Abstract
Darwin-SW (DSW) is an RDF vocabulary designed to complement the Biodiversity Information Standards (TDWG) Darwin Core Standard. DSW is based on a model derived from a community discussion about the relationships among the main Darwin Core classes. DSW creates a new class to accommodate an important aspect of its model that is not currently part of Darwin Core: a class of Tokens, which are forms of evidence. DSW uses Web Ontology Language (OWL) to make assertions about the classes in its model and to define object properties that are used to link instances of those classes. A goal in the creation of DSW was to facilitate consistent markup of biodiversity data so that RDF graphs created by different providers could be easily merged. Accordingly, DSW provides a mechanism for testing whether its terms are being used in a manner consistent with its model. Two transitive object properties enable the creation of simple SPARQL queries that can be used to discover new information about linked resources whose metadata are generated by different providers. The Organism class enables semantic linking of biodiversity resources to vocabularies outside of TDWG that deal with observations and ecological phenomena.
Introduction
Rationale
Biodiversity data, records of the occurrences of organisms in their environment, have traditionally been recorded along with collected specimens preserved in natural history collections. Many specimen-based data have now been exposed electronically by a large number of providers. These data have been aggregated on a large scale and are augmented by observations data and data collected by newer mechanisms such as remote sensing and digital imaging. Expressing these data in the Resource Description Framework (RDF)1 See
Darwin Core [10] is a general-purpose vocabulary designed to facilitate the transfer and integration of biodiversity data. It was developed from 1998 through 2009, when it was ratified as a TDWG technical specification standard.11
Because of its widespread use, and since its terms are defined using RDF, there has been considerable interest in using the Darwin Core vocabulary to describe biodiversity resources in RDF. An RDF guide13 Current version at
In response to needs in our Bioimages15
The development of Darwin-SW was influenced by previous work in modelling the biodiversity domain. An early attempt to develop a standard model was the Association of Systematics Collections (ASC) Information Model for Biological Collections,17
In 2006, the Taxonomic Databases Working Group (TDWG; now Biodiversity Information Standards) began development of a high level technical architecture for information transfer. A central part of this architecture was the creation of an ontology to facilitate the integration of standards.19
The TaxonConcept ontology22
Following the ratification of Darwin Core as a standard, there was a lengthy discussion on the TDWG email discussion list24
Because of our goals to enable the translation of existing Darwin Core-based records into RDF so that they could be easily merged with records from other providers, and to achieve this by using as many existing Darwin Core terms as possible, Darwin-SW was designed essentially to establish a standard mapping of a consensus relational model to RDF. We conducted an extensive review and summary27
In an effort that paralleled the development of DSW, the definitions of several Darwin Core classes were changed officially to make them consistent with the consensus reached during the email discussion.28 http://rs.tdwg.org/dwc/terms/history/decisions/#Decision-2014-10-26_15.
In this paper, class names are capitalized when referenced in English text, e.g. Organism. In diagrams and tables, URIs are abbreviated using standard QNames for namespace prefixes, e.g.
Section 2 of this paper describes the model, design considerations, and features of Darwin-SW. Sections 3 and 4 provide examples showing how DSW facilitates simple forms of reasoning and integration across biodiversity domains. Section 5 describes how DSW may evolve with Darwin Core in the future.

Entity-relationship diagram of the Darwin-SW model using crow’s foot notation with classes and relationships described in English text.
Design considerations
There has been longstanding interest in the TDWG community in leveraging Linked Data and Semantic Web technologies. The barrier caused by the lack of a consensus set of predicates for expressing biodiversity data as RDF would be lowered most easily if most of those predicates originated from terms already in common use. Because of the widespread acceptance of the Darwin Core standard, the choice was made to base Darwin-SW primarily on existing Darwin Core classes and the properties organized under them. The “lingua franca” status of Darwin Core makes integration of triples from multiple institutions more feasible.
Instances of key classes within the biodiversity domain (such as organisms, specimens, and taxa) within or among institutions can be linked using DSW-defined object properties that have no analogues in Darwin Core. Those instances can serve as anchor points to which resources outside the biodiversity community may be linked. Because of this cross-institutional and cross-community anchoring function, DSW expects that most resources will be identified by persistent HTTP URIs or HTTP-proxied versions of globally unique identifiers in accordance with the TDWG GUID Applicability Statement Standard.
Since the primary objective of DSW is to facilitate the linking of real data, and also since some Darwin Core classes have been applied with a wide variety of meaning, DSW takes a practical approach to the definition of classes. In many cases, DSW sees class instances as nodes that group related properties rather than as entities that are heavily constrained ontologically (see Sections 2.2.2 through 2.2.4 for specific examples).
This approach differs significantly from that taken in the development of more formal ontologies with extensive class hierarchies, OWL restrictions, and other features that introduce significant entailments. Although DSW uses properties from OWL in its definitions, these properties were chosen very selectively to generate a limited number of entailments that facilitate a few simple but useful reasoning tasks which can be performed using SPARQL queries (see Section 3 for examples).
Classes of the Darwin-SW model
Relationships among classes
Many existing biodiversity data are managed as part of relational databases where tables contain data related to particular Darwin Core classes. Figure 1 is an entity-relationship diagram that shows how those tables would be related in a standard relational database under the Darwin-SW model. This diagram shows the most normalized database structure likely to be used by a provider that uses Darwin Core as the basis for its records. The table relationships shown in the diagram using “crow’s feet” notation [7] do not represent cardinality restrictions, but rather the relationship types that are likely to exist between instances of the classes. For example, many Events may be associated with a Location, while the reverse is not likely to be true. The structure of Fig. 1 was influenced30
If a provider with such a relational database chose to expose its data as RDF, each record in a table would be represented as a node in the RDF graph. The cardinality of relationships shown in Fig. 1 indicates the way we would expect arcs to link nodes representing instances of the linked classes. For example, the one-to-many relationship between the Location and Event classes indicates that we expect that a node representing a Location instance could be linked to many Event nodes, and not the reverse. Darwin-SW does not enforce cardinality through object property restrictions; Fig. 1 simply indicates the cardinality expected based on data from typical database implementations.
Section 2.2.2. through 2.2.4. provide additional details about key classes in the model.
A key addition of the DSW model was the inclusion of an Organism class. An instance of the Organism class is not restricted to a single biological organism: it can be any sort of organism, clone, colony, or group of organisms that is typically observed or sampled over time. An additional requirement is that Organism instances should be taxonomically homogeneous because they intended be the objects of taxonomic determinations (i.e. Identification instances). A third feature of an Organism is that it serves as an anchor point for resources derived from it, such as specimens, images, and samples. So although an Organism can be described in conceptual terms by comparison with biological organisms, from the standpoint of the DSW model, an Organism is a node that connects Occurrences, Identifications, and derived resources [1].
The existence of this class was implied by the existing Darwin Core term
The Darwin Core Occurrence class (
Taxa are an important component of any biodiversity model and significant effort has been expended towards defining the meaning of the terms “taxon” and “taxon concept” [5]. The TDWG Taxon Concept Transfer Schema (TSC) standard33 The corresponding Darwin-SW-minted term
Object properties that link the main classes in the Darwin-SW model
Notes: Namespace prefixes are defined in Table 3 of the Appendix. Each property has an inverse property linked by an
Evidence-related object properties in the Darwin-SW model
Notes: Each property has an inverse property linked by an
Darwin-SW defines a number of object properties used to link classes in the DSW model (Tables 1 and 2). In most cases, these properties occur in pairs that are related by an
Table 1 shows the object properties defined by DSW to link its main classes (Location, Event, Occurrence, Organism, Identification, and Taxon Concept). Because the primary objective of DSW is to facilitate the linking of real data, these object properties serve primarily as a means to facilitate one-to-many or many-to-many relationships among instances of the main classes as described in Section 2.2.1.
Properties linking to agents
In addition to the object properties intended to link its main classes, DSW also makes use of the object properties

Properties used to link evidence to key classes in the Darwin-SW model. Bold arrows are transitive
A key innovation of Darwin-SW is the recognition of the role of evidence in documenting the occurrences and identifications of organisms. DSW recognizes organism-related evidence by explicitly defining the class
Tokens are linked to the Organism from which they originated by the transitive object property
DSW defines two pairs of object properties that can be used to indicate the evidence on which an assertion about an occurrence is based (Table 2; Fig. 2).
Querying and reasoning facilitated by DSW
Denormalization and inconsistent term use
A design goal of Darwin-SW was to facilitate biodiversity data integration by making it feasible to merge graphs containing RDF data exposed by different providers. The combined graph could then be queried to discover information that would not be apparent through examination of the separate graphs.
One possible impediment to this kind of cross-institutional data integration is inconsistent use of object properties. In this section, the term “normalize” is used in the relational database sense. Depending on the type of resource of interest to particular data providers, they may structure their non-RDF databases using different levels of normalization. When the data in those databases are exposed as RDF, providers may not include instances of classes that are included in the Darwin-SW model.
For example, providers that are interested in recording many Occurrences at an Event and many Events at a Location will create Event instances (Fig. 1). Using Darwin-SW, they can then link Occurrences to Events using
Other providers that are not interested in linking many Occurrences to a single Event may have denormalized their model to eliminate Event. Such providers may inappropriately link Occurrences directly to Locations using
Although in the spirit of the semantic web providers may link the resources they describe in any manner, using Darwin-SW object properties to link class instances in a manner that is inconsistent with Fig. 1 and Table 1 is counterproductive to the design goal of enabling data integration and effective querying. A simple SPARQL query based on a graph pattern that assumed Occurrences were linked to Events and that Events were linked to Locations would fail to bind Occurrences that were linked directly to Locations (Appendix, Query 1). Thus inconsistent use of DSW object properties caused by denormalizing the DSW model inhibits effective querying.
It would be possible to construct a more complex SPARQL query to accommodate alternate degrees of normalization. However, because there are six main classes in DSW, there would be many possible ways that denormalization could occur and it would be difficult to account for all possible permutations using any reasonable query. For this reason, DSW assumes the most normalized model that is likely to be of interest to the biodiversity informatics community. If necessary, providers can create blank nodes representing class instances that are not explicitly present in their database in order to use DSW object properties consistently with the DSW model (Appendix, Example 3).
Detecting inconsistencies using ranges, domains, and disjoint classes in Darwin-SW
Because RDF allows anyone to say anything about anything, there is no simple way to enforce appropriate linking using DSW object properties. However, the properties of DSW terms make it possible to detect inconsistent links of the sort discussed in Section 3.1.
Darwin-SW declares domains and ranges for most object properties that it defines (Tables 1 and 2). A client can infer types that are entailed by these domain and range declarations. Because DSW declares all of its main classes to be mutually disjoint, a client can detect inconsistent DSW object property use when an inferred type of a resource conflicts with a disjoint type that is explicitly declared for that same resource. An inconsistency can also be generated simply by using two DSW object properties in a manner inconsistent with the DSW model. For example, asserting:
generates an inconsistency without any explicit type declarations since the range of
Using this approach, graphs containing data from new providers could be screened for inappropriate use of object properties before they are merged permanently with an existing multi-institution graph. An OWL reasoner could detect the change of the multi-institution graph from consistent to inconsistent upon the addition of the incoming graph if the incoming graph contained DSW object properties used in a manner that conflicted with the DSW model. However, the limited reasoning that is required to screen for inappropriate linking can be done using several simple SPARQL queries (see Queries 2 and 3 in the Appendix). These queries are not computationally intensive and only one needs to be run on the entire merged dataset.
Cross-institutional discovery
The ability to discover previously unknown information that is entailed by asserted triples is one of the most compelling reasons for expressing data using RDF vocabularies instead of more traditional database methods.
The semantics imposed by Darwin-SW term definitions may entail some simple information not explicitly expressed. For example, an asserted triple:
entails
because
Although it may be useful to generate unstated triples of this sort, that hardly constitutes “discovery” of novel information. However, DSW enables the discovery of truly novel information by making it possible to link in a simple and consistent manner data about resources originating from different institutions, conducting limited inferencing, and then querying the graph that is a merge of the inferred and asserted triples. In particular, use of the transitive object property
Linking duplicates
It is common practice in the collections community to collect multiple specimens from the same organism, colony, or local population (a taxonomically homogeneous entity and therefore a
Unfortunately, records of duplicate exchanges are often poor, particularly when the specimens are old. This creates a problem when new information is linked to one duplicate without the knowledge of other institutions holding duplicates.
Despite the colloquial meaning of the word “duplicate”, it would be incorrect to assert equivalence between duplicate specimens since they are distinct entities with different post-collection histories. But because of the nature of the origin of duplicate specimens, it would be correct to state that each duplicate was derived from a single Organism, and it would be appropriate to link each duplicate to the same Organism instance using
For example, if Provider 1 described a specimen like this:
Provider 2 could assert that another specimen was a duplicate using:
If the specimen were collected from the same Organism but as part of a different Occurrence (i.e. at a different time and possibly a different place), Provider 2 could create a different Occurrence instance while still asserting a
In a case where the duplicate status were initially unknown, it would be likely that each institution that databased one of the duplicate specimens would mint a separate URI for the Organism from which its specimen was derived. A later discovery that the specimens were duplicates would imply that the two Organism URIs were simply different identifiers for the same resource.
Two URIs identifying an identical resource can be linked using the predicate
For example, Provider 3 might have described a specimen in its collection like this:
If Provider 3 discovered later that the specimen was a duplicate documenting the same Occurrence as Provider 1’s specimen, it could document that discovery by making the following assertions:
If the specimen were collected from the same Organism, but at a different time or location, Provider 3 could indicate that by asserting only:
Documenting duplicates in this way creates links that facilitate discovery of any new information related to the linked resources. For example, if a new taxonomic determination were made based on Provider 1’s specimen:
Providers 2 and 3 could discover this new information related to their specimens because their specimens were linked to the same Organism instance. The following section describes methods that can be used to make such discoveries.
Organisms and specimens have increasingly become the source of a variety of derived resources obtained through physical and electronic means. Tissue samples, DNA sequences, digital images, telemetry, digital sound recordings, and video may be generated directly from organisms or from resources derived from organisms. As they are generated, these resources may pass to new institutions, and new metadata about the resources may be created without the knowledge of holders of related resources.
The object property
In Fig. 2, the specimen image is linked to its parent specimen by the triple:
and that specimen is linked to its parent Organism (a tree) by the triple:
A reasoner can infer the triple
which links the specimen image directly to its more distant ancestor (the tree) based on the transitivity of
would find all resources derived from the Organism in Fig. 2.
One implication of the transitivity of
would discover new resources derived from the Organism in Fig. 2 that were created after the beginning of 2012.
This is an uncomplicated illustration because it involves a single Organism with few
would discover any new determinations that were made after the start of 2012 that were relevant to any images in the collection identified by the URI
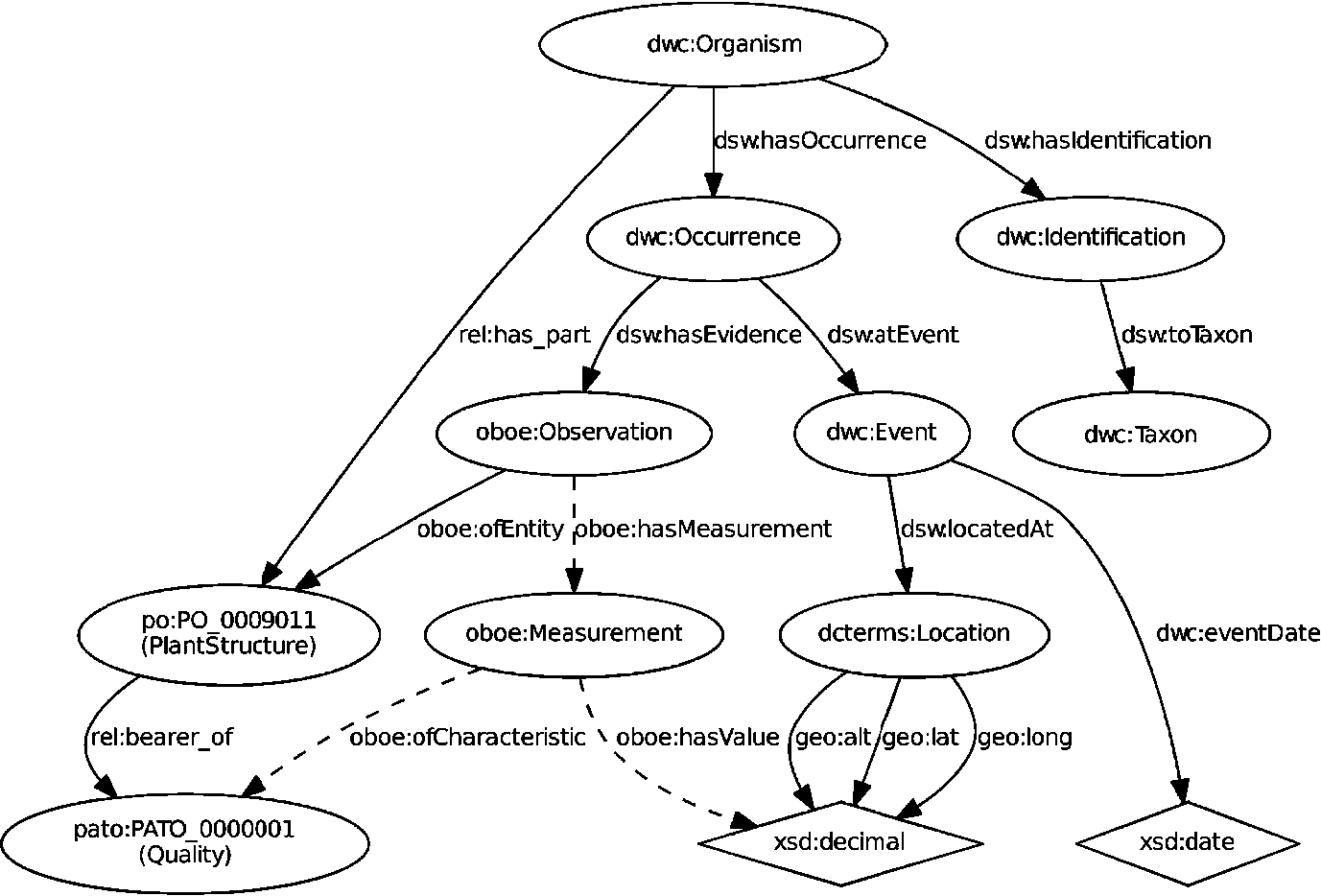
Diagram of a graph structure that can be used to represent observations of organisms that also have identifications. Note that the diagram shows links between
In these two examples, the chain of derived resources was relatively short. However, one can imagine realistic scenarios where the chain of derived resources was much longer. For example, consider a bird that is captured and banded by Institution A. A tissue sample collected during the capture is sent to Institution B which extracts and sequences DNA, with the sequence deposited in the repository of Institution C. At a later time the bird is found dead and given to Institution D which accessions the bird skin and associates the specimen with the earlier collection event based on the information on the band. Institution D loans the skin to Institution E, where an expert makes a determination based on comparison with other skins in the collection at Institution E. If each institution linked its resource to the parent resource from which the institution’s resource was derived using
These examples illustrate how a relatively uncomplicated set of RDF properties combined with simple queries can discover kinds of information that would be very difficult to track using conventional database methods. The challenge therefore becomes more social than technological since the barriers to achieving that kind of discovery require community efforts such as adoption of standard vocabularies, commitment to persistent URI identifiers, reuse of identifiers assigned by others, and establishment of a consensus triple store into which graphs from various institutions would be merged.
Darwin-SW facilitates the linkage of core museum resources (specimens, taxa) to related, ‘external’ RDF resources developed in other knowledge domains (such as genes, locations, publications, agents, media, etc.). In particular, the employment of an Organism class permits simple linkage of taxonomic determination, vouchering information and
As an example, Fig. 3 shows a graph representation of a morphological observation of a named Organism. It employs two additional, existing ontologies: the OBOE observations ontology38
Similarly, because Darwin Core includes a class for Organisms, Darwin-SW facilitates the documentation of interactions among organisms, such as predation, parasitism, and mutualism. Thus it also advances the ability to document ecological interactions using semantic tools.
Because Darwin-SW is based on the Darwin Core standard, it should evolve over time as that standard evolves, with new terms being added or changed as necessary to maintain consistency.
Although changes to Darwin Core may require DSW to change, DSW was designed with stability in mind. The DSW URIs are based on a purl.org namespace so that they will remain independent of any particular server domain. As it becomes necessary to deprecate DSW terms, they will be maintained in the vocabulary with their deprecation noted in the RDF.
Advancing the efforts towards expressing biodiversity data as RDF depends critically on the availability of object properties to link resources described using Darwin Core terms. In this paper we have provided concrete examples of how Darwin-SW object properties can be used to accomplish reasoning tasks in the context of integration of biodiversity instance data. A variety of approaches to creating object properties have been suggested, and achieving a consensus on which approaches are effective requires testing whether those properties can be used to satisfy important use-cases using real data on a realistic scale. Using the examples in this paper, the performance of Darwin-SW in materializing useful entailments can be compared to other approaches. This is an important step in the development of a consensus RDF model for the biodiversity informatics community.
Footnotes
Acknowledgements
The general outline of the Darwin-SW model was suggested in 2010 by Richard Pyle in an email to the tdwg-content email list.![]()
Hilmar Lapp, Mark Schildhauer, and several anonymous reviewers provided very helpful comments on earlier drafts of this paper.
Steve Baskauf’s participation in the Semantics of Biodiversity Symposium at TDWG 2013 was supported by the Research Coordination Network for the Genomic Standards Consortium (RCN4GSC, NSF DBI-0840989) and the Scientific Observations Network (SONet, NSF #0753144, OCI-Interop).
Campbell Webb is grateful for research support by the National Science Foundation (DEB-1020868).
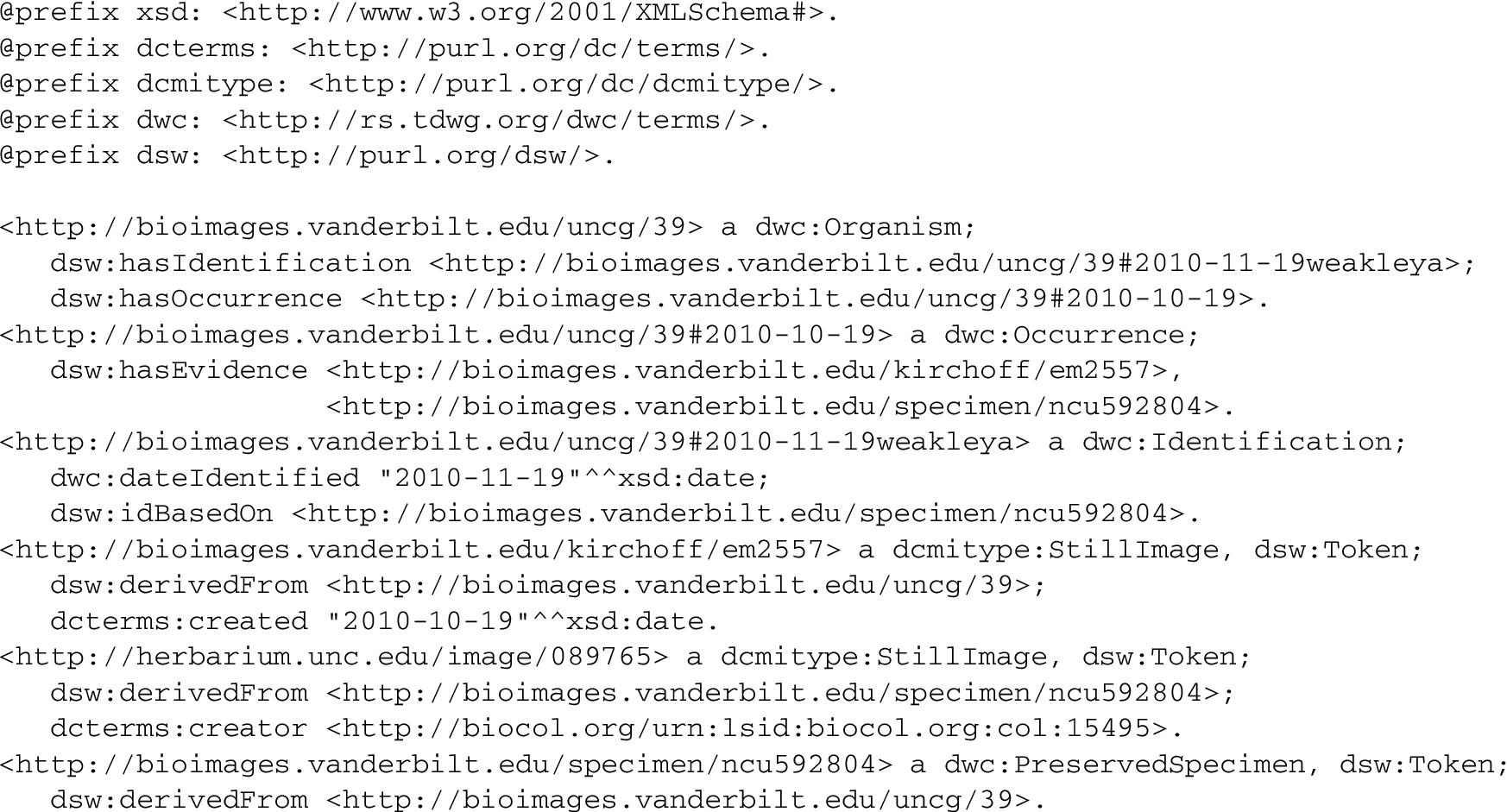
QName namespace prefixes used in this paper Note: Refer to RDF/Turtle serialization based on the graph in Fig. 2.
Vocabulary name
Prefix
URI
Darwin Core (literal values)
Darwin Core (IRI values)
Darwin-SW
Dublin Core
Dublin Core Type Vocabulary
Friend of a Friend
WGS84 Geo Positioning vocabulary
Extensible Ontology for Observations
Phenotypic Quality Ontology
Plant Ontology
Resource Description Framework
RDF Schema
XML Schema
Web Ontology Language