficient. To identify novel microRNAs regulating ErbB/PI3K pathway genes, several bioinformatics tools have introduced ErbB4-miR1 and ErbB4-miR2 as two tandem novel putative miRNAs encoded to form the stem-loop structures located in the fourth intron of
Firstly, RNA-seq data (small RNA seq reads from ENCODE/CSHL and SRA data) analysis, as well as RT-qPCR detection followed by sequencing of the amplicons of mature ErbB4-miR1 and ErbB4-miR2 forms, verified the identity of these novel miRNAs. RT-qPCR analysis showed that ErbB4-miR1/2 were downregulated in both tissue samples and in higher grades of breast and colorectal cancer cell lines. These data suggest that ErbB4-miR1/2 are associated with breast and colorectal cancer progression. Since bioinformatics tools introduced ErbB receptors as bona fide and common target genes of ErbB4-miR1/2, precursors of these miRNAs were individually overexpressed in both breast (SKBR3) and colorectal (SW480) cancer cell lines, and their molecular/cellular effects were investigated. ErbB4-miR1/2 overexpression resulted in reduced transcript levels of ErbB2, ErbB3, AKT1, and AKT3 genes in SW480 and SKBR3 cells, as detected by RT-qPCR. The dual luciferase assay verified that ErbB2-3’UTR is directly targeted by both ErbB4-miR1/2, and ErbB3-3’UTR is targeted by ErbB4-miR2. Finally, western blotting against phospho-AKT and total-AKT levels confirmed the inhibitory effect of ErbB4-miR1/2 on the ErbB/PI3K signaling pathway.
Multiple studies have supported the role of ErbB/PI3K in cell cycle and proliferation regulation by modulating the downstream genes involved in these pathways [37]. Consistently, when ErbB4-miR1/2 were overexpressed in SKBR3 and SW480 cells, the results indicated that these miRNAs inhibited cell proliferation and viability, as evidenced by PI flow cytometry, colony formation, MTT assays, and RT-qPCR. Therefore, the growth-inhibitory effect of ErbB4-miR1/2 could be attributed to the suppression of ErbB/PI3K signaling components.
Because an increase in intracellular ROS can induce cancer cell cycle arrest [38], ErbB4-miR1/2 overexpression effect on ROS levels was investigated. The results of their overexpression indicated that these miRNAs increased ROS levels in the cells, along with their effect against ErbB/PI3K signaling and inhibition of cell proliferation, resulting in cell cycle arrest through ROS production. Taken together, these data suggest the inhibitory effect of ErbB4-miR1/2 on breast and colorectal cancer cell viability and proliferation.
The PI3K signaling pathway can affect tumor aggression by controlling the EMT process [39, 40], and is considered to be a possible target for tumor prevention and treatment. In this study, the scratch and transwell assays showed that ErbB4-miR1/2 overexpression in both SKBR3 and SW480 cells resulted in decreased cell migration. In addition, altered expression of EMT-related genes was observed following ErbB4-miR1/2 overexpression. These data indicated that ErbB4-miR1/2 overexpression retarded cell cycle progression and inhibited migration in breast and colon cancer cells, suggesting that these miRNAs may also function as metastasis suppressors by blocking these pathways.
Collectively, the present study demonstrated that ErbB4-miR1/2 are novel miRNAs encoded by the

Schematic representation of ErbB/PI3K regulation by ErbB4-miR1/2. The present study demonstrated that ErbB4-miR1/2, are novel miRNAs encoded from the
Footnotes
Acknowledgments
This work was supported by Tarbiat Modares University. We thank 4402 Lab members, Motamed Cancer Institute (MCI), and Imam Khomeini Hospital for their cooperation.
Author contributions
CONCEPTION: ZG, BMS and SJM
DATA CURATION: ZG
ANALYSIS OF DATA: ZG
PREPARATION OF THE MANUSCRIPT: ZG and BMS
SUPERVISION: BMS
Conflict of interest
The authors declare no conflicts of interest.
