Abstract
BACKGROUND:
Type 2 diabetes mellitus (T2DM) is a complex disease with high incidence and serious harm associated with polygenic determination. This study aimed to develop a predictive model so as to assess the risk of T2DM and apply it to health care and disease prevention in northern China.
OBJECTIVE:
Based on genotyping results, a risk warning model for type 2 diabetes was established.
METHODS:
Blood samples of 1042 patients with T2DM in northern China were collected. Multiplex polymerase chain reaction and high-throughput sequencing (NGS) techniques were used to design the amplification-based targeted sequencing panel to sequence the 21 T2DM susceptibility genes.
RESULT:
The related key gene KQT-like subfamily member 1 played an important role in the T2DM risk model, and single-nucleotide polymorphism rs2237892 was highly significant, with a
CONCLUSIONS:
Susceptibility genes in different populations were examined, and a model was developed to assess the risk-based genetic analysis. The performance of the model reached 92.8%.
Introduction
The prevalence rate of diabetes in China has increased rapidly, and the increase in medical expenses has brought a heavy burden to families and society in the last two decades. At present, the prevalence rate of diabetes in China is 9.1%. China has more than 100 million patients and 400 million potential patients [1]. Hence, diabetes has become a prominent public health and social problem. Type 2 diabetes mellitus (T2DM) occurs mainly in adults and is the main type of diabetes, accounting for more than 95% of the total patients with diabetes. Recent genome-wide association studies (GWAS) and improved single-nucleotide polymorphism (SNP) analyses have revealed hundreds of common genetic mutations closely related to T2DM. However, the relationship between gene polymorphism and the occurrence of T2DM, as well as its underlying mechanism, is still unclear [2, 3, 4, 5]. Consequently, the etiology and pathogenesis of T2DM have not been fully elucidated. Further, differences exist in the genetic background, living environment, and behavioral patterns. The T2DM susceptibility gene mapping of different regions and nationalities is different. Also, no accurate detection and evaluation model is available to screen populations at high risk of diabetes. Therefore, the occurrence of T2DM and its correlation with the SNP of the T2DM susceptibility gene have become a hot spot in recent years [6, 7, 8].
In 2007, a study entitled “risk assessment method of diabetes in adults in China” published in the Chinese Journal of Health Management proposed the first DM risk assessment model in China [9]. The model was not based on population data, but was a composite model estimated by literature and expert experience. However, it was an effective method in the absence of prospective cohort studies in China. Unfortunately, the model had no data validation results. In 2009, Chien [10] established a prediction model for DM in the Taiwan community population. This model was the first individual risk score model of DM in a Chinese population based on Framingham cardiovascular prediction model. The cohort data of people aged more than 35 years were tracked for 10 years using the Cox proportional hazards model. The risk assessment score system was established using the Framingham risk score equation published by Sullivan [11] in 2004. The indicators included age, fasting blood glucose level, BMI, TG, i-idl-c, and blood leukocyte count. After the establishment of the model, the statistical methods of net reclamation improvement and integrated discrimination improvement were used for training. The AUC reached 0.702, which was better than that of the classic models such as San Antonio, Framingham, PROCAM, and Cambridge. To a large extent, the model covered the risk assessment of DM in China. However, the application of the model was limited due to the close relationship between leukocyte count and infection.
This study was performed to develop a predictive model so as to assess the risk of T2DM and apply it to health care and disease prevention in northern China.
Materials and method
Screening of susceptibility genes
Data resources from the following databases were used: OMIM online [12], GWAS Catalog [13], GeneCards [14], and HGMD [15]. More than 350 susceptibility genes related to T2DM were statistically analyzed using the aforementioned databases. Also, the distribution of different populations was different. The susceptibility genes reported in European populations were TCF7L2, CDKAL1, SLC30A8, FTO, CDKN2B, and so forth, while the susceptibility gene reported in East Asian populations was KQT-like subfamily member 1 (KCNQ1) [16, 17, 18, 19, 20, 21, 22, 23]. The frequency of the report is shown in Fig. 1. Furthermore, 21 of the 16 susceptibility genes were selected in combination with the reported frequency of the susceptibility genes or loci on the basis of the analysis of population differences, IPA gene, and mind pathway, as shown in Table 1.
Screening susceptibility genes and locus as well as the function of genes
Screening susceptibility genes and locus as well as the function of genes
Reported frequency of susceptibility genes among Asians (left) and Europeans (right).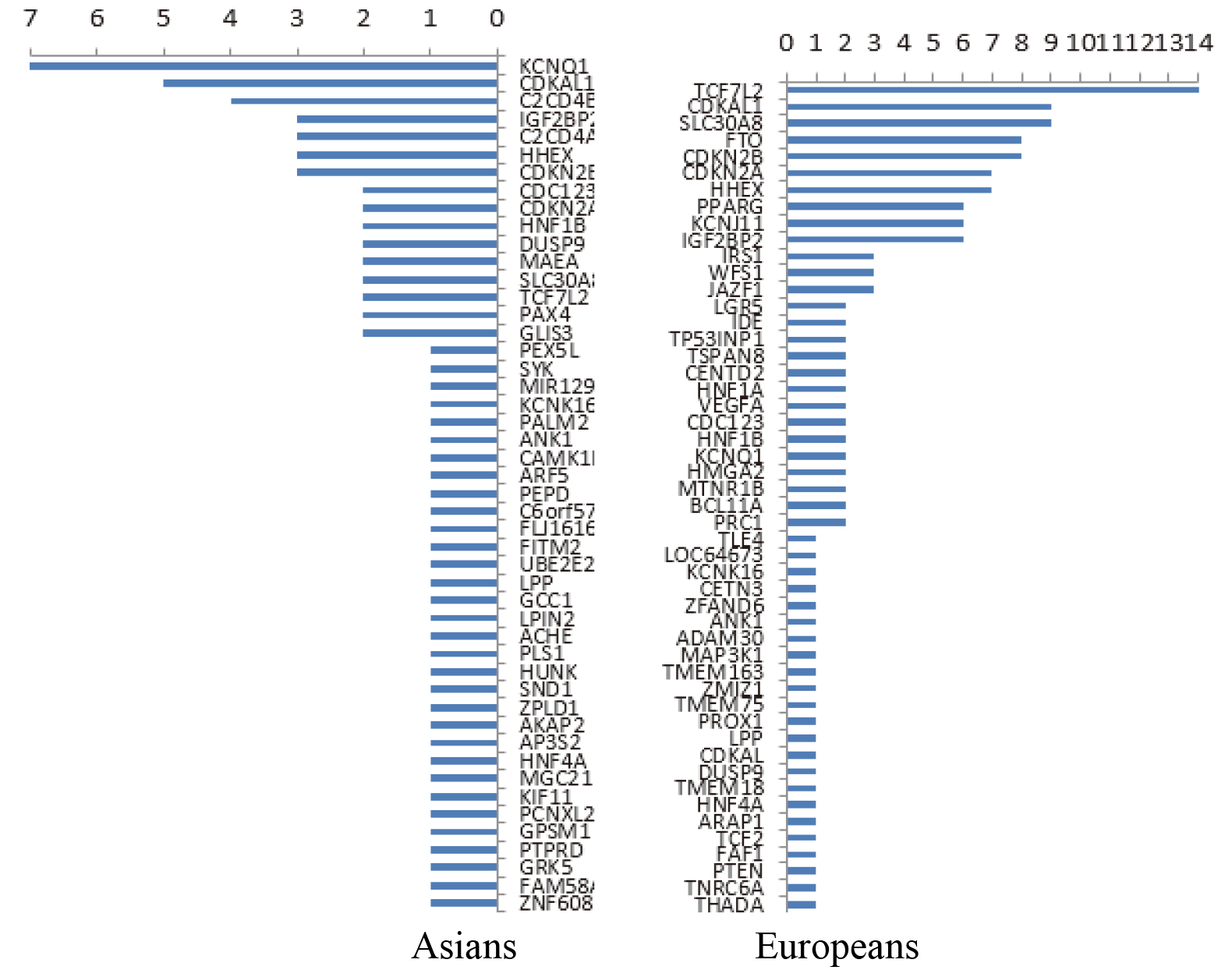
In 2014 and 2015, 1042 peripheral blood DNA samples of patients diagnosed with T2DM in Hebei Yiling Hospital were selected, which met the 1999 World Health Organization diagnostic criteria for diabetes. This study was approved by the ethics committee of each hospital, and all participants signed informed consent. Each individual sample should be no less than 5 mL to meet the requirements of subsequent DNA extraction and gene mutation identification. Vacuum containers with EDTA anticoagulant were collected. The samples are stored at
Results and analysis
Statistical analysis of sequencing results
High-quality sequencing data were obtained through second-generation sequencing. The software analyzed the mutation frequency of 21 SNP sites in the samples of 1042 patients, as shown in Table 2, in accordance to a previous study [24]. A comparison of the genotypes of the SNP sites showed that the distribution frequency of the genotypes was as follows: PPARG rs1801282, IGF2BP2 rs4402960, HHEX rs1111875, HNF1 rs4430796, and WFS1 rs10010131 with no significant difference (T2D
Statistics of mutation frequency of 21 SNP loci
Statistics of mutation frequency of 21 SNP loci
The logistic regression model was used to predict the risk of T2DM. The criteria for T2DM were used in the construction of the analytical model. The classification variance was assumed as
A total of 1563 people were statistically analyzed based on the susceptibility typing results of 21 loci. After constructing the data set, the logistic regression analysis was used to establish the classification prediction model.
In the aforementioned model,
In practical application, the overall evaluation could be judged based on the calculation. When
Refinement evaluation: A
Discussion and conclusion
Genetic testing was performed on 1042 patients with diabetes. A total of 1028 confirmed cases of T2DM were detected through model verification. The detection accuracy was 97.5%. Further, 512 Asian health population data were acquired from the 1000G project public database. Based on the analysis results, the aforementioned formula was used for calculation. Of these, 451 individuals had no T2DM. The detection accuracy was 88.1%. The overall detection accuracy was 92.8%.
Previous studies found that early interventions (diet, exercise, medications, and so forth) could slow or even reverse the T2D development because the early onset of T2D was mild, and the current diagnostic criteria failed to detect and diagnose diabetes in most patients as early as possible. This not only delayed prevention and treatment but even worsened the disease. The pathogenesis of T2D is related to not only environmental factors such as high sugar intake and lack of exercise but also genetic factors. It is a complex genetic disease caused by multiple genetic mutations. Many SNPs related to the pathogenesis of T2D have been found with the development of GWAS and meta-analyses. The genes for these loci are located in the cells of the pancreas. They affect cell function by acting on different physiological and pathological processes. Screening for these genetic variants to assess the risk of diabetes is a hot topic in genetic diagnosis. Sequencing and genotyping statistical analysis of genomic DNA extraction in patients with type 2 diabetes with normal blood control group Blood examination and clinical biochemical examination specimen collection to screen susceptible gene SNP site design primers [25, 26, 27, 28].
This study further detected the T2D-related loci in northern China so as to establish the early genetic screening model of Chinese people. A total of 1563 people participated in the study. The T2D case group (Han population in northern China) and the normal control group (1000 Genomics) comprised 1042 and 512 cases. A risk warning model was established for T2DM. The risks of rs10425678, rs10811661, rs10886471, rs1111875, rs12243326, rs13266634, rs17584499, rs1800629, rs1800795, rs1801282, and T2D were closely related. Also, rs2237892, rs2237895, rs2237897, rs4402960, rs5219, rs7172432, rs7403531, rs7903146, rs8050136, rs864745, and rs9939609 were included. First, this study verified that the SNP locus rs2237892 of the KCNQ1 gene was at the start of the T2D risk gene in the T2D population of Han nationality in China. The logistic regression analysis model results were more accurate compared with other models. Moreover, a software tool was developed for diabetes risk assessment in northern China, thus laying the foundation for the development of subsequent gene detection products.
However, a limitation of this study was the insufficient sample size or different frequencies of different susceptible genes in different populations [29, 30, 31, 32]. Therefore, further verification and improvement are needed in this regard. In this study, the SNP locus rs5219 of KCNJ11 in the Chinese population was the same as that reported in the European population, and therefore it was believed that this locus has not been repeated in more samples.
Footnotes
Acknowledgments
This study was financially supported by the Beijing Science and Technology Project (No. Z181100001918015) and the Beijing Municipal Financial Project (No. PXM2019_178305_000019).
Conflict of interest
None to report.
