Abstract
The tri-organotin(IV) complex of composition [n-Bu3Sn(C6H5OCH2CONHO)] has been synthesized by using Bu3SnCl and potassium phenoxyacetohydroxamate ligand (KPAH) in pre-determined molar ratio of methanol and tetrahydrofuran. These are characterized by spectral techniques like FTIR, 1 H NMR and mass spectrometry. Studies on electrochemical processes are carried out using cyclic voltammetry. The in vitro antibacterial and antifungal activities performed by using minimum inhibitory concentration (MIC) method has revealed complex as biologically active. Molecular docking of the ligand and complex has been performed against the fungal receptor (PDB ID: 1UKC) to study the interactions between compound and receptor.
Keywords
Introduction
Due to structural complexities, the organotin(IV) compounds have drawn more research interest among various organometallic compounds [1, 2]. In pharmaceutical industries, the synthesis of organotin(IV) compounds as an effective drug with low toxicity and high activity for the medicinal applications has continuously been increasing. The majority of applications of organotin compounds are in catalytic, biological, medicine and material science [3–7].
Hydroxamic acids, R’(CO)N(OH)R” (R’ = alkyl/aryl; R” = H or alkyl/aryl); a class of weak organic acids have much biochemical interest as an active pharmacological agent as they exhibit wide variety of biological activities [8, 9]. Due to metal-binding property, the hydroxamic acids are used by micro-organisms and they substitute for antibacterial, anti-inflammatory, antihypertensive, antitumor, psychotropic and insecticidal [10–12].
Thus, on account of biological importance of both organotin compounds and hydroxamic acids, we report herein the synthesis of tri-butyltin(IV) phenoxyacetohydroxamate [13–16] by the reaction of Bu3SnCl with biologically significant functionally substituted potassium phenoxyacetohydroxamate (KPAH) ligand. The antimicrobial potential against standard bacteria and fungi of potassium phenoxyacetohydroxamate (KPAH) ligand and tri-butyltin(IV) phenoxyacetohydroxamate [n-Bu3Sn(C6H5OCH2CONHO)] complex has been studied thoroughly.
Experimental
Materials and method
Before use, all the reagent and solvents were dried and distilled. The synthesis of potassium phenoxyhydroxamate (KPAH) was carried by reported method [13]. At 18±1°C, the molar conductance of the complex solution (10-3 M) was measured using a CON510; Bench conductivity/ TDS meter.
The FTIR (Fourier Transform Infrared) spectra of ligand and complex were recorded on a Nicolet-5700 spectrophotometer using potassium bromide (KBr) pellets. The 1 H NMR spectra of the complex was recorded on Bruker Avance II 400 spectrometer using DMSO-d6 as solvent and TMS as an internal standard. The mass spectrometer Waters QTOF-MICROMASS is used to record an Electrospray ionization mass spectrum (ESI-MS) of complex. Autolab Potentiostat 128 N electrochemical analyzer was used to perform cyclic voltammetric measurements. A negative scan was initiated in positive direction at a scan rate 100 mVs-1 with the scan in –2 to +2 V range in 10-4 M methanolic solution with 10-2 M KCl as supporting electrolyte. The potentials were measured against reference electrode Ag/AgCl, Pt-working and auxiliary electrodes.
Synthesis of tri-butyltin(IV)phenoxyacetohydroxamate
The solution of n-Bu3SnCl (0.006 mol) was added to potassium phenoxyacetohydroxamate (0.013 mol) solution. The reaction mixture was refluxed for 24 h where formation of a white solid (KCl) was observed. By distillation, the excess of solvent was removed from the filtrate and then by using distilled petroleum ether concentrate was treated. The petroleum ether addition immediately separates the solid cream which was recrystallized after drying appropriately under vacuum (Yield 78%).
Antimicrobial activity assay
The in vitro antibacterial and antifungal activities of potassium phenoxyhydroxamate (KPAH) and tri-butyltin(IV) phenoxyacetohydroxamate complex were evaluated by MIC method against bacteria and fungi using two fold serial dilution and Resazurin as oxidation-reduction colorimetric indicator [17]. For both ligand and complex, the stock solutions were prepared in DMSO (100μg/mL). The standard antibacterial drug: chloramphenicol for S. aureus, S. typhi, E. Coli and B. cereus and antifungal drug: Nystatin for A. niger and A. alternate were used for comparison of results [18]. These studies were performed in triplicate.
Molecular modelling study
To find suitable interactions of ligand and complex against crystal structure of fungal receptor A. niger (PDB ID: 1UKC), molecular docking study was performed. For docking, Autodock tool 4.2 was used and chain-A was selected. Gasteiger charges were assigned to the ligand and complex before docking. Most of the docking parameters were kept as default. The distance measurements among the interacting residues were completed on PyMol [19–21].
Results and Discussion
The reaction of tri-n-butyltinchloride (Bu3SnCl) with potassium phenoxyacetohydroxamate in pre-determined molar ratio 1:1 (metal:ligand) in anhydrous methanol and benzene under reflux at 68°C result in tri-butyltin(IV) phenoxyacetohydroxamate complex of composition [n-Bu3Sn(C6H5OCH2CONHO)] shown by the equation (Fig. 1).
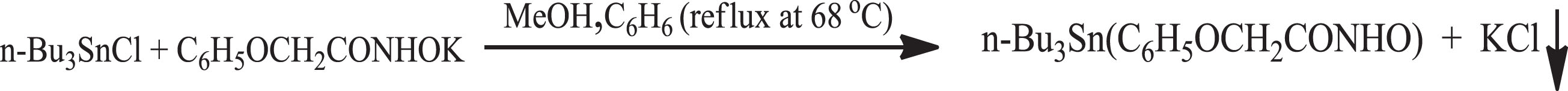
Synthesis of tri-butyltin(IV)phenoxyacetohydroxamate complex.
This complex is a cream colored solid with sharp melting point and easily soluble in nearly all usual organic solvents. The recorded molar conductance is 1.65 and 1.70 mathchar " 1356mScm2mol-1 in 1–3 M methanol for ligand and complex respectively (Table 1) representing their non-electrolytic nature [22]. The structure of complex [n-Bu3Sn(C6H5OCH2CONHO)] is shown in Fig. 2.
The analytical data of the ligand and complex
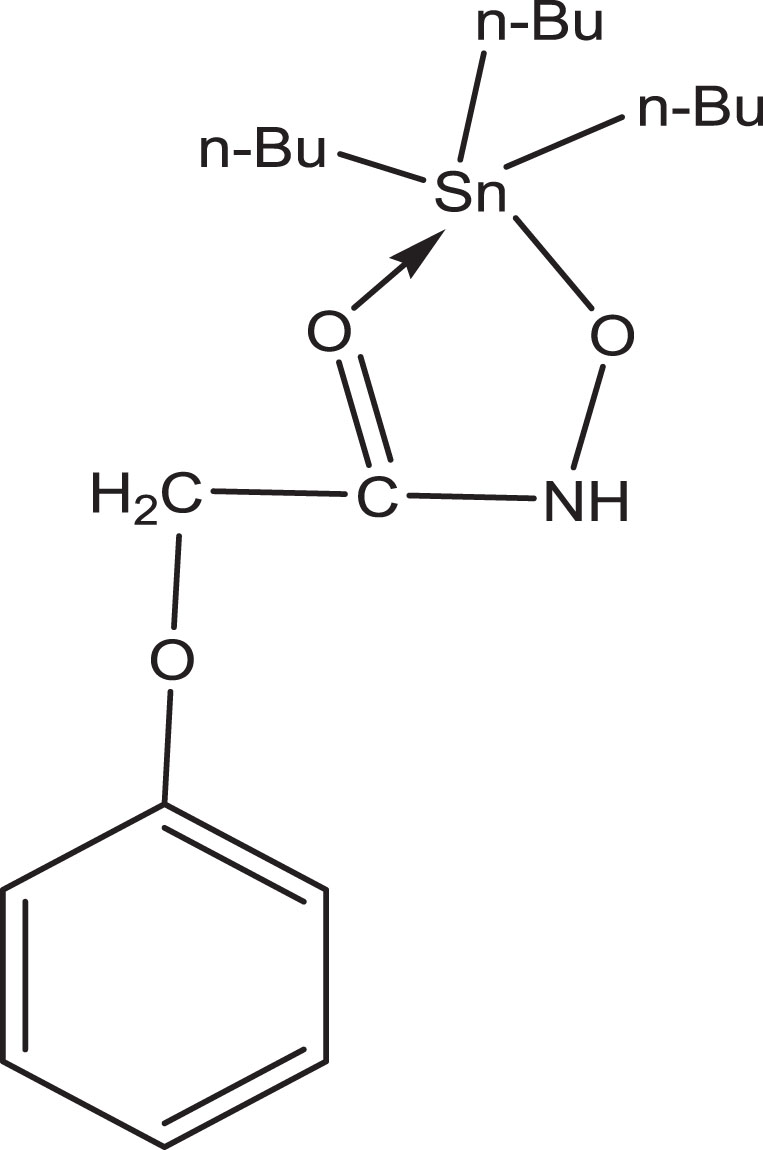
Structure of tri-butyltin(IV) phenoxyacetohydroxamate complex.
The synthesis of complex tri-butyltin(IV) phenoxyacetohydroxamate has been ascertained through carefully assessing its IR spectra with KPAH ligand recorded in 4500–450 cm-1 region (Table 2 and Figures S1, S2). A sharp absorption band at ∼3178 cm-1 in free ligand corresponds to ν(N-H) mode. The bands appearing at 1687 cm-1, 1276 cm-1 and 987 cm-1 correspond to ν(C = O), ν(C-N) and ν(N-O) modes respectively and are in agreement with the reported literature [23]. The absorption band at 3211 cm-1 in [n-Bu3Sn(C6H5OCH2CONHO)] indicates the characteristic ν(N-H) mode. The absorption bands due to ν(C = O), ν(C-N) and ν(N-O) modes have been obtained at 1682 cm-1, 1247 and 998 cm-1 respectively.
IR spectral data for ligand and complex
IR spectral data for ligand and complex
The retention of band due to ν(N-H) modes at 3211 cm-1 in complex has indicated that nitrogen does not participate in bonding. The ν(C = O) absorption band has been shifted to lower frequency in tri-butyltin(IV) phenoxyacetohydroxamate complex. The shift observed in various modes such as ν(C = O), ν(C-N) and ν(N-O) has shown bonding by carbonyl and hydroxylamine oxygen atoms [O,O coordination] which signify that ligand is of bi-dentate nature. The complex formation and bonding by carbonyl oxygen and hydroxylamine group has further been assured by new band appeared at 435 cm-1 which is assigned to ν(Sn-O) mode [13, 24].
The formation of tri-organotin(IV) phenoxyacetohydroxamate has further been validated by comparison of its room temperature 1H NMR with free ligand [KPAH] (Table 3 and Figures S3, S4). The ligand [KPAH] show signals due to –NH proton, aromatic protons and –CH2 at chemical shift (δ) values 10.69–10.95 ppm, 6.91–7.59 ppm and 3.37–3.75 ppm respectively. The complex of composition [n-Bu3Sn(C6H5OCH2CONHO)] exhibits –NH, aromatic proton and -CH2 proton signals respectively at 11.89 ppm, 7.06–7.67 ppm and at 3.46–4.35 ppm. The aromatic protons has shown downfield shift in complex as compared to ligand. The N-H proton resonance in ligand as well as in complex has shown that nitrogen does not participate in coordination. The DMSO-d6 exhibits a sharp signal in 2.50–2.51 ppm range for both ligand and complex. The signal appeared in 1.13–1.35 ppm range due to n-butyl group in complex which is absent in free ligand has further substantiated the complex formation. The resonance observed in 1H NMR spectra of the complex substantiated their formation.
The chemical shift δ (in ppm) for ligand and complex obtained from 1H NMR
The chemical shift δ (in ppm) for ligand and complex obtained from 1H NMR
The ESI mass spectrum of newly synthesized [n-Bu3Sn(C6H5OCH2CONHO)] complex has no molecular ion peak but it has shown di-tin fragment ions at m/z (%) 955 (2), 812 (12), 764 (10) and 634 (6.5) corresponding to [n-Bu6Sn2(C6H5OCH2CONHO)2 + K +]+, [n-Bu6Sn2(C6H5OCH2CONHO) + K+ + Na+]+, [n-Bu3Sn2(C6H5OCH2CONHO)2 + Na+]+ and [n-Bu4Sn2(C6H5OCH2CONHO) + Na+]+ respectively. The fragment ions at m/z (%) 523 (23), 485 (7) and 263 (21) have been accounted for [M+ + 3Na+]+, [M+ + Na+ + 6H+]+ and [LK + K+ + Na+ -5H+] respectively. The fragment ion corresponding to [C6H5OCH2CON]+ has been appeared as a base peak at m/z (%) 150 (100). (Tables 4 and Figure S5).
Mass spectral data of complex
Mass spectral data of complex
The cyclic voltammetric measurements for the ligand [KPAH] and respective tri-organotin(IV) hydroxamate of composition [n-Bu3Sn(C6H5OCH2CONHO)] has been carried out. The voltammetric scans are shown in (Figures S6, S7). The ligand [KPAH] has shown reduction wave at +0.643 V and oxidation wave at –0.514 V. The complex of composition [n-Bu3Sn(C6H5OCH2CONHO)] has shown reduction wave at +0.625 V and one oxidation wave at –0.576 V. The observed voltammograms are suggestive of electrochemically reversible reduction reaction which is herein seemed to be metal centered.
Antimicrobial activity
The biological potential of tri-organotin(IV) compounds have been affected by the hydroxamate ligand’s nature and the number of organic groups coupled with tin [25]. The organotin(IV) complexes have a large number of biological applications with different ligands [26–29].
Antibacterial activity
The inhibitory effect of ligand [KPAH] against B. cereus and S. aureus has been observed at MIC 19.38 and 18.70μg/ml whereas for E. coli and S. typhi at 18.70 and 4.60μg/ml MIC respectively. The complex of composition [n-Bu3Sn(C6H5OCH2CONHO)] has shown MIC at 8.81, 8.81, 14.86 and 3.35μg/ml against B. cereus, S. aureus, E. coli and S. typhi respectively. The observed MIC values indicate that complex exhibit promising antibacterial activity against all tested bacteria which is even better than the free ligand [KPAH]. The commercial antibiotic Chloramphenicol has been chosen as a standard for comparing the results (Table 5 and Fig. 3).
Antibacterial and Antifungal activity data for ligand, complex and standard antibiotic using MIC (in μg/mL) method
Antibacterial and Antifungal activity data for ligand, complex and standard antibiotic using MIC (in μg/mL) method
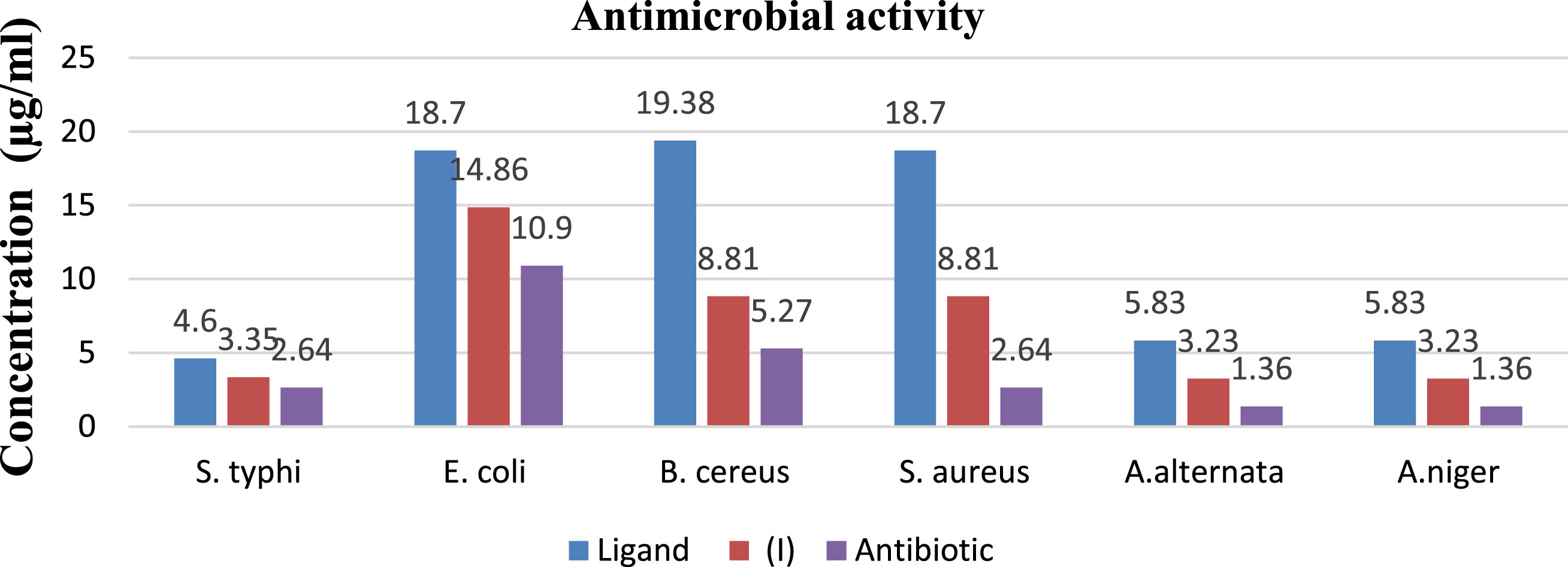
In vitro antibacterial, antifungal spectra of ligand [KPAH], tri-butyltin(IV) phenoxyacetohydroxamate complex (I) and antibiotic.
The inhibition effect against A. alternata and A. niger of the ligand [KPAH] has been observed respectively at MIC 5.83μg/ml and 5.83μg/ml. The complex of composition [n-Bu3Sn(C6H5OCH2CONHO)] has shown inhibitory activity at 3.23μg/ml against both A. alternata and A. niger. The Complex has shown increased inhibitory activity compared to the free ligand [KPAH] against both fungi. The chelation theory may explain the enhanced antimicrobial activity of the complex with respect to the ligand. Upon complexation, the polarity of central metal ion is partially reduced due to sharing of its positive charge with ligand, favoring complex permeation via lipid cell membrane [30].
Molecular docking study
The interactions between drug molecule and receptor have been predicted by Molecular docking study. The affinity of ligand [KPAH] and tri-butyltin(IV) phenoxyacetohydroxamate complex towards A. niger fungal receptor (PDB ID: 1UKC) has been evaluated by determining their binding energies, inhibition constant as well as bonding interactions. The estimated free binding energy of the ligand and complex has been found to be –2.59 kcal/mol and –3.94 kcal/mol whereas inhibition constant (Ki) values are 1.30 and 12.71 mM respectively (Table 6).
The estimated binding energy and inhibition constant from docking of ligand and complex with A. niger fungal receptor
The estimated binding energy and inhibition constant from docking of ligand and complex with A. niger fungal receptor
The amino acid residues Glu 338, Gly 339, Thr 401 and Asp 397 of A. niger fungal receptor have shown hydrogen bonding interactions with phenoxy -
Distances between the interacting residues of A. niger fungal receptor and Ligand
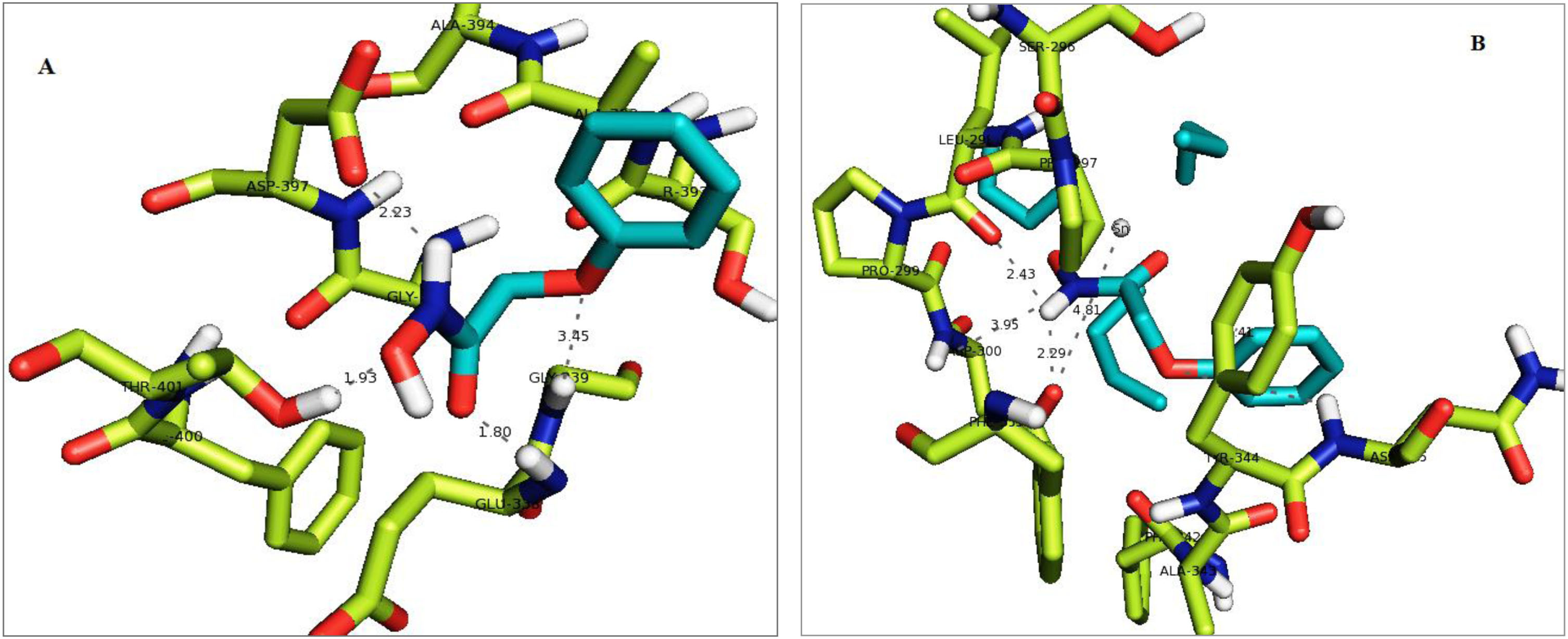
(A) Secondary structure of A. niger fungal receptor (PDB ID: 1UKC) where interacting residues are represented by sticks (green colour) and ligand is shown by cyan colour sticks. The interactions of ligand with amino acid residues are shown by dotted lines. The interacting residues are named with their respective three letters code. (B) Secondary structure of A. niger fungal receptor (PDB ID: 1UKC) where interacting residues are represented by green colour sticks and complex is shown by cyan colour sticks. The interactions of amino acid residues (represented with three letters code) of receptor with complex are shown by dotted lines.
Distances between the interacting residues of A. niger fungal receptor and Complex
Thermal energy (Internal energy) of the molecule can be related with binding energy of the molecule. The molecule with lower internal energy has more binding ability with the receptor. The molecular docking study reveals that complex has lower internal energy as compare to the ligand which indicate that complex has more binding affinity.
The tri-butyltin(IV) phenoxyacetohydroxamate complex has been synthesized by reaction of Bu3SnCl with potassium phenoxyacetohydroxamate [KPAH]. Various spectral techniques like FTIR, 1H NMR and mass spectrometry are used for characterization purpose. The carbonyl and hydroxylamine oxygen atoms [O,O coordination] bonding is specified from IR spectra. From different spectral studies, geometry proposed around tin is five-coordinate distorted trigonal bi-pyramidal. Electrochemical properties studied using cyclic voltammetry suggest metal-centered, two electron quasi-reversible reductions.
The antimicrobial activity of ligand and complex studied using MIC method against bacteria and fungi has revealed complex [n-Bu3Sn(C6H5OCH2CONHO)] as biologically active. Molecular docking of ligand and complex against A. niger fungal receptor has shown more negative binding energy and increased inhibition constant value further establishing more strong binding (interactions) with complex.
Conflict of interest
The authors have declared no conflict of interest.
