Abstract
Background:
Studies have shown different gut microbiomes in patients with Parkinson’s disease (PD) compared to unaffected controls. However, when the gut microbiota shift toward dysbiosis in the PD process remains unclear.
Objective:
We aim to investigate the changes in gut microbiota, locomotor function, and neuropathology longitudinally in PD rodent models.
Methods:
Fecal microbiota were longitudinally assessed by sequencing the V4–V5 region of the 16S ribosomal RNA gene in a human mutant α-synuclein over-expressing mouse model of PD, SNCA p.A53T mice, and the non-transgenic littermate controls. The locomotor function, neuronal integrity, and α-synuclein expression in the different brain regions were compared between groups. Human fecal microbiota communities from 58 patients with PD and 46 unaffected controls were also analyzed using metagenomic sequencing for comparison.
Results:
Compared to non-transgenic littermate controls, the altered gut microbiota of the SNCA p.A53T mice can be detected as early as 2 months old, and the diurnal oscillation of the gut microbiome was dampened throughout PD progression starting from 4 months old. However, neuropathology changes and motor deficits were observed starting at 6 months old. Similar changes in altered gut microbiota were also observed in another PD genetic mouse model carrying the LRRK2 p.G2019S mutation at 2 months old. Among the commonly enriched gut microbiota in both PD genetic mouse models, the abundance of Parabateroides Merdae and Ruminococcus torques were also increased in human PD patients compared to controls.
Conclusion:
These findings revealed the altered gut microbiota communities and oscillations preceding the occurrence of neuropathy and motor dysfunction in the PD process.
INTRODUCTION
Parkinson’s disease (PD) is a common neurodegenerative disorder affecting 7 to 10 million people worldwide, a number expected to double by 2030 [1]. The pathological hallmark of PD, neuronal accumulations of alpha-synuclein named Lewy body, has been identified within the enteric nervous system (ENS) early in the disease process [2, 3]. Studies in rodent PD models have shown that Lewy bodies could propagate from the gut ENS to the brainstem through the vagus nerve [2, 4]. Consistently, gastrointestinal dysfunction, especially constipation, is the earliest prodromal symptom before motor disability by decades [5]. PD patients have an altered gut microbiota composition compared with unaffected controls, and changes in the gut microbiota communities correlate with disease severity and progression in patients with PD [6]. In addition, gut microbiota transplantation from human PD patients could aggravate motor dysfunction in a transgenic PD mouse model [7]. These observations have prompted the hypothesis that gut dysbiosis with an altered gut microenvironment may trigger the disease process of PD.
The gastrointestinal tract is the residence of trillions of microorganisms. The composition of gut microbiota could be modulated by diet, genetic factors, ages, and the circadian clock of the host [8–14]. On the other hand, the composition of gut microbiota could influence the digestion, metabolism, and immune function of the host [15–21]. This intense crosstalk maintains the host-microbial homeostasis. Gut dysbiosis, an imbalance in the composition and function of the intestinal microbiome, leads to dysregulation between host immunity and the gut microbiota resulting in the leaky gut phenomenon [22]. This increased intestinal permeability allows for the leakage of bacteria and their potential metabolites into the systemic circulation and promotes central dopaminergic neurodegeneration through the non-vagal microbiota-gut-brain axis [23]. Recent in vivo studies have shown that changes in gut microbiota could advance the early disease process of neurodegeneration in PD [23] and accelerate the neuropathology and motor dysfunction in a transgenic PD mouse model [7]. However, when the gut microbiota shift toward dysbiosis status remains unclear. In addition, gut bacterial oscillations without changes in microbial abundance have recently been observed to link to an increased susceptibility to neurodegeneration [24]. Gut bacterial rhythms are coordinated by the host circadian rhythms and the external light dark cycle [11, 25]. Circadian rhythm is driven by the suprachiasmatic nucleus (SCN), which is located in the brain’s central core involving the brain-to-gut axis [24]. The circadian clock or light signals could induce rhythmic neurotransmitter secretions, including vasoactive intestinal polypeptide [26], which regulate gut motility leading to gut microbial oscillations and functions [11, 25]. As growing evidence reveals marked circadian rhythm disruption in patients with PD [27–30], it is still unclear whether or when the daily oscillation of the gut microbes is affected in the PD process.
Here we report that the dysbiosis in a PD mouse model carrying the PD-causative SNCA p.A53T mutation [31] can be detected as early as 2 months old, and dampening of daily oscillation of gut microbes was exhibited in 4 months old. In contrast, the classical neuropathology changes and motor deficits of PD are observed starting at 6 months old. Early dysbiosis could also be observed in another PD genetic mouse model, LRRK2 p.G2019S mutation, at 2 months old. Finally, some commonly enriched genus of gut microbes in both PD models matches the changes of specific species of gut bacteria in human PD patients.
MATERIALS AND METHODS
Animal study
Mice
The transgenic SNCA p.A53T mice (B6.Cg-2310039L15RikTg(Prnp-SNCA*A53T)23Mkle/J) [31] and the transgenic LRRK2 p.G2019S (FVB/N-Tg [LRRK2*G2019S]1Cjli/J, JAX009609) [32] were purchased from The Jackson Laboratory (Bar Harbor, ME USA). The heterozygous transgenic mice were then intercrossed to obtain non-transgenic (NTG) littermate control animals. All animals were kept in a light-tight box with air ventilation for at least two weeks before behavioral testing and throughout the experiment. White LED lights were used to provide 800–100 lux of luminance at the home cage bedding level. The light-dark cycle was controlled by a timer for 12 h of light and 12 h of total darkness. Fecal samples were collected after behavioral tests. Autoclaved normal chew food (LabDiet, US, #5010) and filtered non-acidified water were provided ad libitum. To reduce disturbance, cages including food and water were changed once per week during experimental period. The animal studies were approved by the Institutional Animal Care and Use Committee at the Laboratory Animal Center, College of Medicine, National Taiwan University.
Immunofluorescence staining of TH, NeuN, alpha-synuclein and Iba1
Whole brains were dissected from NTG littermate controls and SNCA p.A53T mice at 2 months, 6 months, and 10 months after birth and fixed in 4% paraformaldehyde for 20 h. Before sacrificing the mice, adult mice were first deeply anesthetized with isoflurane (Baxter, Deerfield, IL) administered in a small chamber. Then, mice were perfused with cold 1X phosphate buffer (PBS, pH = 7.4), followed by 10 min of cold 4% paraformaldehyde through intra-cardiac perfusion.
The substantia nigra was cut into free-floating sections 20μm thick (anterior–posterior, –2.30 to –4.16 mm from the bregma) using a cryomicrotome (Leica, Wetzlar, Germany). The sections were rinsed in PBS, incubated for 30 min at room temperature with a blocking solution (1X PBS, BSA 1:50, 0.3% Triton 100), and then incubated overnight at 4°C with a rabbit polyclonal antibody raised against tyrosine hydroxylase (TH, AB152, 1:300 in BSA; Millipore, Dundee, UK), alpha-synuclein (1:200 in BSA, 10842-1-AP, Proteintech, IL, USA) and polyclonal anti-goat against Iba1 (1:200 in BSA, GTX100042, GeneTex) for 16 h at 4°C. Sections were then washed with PBST and incubated with Cy3-conjugated secondary antibodies (donkey anti-rabbit and Cy2-conjugated donkey anti-mouse) for 1 h at room temperature in 1% BSA blocking solution. Three washes were performed, followed by mounting medium with DAPI (GTX30920, GeneTex). In addition, each brain was sectioned serially in the coronal plane using a sliding microtome at 40μm throughout the entire hippocampal formation (from bregma: anterior-posterior –5.6 mm, medial-lateral 4.9 mm, dorsal-ventral –6.8 mm from dorsal dural surface) and the frontal motor cortex (Bregma = 1.72 to 1.48 mm). The sliced sections were incubated with mouse anti-neuron-specific nuclear protein (NeuN; 1:200; Merck Millipore, Temecula, CA) in PBS at 4°C overnight, and incubated with a secondary antibody (Jackson ImmunoResearch) conjugated to Cy3 at 1:200 in PBS. After washing in PBS, sections were counterstained with 4′6-diamidino-2-phenylindole (DAPI) to identify nuclei. A fluorescent microscope Nikon (Thermo Fisher Scientific) was used to capture the fluorescent images of the sections. The total number of TH-immunoreactive neurons in the SNc, or NeuN-positive cells in the hippocampus or motor cortex was quantified using Stereo Investigator 10.0 software (MicroBrightField Inc, Williston, VT) [33]. All stereological analyses were performed under the 100 X objective of a Zeiss Axio microscope (Imager A1, Zeiss).
Motor function assay
Motor function assays, including beam balance test and rotaroad test, were examined every 2 months starting from the age of 2 months old (aged 2, 4, 6, 8 months), and ended at 10 months old before sacrifice for longitudinal analysis.
For the beam balance test, an acrylic glass beam with a flat surface of 3 * 100 cm2 was used to connect the start platform and the home cage. Mice were trained to cross the beam for 3 days before the testing day, and they received 10–15 trails on each training day. On the testing day, the traveling time for the central 80 cm was recorded. Each mouse was recorded for five trails and average time was used for analysis. The beam was cleaned with tissue soaked with 70% ethanol between each trial. All tests were performed at ZT10-12 with a 20-min acclimation period in the experimental room on both training and testing day.
For the rotaroad test, mouse was placed on a 6 cm length and 1.5 cm diameter cylindrical rod, which is spanned by 20 grooves with depth 0.15 mm for the subject to grab. The cylindrical rod rotated along its long axis starting at 5 rpm and gradually accelerated 0.5 rpm every 1.5 s to maximum 40 rpm. The speed of the mouse falling off from the rod was recorded. Results from each individual were calculated by the average of 5 trails. This Rotarod apparatus was constructed by a 3D printer and programmed with Python on the Raspberry Pi. All tests were performed at ZT10-12 with at least 10 min of acclimation to the experimental room.
Fecal sample collection
Fecal samples were collected approximately every 2 months starting from the age of 2 months old (aged 2, 4, 6, 8 months), and ended at 10 months old for longitudinal analysis. For every collection age point, fecal pellets were collected over a 48 h period from each mouse at ZT3, ZT7, ZT11, ZT15, ZT19, ZT23 for JTK analysis. Fecal pellets were flash frozen (–80°C) until DNA extraction.
Metagenomic library preparation and 16S sequencing
The DNA was extracted from around 25 mg of feces, using a QIAamp DNA Mini Kit (Qiagen, Germany) according to the manufacturer’s instructions. And 40 ng of DNA was used in library preparation for 16S rRNA gene sequencing. Amplicons spanning the variable region 4/5 (V4/V5) of 16S rRNA gene were generated using the following primers: 5’-CGT-CGG-CAG-CGT-CAG-ATG-TGT-ATA-AGA-GAC-AGA-YTG-GGY-DTA-AAG-NG-3’ and 5’-GTC-TCG-TGG-GCT-CGG-AGA-TGT-GTA-TAA-GAG-ACA-GCC-GTC-AAT-TYY-TTT-RAG-TTT-3’. The PCR products were cleaned and barcoded using Nextera® Index Kit (Illumina Inc, US). Final products were purified, concentrated and DNA quality verified. The library was sequenced on an Illumina MiSeq platform, with 250 bp paired-end sequencing.
Microbiota sequence analysis
All analysis is following the procedure described in the Microbiome Helper protocol [34]. Reads were processed with QIIME 2 v2022.2 (Quantitative Insight into Microbial Ecology) [35]. In brief, V4/V5 primer sequence were trimmed using the cutadapt QIIME 2 plugin. Paired-end reads were joined and denoised into amplicon sequence variants (ASVs) with the Divisive Amplicon Denoising Algorithm 2 (DADA2) [36]. Taxonomy was assigned using the Silva 138 database (SSU Ref NR 99) [37], and an ASV table was created. For beta diversity analysis, randomly picked 13,000 reads from each sample were used for pairwise distance calculation between all pairs of samples (Bray-Curtis and UniFrac distances). To visualize the dissimilarity matrix, principal coordinate analysis (PCoA) plots were generated using QIIME 2. Shannon index from QIIME 2 were used to compare the species diversity (alpha diversity) of each sample. To determine diurnal fluctuations of each ASV, percentages of ASV reads from the same mouse collected at different time points were used for the JTK_CYCLE analysis [38], a non-parametric statistical algorithm designed to identify and characterize cycling variables in large datasets. In brief, the relative abundance of each ASV from 6 time points was tested for rhythmicity with an assumption period between 22–26 h. ASV with daily oscillation was defined by having both permutation-based adjusted p values and Benjamini-Hochberg q-values < 0.05. Linear discriminant analysis effect size (LEfSe) [39] was used to detect significant difference in relative abundance of microbial taxa between groups via online Galaxy Browser (https://huttenhower.sph.harvard.edu/galaxy/). Significant thresholds were set as default settings: alpha < 0.05 for a factorial Kruskal-Wallis test among classes and LDA > 2.0 for logarithmic LDA score.
Human study
Participants
A total of 58 PD patients and age, gender, and diet habit-matched 46 control participants were recruited from National Taiwan University Hospital. PD was diagnosed according to the United Kingdom PD Society Brain Bank Clinical Diagnostic Criteria [40]. Controls were neurologically unaffected participants who were spouses or accompanying friends of the PD patients. Participants were excluded if they had a history of inflammatory bowel disorders, irritable bowel syndrome, colitis, colon cancer, use of antibiotics or probiotic supplements within 3 months of enrollment; or were vegetarians. A comprehensive dietary history was collected using the Food Frequency Questionnaire [41].
Gut microbiota analyses with shotgun metagenomic sequencing
Fecal samples from participants were collected into stool specimen collection tubes containing DNA stabilizer (Sarstedt), which were immediately flash-frozen on dry ice and stored at –80°C before analyses. Total fecal DNA was extracted using a QIAamp DNA Stool Mini Kit (Qiagen, Hilden, Germany) and gut microbiota was analyzed with shotgun metagenomic sequencing (Illumina NovaSeq 6000) as previously described [42]. Briefly, paired-end reads of the metagenomic datasets from both PD and control samples were trimmed using Trimmomatic v0.38 [43]. To perform the taxonomic and functional profiling of the gut microbial community, MetaPhlAn 3 was run by running HUMAnN 3 [44] integrated with the Bowtie2, Diamond alignment tool, UniRef90 protein database, and the ChocoPhlAn pangenome database [45]. Pathway definitions from gene family were provided by MetaCyc database [46].
Statistical analysis
Difference of group comparisons were performed with Student’s t tests and chi-square tests for quantitative and categorical variables, respectively. The homogeneity of variances was tested by using Levene’s test. Variables that followed a Gaussian distribution were compared with two-tailed t tests or analysis of variance (ANOVA). For variables that violated the assumptions of normality or homoscedasticity, the groups were compared with nonparametric Mann-Whitney U test (for two groups) or Kruskal-Wallis test (for more than two groups). Gut microbiota from different samples were compared using the nonparametric statistical methods ADONIS (PERMANOVA; test for source of variation) and ANOSIM (test for similarity) using QIIME 2. F/B ratio, alpha diversity, beta diversity, behavior test and summation of daily oscillation were compared using 2-way ANOVA with Bonferroni’s multiple comparisons test. All analyses were performed with Stata (StataCorp LP, College Station, USA) software, R software (version 3.1.0, the R Project for Statistical Computing), LEfSe [39], or STAMP [47]. A p value < 0.05 was considered significant.
RESULTS
The SNCA p.A53T mice exhibit an age-dependently nigral TH-positive neuronal loss followed by locomotor dysfunction starting from 6 months old.
Recent evidence demonstrates that gut microbiota could influence the motor dysfunctions in a transgenic PD mouse model [7]. However, when gut microbiota composition is shifted to a disease-associated form remains unclear. To track the transformation of gut microbiota in PD, we performed a longitudinal study to collect fecal samples from transgenic heterozygous SNCA p.A53T mice and their littermate controls every 2 months from early adult age at 2 months old to 10 months old. We also concomitantly examine the locomotor function and neuropathology changes with time-sequential multiple measurements.
The neuronal integrity of TH-positive neurons in the substantia nigra was more reduced with age in SNCA p.A53T mice compared to littermate controls (Fig. 1A).
At the age of 2 months old, the number of nigral TH-positive cells was comparable between SNCA p.A53T mice and NTG littermate controls in this early adult stage (Fig. 1A, B and statistics in Fig. 1C). Both groups had progressively reduced number of TH-positive cells with age, while SNCA p.A53T mice showed a significantly lower number of TH-positive neurons in the substantia nigra than their littermate controls at the age of 6 months (2031.38±142.19 vs. 2528.88±177.02, p = 0.01, Fig. 1B and statistics in Fig. 1C). At the age of 10 months old, the SNCA p.A53T mice showed a further lower number of nigral TH-positive neurons than their littermate controls (1195.25±83.67 vs. 2017.50±161.43, p < 0.01, Fig. 1B and statistics in Fig. 1C). Consistently, the neuronal alpha-synuclein expression also revealed an age-dependent increase in nigral TH-positive neurons in the SNCA p.A53T mice compared to littermate controls (Fig. 1B and statistics in Fig. 1D).
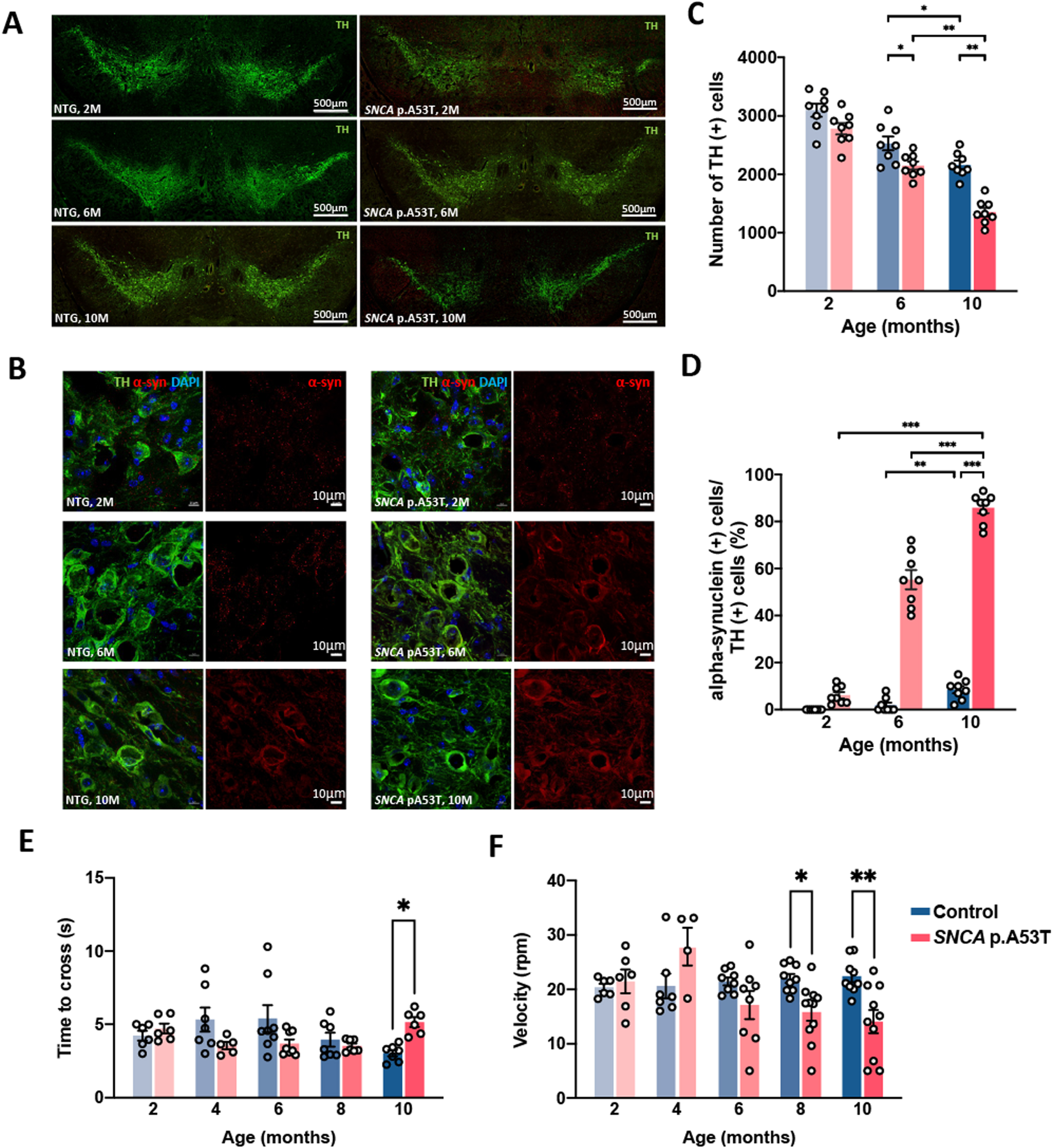
SNCA p.A53T mice showed age-dependent dopaminergic neurodegeneration, neuronal alpha-synuclein accumulations, and locomotor dysfunction. A) Representative mid-brain sections show immunofluorescence staining of tyrosine hydroxylase (TH)-positive cells in substantia nigra in age/sex-matched non-transgenic (NTG) littermate controls (left panel) and the transgenic SNCA p.A53T mice (right panel) at different ages. B) Enlarged representative images of immunofluorescence staining of TH (green), alpha-synuclein (red), and DAPI (4′,6-diamidino-2-phenylindole, blue) in age/sex-matched non-transgenic littermate controls (left panel) and the transgenic SNCA p.A53T mice (right panel) at different ages. C) Average numbers of TH-positive cells of NTG controls and the transgenic SNCA p.A53T mice at different ages. D) The percentage of alpha-synuclein-containing TH-positive cells relative to total TH-positive cells in individual groups of animals at different ages. E, F) Locomotor ability of SNCA p.A53T (red) and control littermate mice (blue) was tested using beam balance (E) and rotarod tests (F). SNCA p.A53T mice completed beam balance tests with a similar time to control littermates at early adult age but were significantly slower at 10 months old. SNCA p.A53T mice showed similar rotarod performance to control mice at early adult age but displayed a significant deficit after 8 months old. n = 4–11 in each group, error bars represent the mean and standard error. *p≤0.05, **p≤0.01 by 2 ways ANOVA with Bonferroni’s multiple comparisons test.
In addition, although there was no gross morphological change in total brain volume between groups (Supplementary Figure 1A), a more reduced hippocampal volume was observed in the SNCA p.A53T mice than in littermate controls in an age-dependent manner (Supplementary Figure 1E). The density of NeuN-positive neurons measured in the primary motor cortex and hippocampus showed a reduced cell number in aged SNCA p.A53T mice compared to littermate controls at the age of 10 months old (Supplementary Figure 1B, F and statistics in Supplementary Figure 1C, G). Notably, the neuronal alpha-synuclein expression was also consistently increased in the motor cortex in aged SNCA p.A53T mice at the age of 10 months old (Supplementary Figure 1B, D). In the hippocampus, however, despite a reduced count of NeuN positive cells in the molecular layer of the hippocampus, there was no apparent alpha-synuclein retention detected in hippocampal neurons in aged SNCA p.A53T mice (Supplementary Figure 1F).
Aside from progressive nigral TH-positive neuronal loss with increased expression of neuronal alpha-synuclein starting from the age of 6 months old, the rotarod test and beam balance test showed that SNCA p.A53T mice had no observable motor dysfunction at 2, 4, and 6 months old. The SNCA p.A53T mice started to show significantly worse performance in the rotarod test at 8 months old and in both rotarod and beam-crossing tests at 10 months old compared to littermate controls (Fig. 1E, F). Together, our results demonstrate that PD neuropathology could be observed at 6 months old, then followed by progressive motor dysfunction in SNCA p.A53T mice.
Altered gut microbiota are observed in young adult PD mice at 2 months old
To investigate the relationship between changes of gut microbiota and PD progression, fecal samples from SNCA p.A53T mice and littermate controls were serially collected at 6 time-points within a 48-hour time period every 2 month from 2 to 10 months old. We found that the average relative abundances of bacterial phyla in fecal microbial communities such as the Firmicutes and the Bacteroidetes did not change significantly throughout the experiment (Supplementary Figure 2A). The ratio of Firmicutes to Bacteroidetes (F/B ratio) showed a significant difference only at the age of 10 months between littermate controls and SNCA p.A53T mice (Supplementary Figure 2B). Thus, the composition of gut microbiota on the phyla level did not show drastic change before the onset of motor symptoms.
To further compare the gut microbiota from SNCA p.A53T and littermate control mice, we performed Principal Coordinate Analysis (PCoA) based on the Unweighted UniFrac, Weighted UniFrac and Bray-Curtis distance matrix at different ages. UniFrac distance includes phylogeny information while Bray-Curtis does not. The gut microbiota of SNCA p.A53T and control mice from all age points were clustered into two different regions in PCoA plots (Fig. 2A-E, unweighted UniFrac). The ANOSIM (mean) and ADONIS (dissimilarity, beta diversity) tests for gut microbiota were significantly different at all time points between SNCA p.A53T and control mice at 10 months old and some time point at 8 months old when motor impairments were observed (Supplementary Table 1). Surprisingly, the gut microbiota was also significantly different at some time point even at the 2, 4, and 6 months old when there were no observable motor deficits (Figs. 1E-F and 2A-C, Supplementary Table 1). There were more and more time points that show significant different at later age. Furthermore, the gut microbiota composition from SNCA p.A53T mice and control seems to deviate from each other in different directions in the PCoA plot (Fig. 2F). At many early age time points, we observed significant difference in ANOSIM and ADONIS tests with Bray-Curtis but not weighted Unifrac distance matrix, suggesting that the difference of gut microbiota between control and SNCA p.A53T mice may start from shifting specific microbes with closer phylogenetic relationship. There was no significant difference in alpha diversity between SNCA p.A53T mice and littermate controls (Fig. 2G), suggesting that the gut micro-environment did not favor specific gut microbe yet at early PD progression. Together, these results showed that the diversity of gut microbes from SNCA p.A53T mice is significantly different from their littermate controls at the early adult stage.
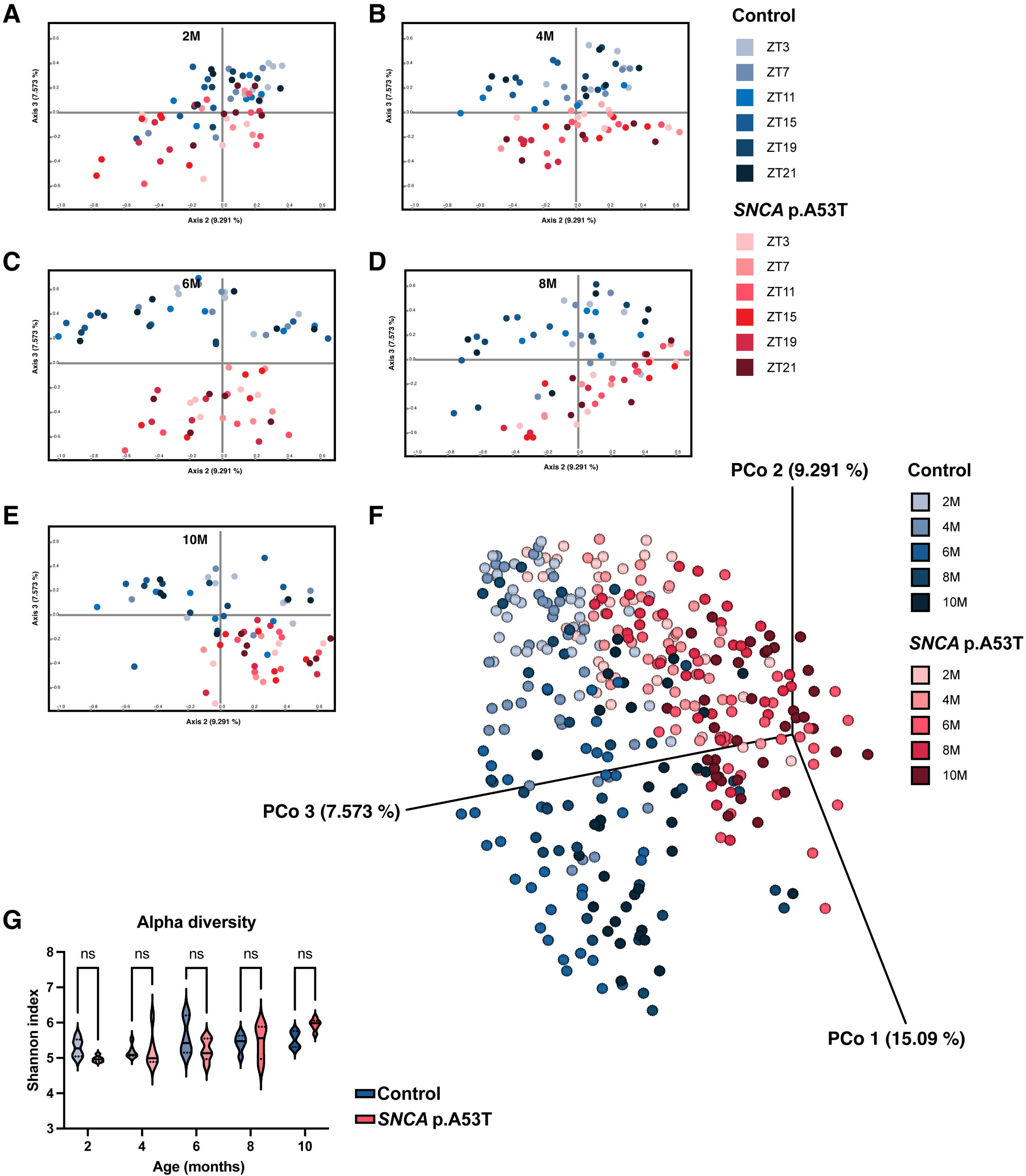
SNCA p.A53T mice had significant different gut microbiota at early adult age. A-E) Principal coordinate analysis (PCoA) of gut microbiota between SNCA p.A53T (red) and littermate control (blue) mice at 2 (A), 4 (B), 6 (C), 8 (D), and 10 (E) months old. At all time points, gut microbes from SNCA p.A53T and control mice were clustered at different areas. F) PCoA plot including all data from different ages. The clusters of gut microbes between SNCA p.A53T and control were moving in different directions over time. G) Gut microbial alpha diversity between SNCA p.A53T and control mice. The diversity with Shannon index showed no significant difference between SNCA p.A53T and control mice. Shannon indexes were average from 6 time points for each mouse for 2-way ANOVA test. n = 5 in SNCA p.A53T 6 month data, n = 6 in all other groups, violin plot shows the distribution of data at different values, and three markers are 25th, median, and 75th percentiles. *p≤0.05, **p≤0.01, ***p≤0.001, ****p≤0.0001 by 2 ways ANOVA with Bonferroni’s multiple comparisons.
To determine which gut microbes are enriched in either SNCA p.A53T or control mice, we performed Linear discrimination analysis Effect Size (LEfSe) at 5 different age points (Fig. 3A-E). We found relative abundance of some microbes were downregulated in SNCA p.A53T mice compared to littermate control. Interestingly, the number of PD enriched features is trending up from young to old mice. This age-related progressively shifting of gut microbe features could be easily observed in cladograms with expanding PD enriched branches (Fig. 3). We also found that the family of Rikenellaceae is significantly higher in control mice, while the genus of Lactobacillus is significantly higher in SNCA p.A53T mice of all ages. Furthermore, some genus such as Parabacteroides and Ruminococcus was higher in the SNCA p.A53T mice only at later stages (8 and 10 months for Parabacteroides and 10 months for Ruminococcus). Therefore, our results showed that dysbiosis could be observed in PD mice even at the early adult age of around 2 months old. We also performed multiple regression analysis using the relative abundance of 51 identified microbes at order level from SNCA p.A53T mice and littermate control at all ages and their corresponding rotarod behavior data. We found that overall gut microbes composition has a linear relationship with rotarod performance. The p-value is 0.028, and the R2 is 0.7876. Although we could not find a single gut microbe that could predict the rotarod performance from the multiple regression analysis, our result suggested that global gut microbe composition is highly correlated to motor function impairment in PD mice.
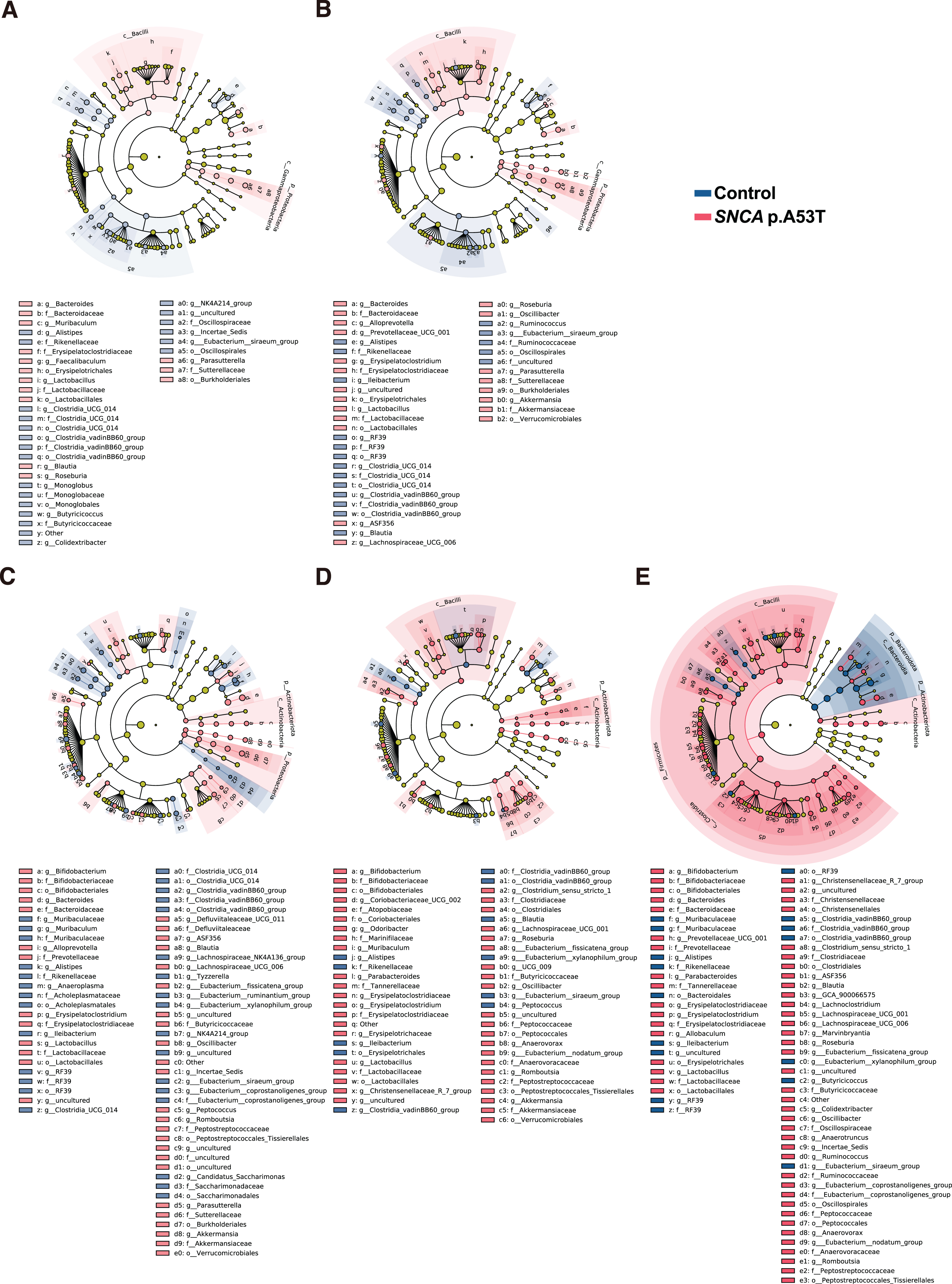
SNCA p.A53T mice showed age-related gut microbes changes. A-E) List of enriched gut microbes at different taxonomic levels in SNCA p.A53T (red) and control mice using Linear discrimination analysis Effect Size (LEfSe) test at 2 (A), 4 (B), 6 (C), 8 (D), and 10 (E) months old. n = 5–6 in each group, p < 0.05, LDA score < 2.0 by LEfSe.
To test whether this early dysbiosis is specific to SNCA p.A53T PD model, we next examine the gut microbiota in another PD genetic model, which harbors the most prevalent LRRK2 p.G2019S mutation in PD patients. Similar to SNCA p.A53T, LRRK2 p.G2019S mice exhibited progressive motor dysfunction at the age of 10 months old [32]. At 2-month-old, while the overall phylum level of Firmucutes and Bacteroidetes were comparable between LRRK2 p.G2019S and littermate control mice (Fig. 4A), the gut microbiota composition from the LRRK2 p.G2019S mice and their littermate controls were clustered in two group using PCoA analysis (Fig. 4B). There were significant differences in both alpha and beta diversity between LRRK2 p.G2019S and control mice during the night time (Fig. 4C, D). The LEfSe analysis also shows that bacteria genera such as Anaerotruncus, Lachnoclostridium, Lachnospiracease UCG001, Anaerovorax, Eubacterium nodatum group, Eubaterium coprostanoligenes group, and Ruminococcus were enriched in LRRK2 p.G2019S mice (Fig. 4E), similar to the findings observed in SNCA p.A53T mice. Interestingly, many of these gut microbes were enriched in aged SNCA p.A53T mice. Together, our results showed that in two different PD mouse models, the dysbiosis could be observed as early as they entered the adult stage, which was earlier than the occurrence of neuropathology and the onset of motor dysfunction. Therefore, it suggests that a common mechanism that influences the gut microenvironment may be involved in causing early dysbiosis in PD.
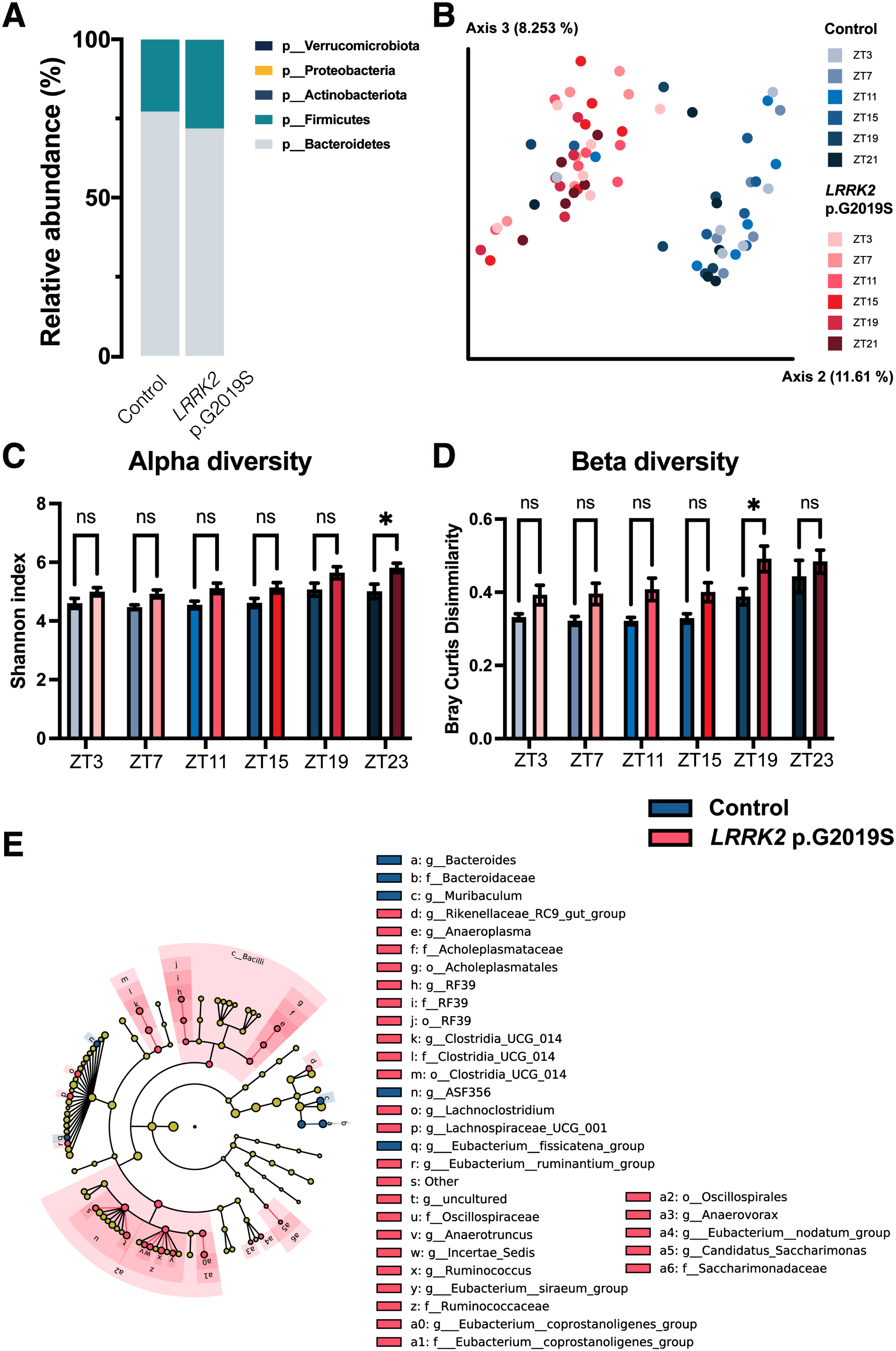
LRRK2 p.G2019S mice showed dysbiosis at 2 months old. A) Gut microbe’s relative abundance at phylum level from control and LRRK2 p.G2019S mice. B) Principal coordinate analysis (PCoA) of gut microbiota using unweighted unifrac distance for LRRK2 p.G2019S (red) and littermate control (blue) mice at 2 months old. Gut microbiota samples from 2 groups of mice were clustered at different areas. C, D) Bar graph of gut microbial alpha (C) and beta (D) diversity obtained from LRRK2 p.G2019S and control mice. Alpha diversity from all time points showed an increased trend with significant different at ZT23. Beta diversity also displayed an increased trend with significantly different at ZT19. n = 6 in each group. * p≤0.05 by two-way ANOVA test. E) List of enriched gut microbes at different taxonomic levels in LRRK2 p.G2019S (red) and control mice (blue) using Linear discrimination analysis Effect Size (LEfSe) test at 2 months old. Cladograms showing enriched gut microbes in LRRK2 G2019S mice and littermate controls at 2 months old. n = 6 in each group, p < 0.05, LDA score < 2.0 by LEfSe.
The daily oscillation of gut microbes is dampened in SNCA p.A53T mice
Another interesting aspect of gut microbes is that 15–30% of them show daily oscillation [11, 25]. Since circadian disorder could be observed in PD patients, we would like to investigate the daily oscillation of gut microbe in SNCA p.A53T mice. We analyzed the daily oscillation of gut microbes from SNCA p.A53T mice and littermate control from 2 months to 10 months old by sequencing fecal samples collected from 6 time-points (ZT3, 7, 11, 15, 19, and 23). The JTK_cycle analysis was used to detect the 24 h oscillation of each bacterial operational taxonomic unit (OUT) within the same groups of animals (Fig. 5A-E). The relative abundance of each oscillating OUT was combined to indicate the total amount of gut microbes with daily oscillation (Fig. 5F). At 2 months old, the PD mice showed higher amounts of microbes with daily rhythm but quickly dampened after 4 months old, and then lost all microbial oscillation at 10 months old. In comparison, the control mice exhibited a relatively stable number of oscillating microbes from 2 months old to 8 months old. The partially dampened daily microbial oscillation of control mice could only be observed at 10 months old. Together, our results showed that the gut microbiota from SNCA p.A53T mice is different from control littermate at the earliest adult age at 2 months old. The composition and daily oscillation showed differences much earlier than the onset of motor symptoms in the process of PD development.
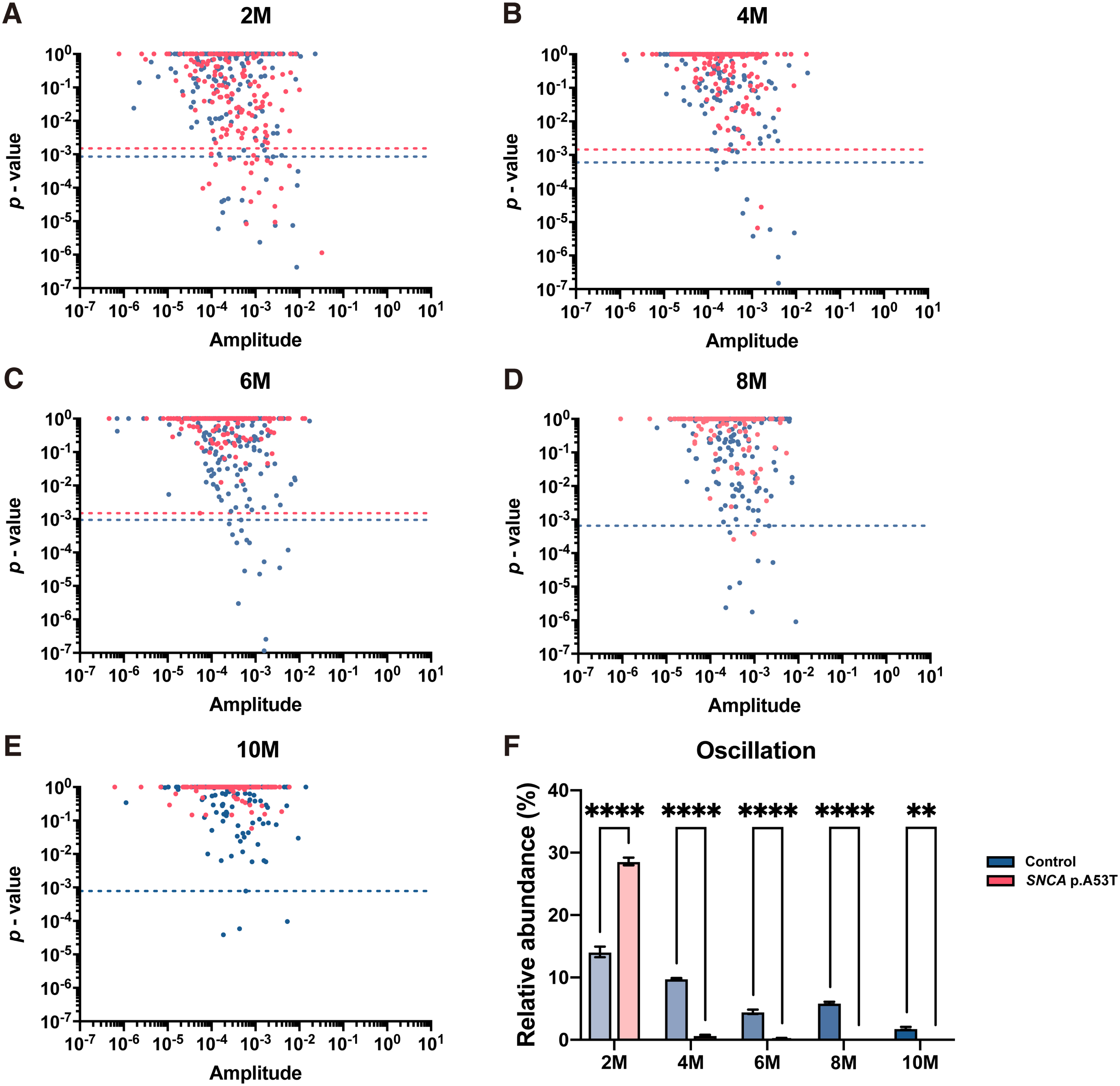
Gut microbiota daily oscillation in SNCA p.A53T mice. A-E) ASV-based JTK_CYCLE analysis shows diurnal oscillating microbes from SNCA p.A53T (red) and control (blue) mice at 2 (A), 4 (B), 6 (C), 8 (D), and 10 (E) months old. Dash lines indicate the Benjamini-Hochberg q-value < 0.05. F) The total percentage of gut microbe displaying daily oscillation for each mouse was summed from the relative abundance of all microbes with both q-value and p-value smaller than 0.05. From 4 months old, the percentage of oscillating microbes in SCNA p.A53T mice was significantly dampened compared to control mice. n = 5–6 in each group, error bars represent the mean and standard error. **p≤0.01, ****p≤0.0001 by 2 ways ANOVA with Bonferroni’s multiple comparisons test.
Dysbiosis of PD patients using metagenomic sequencing
Finally, to see whether the dysbiosis in PD mice could be observed in human PD patients. We compare the difference in gut microbiota from 58 PD patients (mean age: 64.16±10.43 years old, 62.1% were men) and 46 control participants (mean age: 63.61±10.96 years old, 60.9% were men) using fecal metagenomic sequencing analysis (Table 1). Here we found several PD patients enriched gut microbes, including Methanobrevibacter Smithii, Allistipes Shahii, Parabateroides Merdae, Cloacibacillus Porcorum, Ruminococcus torques, and Eubacterium Hallii (Fig. 6A). Many of the above microbes have been reported previously as PD patients enriched gut microbes using 16S rRNA sequencing at the genus or family level [48]. Finally, using metabolic pathway analysis, we found that the peptidoglycan maturation pathway is downregulated in PD patients. Interestingly, 20 metabolic pathways are significantly higher in PD patients than in controls, including pyruvate fermentation to propanoate II pathway (Fig. 6B). Together, our results showed that gut microbiota and their metabolic pathways are different between PD patients and controls. As we showed earlier that genus Parabacteroides and Ruminococcus are higher in both PD mouse models than in their littermates, our study consistently observes enrichment of Parabacteroides and Ruminococcus in human PD patients and 2 PD mouse models using different sequencing methods. These data suggest a strong association between PD and two microbes, Parabacteroides and Ruminococcus.
Clinical characteristics of participants in the current study
BMI, body mass index; LEDD, levodopa equivalent daily dose; MDS-UPDRS, Movement Disorder Society Unified PD Rating Scale; MMSE, Mini-Mental State Examination; N.A., not available; PD, Parkinson’s disease.
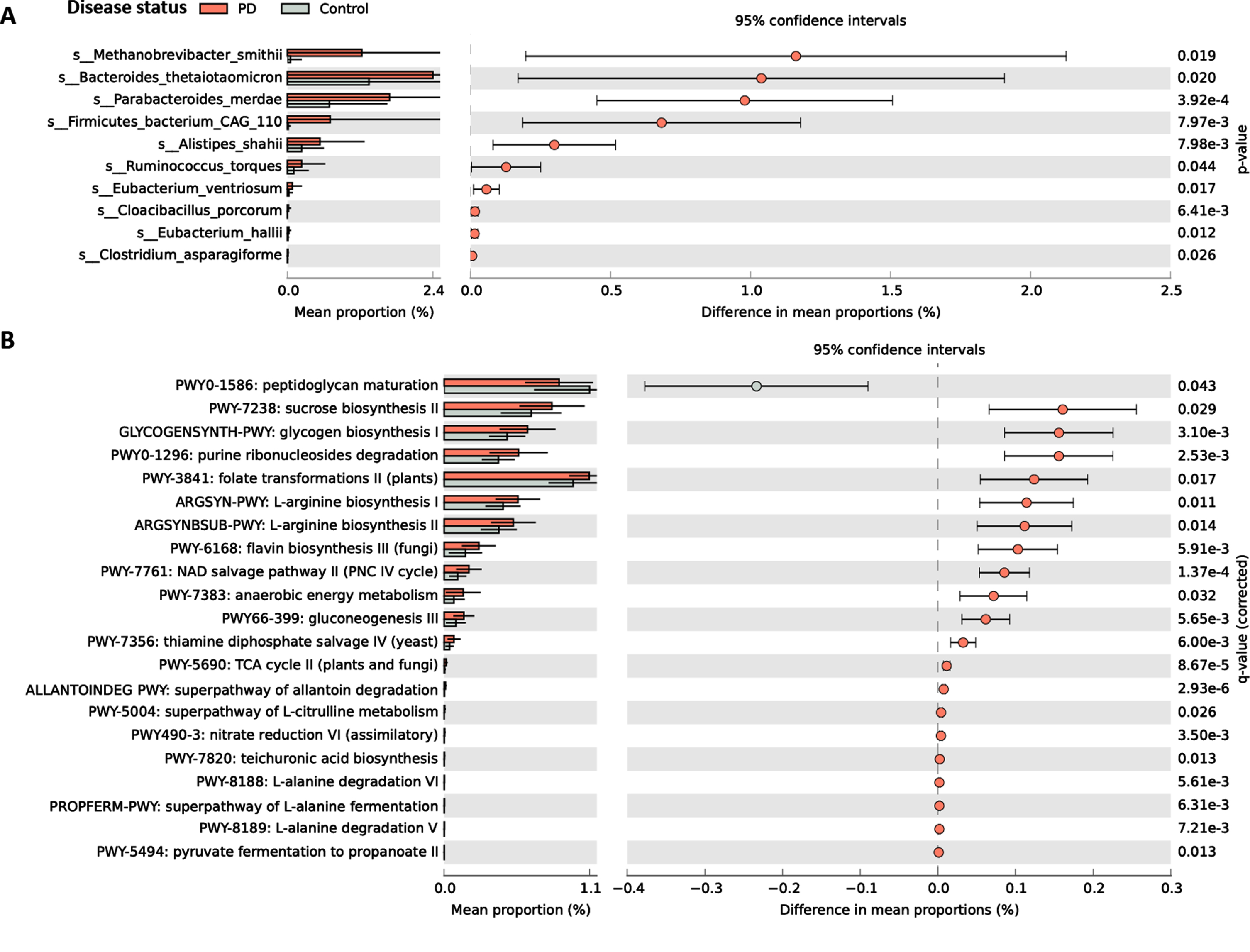
Gut microbe and metabolic pathway difference in PD patients and control subjects. A) The mean relative abundances of the gut bacterial species with significant difference between PD patients and control participants from shotgun metagenomic data. b) The mean relative abundances of the MetaCyc pathways with significant difference between PD patients and control participants from shotgun metagenomic data. n = 58 PD patients and 46 controls, Benjamini-Hochberg q-value < 0.05 by STAMP.
DISCUSSION
This study provides longitudinally follow-up evidence that dysbiosis and dampened daily oscillation of gut microbiota can be observed at an early age in advance of neuropathology and motor symptoms in the PD mouse model. Our study showed that the gut microbiota not only is different at 2 months old mice between SNCA p.A53T and control mice, the composition of the gut microbiome seems to deviate through age. We also found that daily oscillations of gut microbes were also highly blunted with loss of oscillations in SNCA p.A53T mice starting from 4 months old, while the motor deficits are observed starting at 8 months old. Similar changes in altered gut microbiota were also observed in another LRRK2 p.G2019S PD genetic mouse model. Our results revealed the altered gut microenvironment and dampened microbiota oscillations preceding the occurrence of neuropathy and motor dysfunction in the PD process.
Several neurotoxin and transgenic rodent models have demonstrated the changes of gut microbes in relation to PD motor dysfunction and neuropathology (summarized in the Supplementary Table 3). However, most of the studies exploring the changes of gut microbes in PD rodent models are cross-sectional study designs. Only one study examined the weekly change of gut microbes in a short time period of 4 weeks [49]. In this study, we serially analyzed monthly changes of gut microbiome composition starting from the age of 2 months to the age of 10 months in two commonly used PD genetic rodent models, which has been rarely assessed previously in the literature. Although we could not collect fecal samples from the same animal from 2 to 10 months, the age and sex-matched littermate control pairs were strictly used in this study. We demonstrated that dysbiosis in a PD mouse model carrying the PD-causative SNCA p.A53T mutation can be detected as early as 2 months old and dampening of daily oscillation of gut microbes was exhibited in 4 months old. There is no study about gut microbiota in the PD mouse model at the early adult stage.
Although we found significant differences in gut microbiome compositions between groups at 2 months old, it is possible that earlier changes, e.g., from birth itself, existed. The development of the gut microbiome begins in utero although the actual birth marks the first major colonization of the infants [50]. The host genetic background could be one of the factors that modulates the gut microbiome ecosystem. Genome-wide association studies have identified numerous genetic risks that increase susceptibility to PD, and several genes are related to gut microbial regulation and intestinal inflammation, including toll-like receptors, peptidoglycan recognition protein, and MUC2, a component of the mucosal layer that protects the intestinal epithelial barrier [51]. Some of the identified genetic susceptibility to sporadic PD also increases the risk of inflammatory bowel disorders (IBD), among which, NOD2 is known to be a strong predictor of IBD and to interact with LRRK2. Therefore, PD-related mutations such as SNCA p.A53T and LRRK2 p.G2019S may promote the digestive tract of the mice to favor the colonization of these pro-inflammation microbes through an unknown mechanism. Future studies utilizing germ-free housing techniques and colonization with specific gut microbes are needed to confirm the causal-link relationship between the early gut dysbiosis and PD neuropathology in both ENS and brain.
It has been shown that diurnal oscillation in gut microbiota can be regulated by the host circadian clock [11, 25], feeding pattern [11], diet [53], and light dark cycle [54]. Loss of diurnal oscillation in gut microbiota also happens in diseases like type 2 diabetes [55, 56]. Besides, the alpha-synuclein overexpressing transgenic line mice display disrupted circadian rhythm and abnormal sleep-wake cycle from an early age around 3-4 months old [57]. Here we also showed that gut microbiota daily oscillation in SNCA p.A53T mice is significantly dampened compared to littermate controls from 4 months old to 10 months old. Since the light-dark cycle is an important zeitgeber for circadian photoentrainment and could influence gut microbiota, it is possible that signals for the environmental light-dark cycle may be compromised in SNCA p.A53T mice at an early age. Notably, we paradoxically observed that that the transgenic SNCA p.A53T group had a higher proportion of gut microbiota showing diurnal variation compared to the littermate controls at 2 months old, which is earlier than the disrupted circadian rhythm and abnormal sleep-wake cycle previously observed in alpha-synuclein overexpressing transgenic mice [57]. In addition to the central circadian clock in SCN, intestinal epithelial cells (IECs) also exhibit unique transcription oscillations of the core clock genes [58]. These peripheral clocks can display local control and receive external signals independent of the SCN to manipulate variable gut functions such as intestinal motility [59]. Therefore, the early transcriptional regulation of gut intrinsic clock genes in IECs of the SNCA p.A53T mice may be altered to partially explain the activated diurnal oscillation of gut microbiota at the age of 2 months. In addition, previous studies showed that immune signaling in the intestine epithelium displays daily oscillation is contributed by gut microbes [19, 60]. Therefore, early dysbiosis and altered gut microbes oscillation may perturb the gut innate immunity and cause intestinal cell dyshomeostasis, leading to gut barrier impairment and intestinal inflammation, which were commonly observed in both PD rodent models and patients with PD [61]. Nevertheless, how did the daily oscillation of gut microbiota start to dampen around 4 months old in SNCA p.A53T mice requires further investigation. The causal relationship between early dampened gut microbiota oscillations in the process of ENS neurodegeneration and gut a-synucleinopathy needs further investigation incorporating germ-free model system with candidate microbiota transplantation.
Recent evidence has shown that the aggregation of alpha-synuclein in the gut ENS is earlier than in the central nervous system in PD patients [2, 3]. Since the ENS regulates the major digestive tract functions such as gut motility, immune response, and intestinal barrier function which could all modulate gut microbes [62, 63], therefore, the potential early ENS dysfunction in SNCA p.A53T mice may contribute to the gut microbiota difference we observed as early as 2 months old age. Alternatively, the dysbiosis in SNCA p.A53T mice may precede the ENS dysfunction. Earlier studies have shown that the pathological alpha-synuclein aggregation in ENS could be observed as early as 3 months old in PD mice model with alpha-synucleinopathy [64, 65]. These previous findings and our current study are in line with one recent study in human PD patients that showed different gut microbiome signatures in the prodromal phase of PD [66]. Therefore, it is possible that hosts harboring the PD susceptibility genetic factors, for example, SNCA and LRRK2 mutations, combined with early gut microenvironmental changes, such as early gut dysbiosis, may trigger the disease process of PD. This dual-hit hypothesis suggests that altered gut microbiota communities may act as an environmental trigger in susceptible hosts to allow neurotoxic pathogen across the gut barrier and then enters the brain through the vagus nerve to cause PD [67]. In support of this hypothesis, gut microbiota transplantation from human PD patients in alpha-synuclein overexpressing mice aggravating PD neuropathology and motor dysfunction highlight the involvement of the gut microbiota in the induction of PD motor symptoms [7]. Our results in this longitudinally study design identifies that gut microbiota changes could be detected as early as the age of 2 months old, which is ahead of neuropathology in the central nervous system by 4 months, provide further support that altered gut microenvironments may act as a trigger in the PD process. Further studies are needed to untangle a myriad of factors, such as specific gut microbiota, gut metabolites, and receptors, which are critical for the dyshomeostasis of gut barriers and maintenance of intestinal immunity, to the pathogenesis of PD.
Previous studies have showed some upregulated bacteria, such as Ruminococcus and Lactobacillus, and some down-regulated bacteria in human patients with PD compared to unaffected controls [48]. Our current metagenomic data from human PD patients also showed enrichment of Ruminococcus torques and Parabateroides Merdae in the patients. Here we also found that genera Ruminococcus and Parabateroides are higher in SNCA p.A53T mice and LRRK2 p.G2019S mice than in their littermate controls. These observations indicate a specific association of Ruminococcus and Parabateroides with PD cross-species in different genetic PD mouse models and PD patients. Additionally, we also observed that Lactobacillus is enriched in SNCA p.A53T mice of all ages. Consistent with our findings, the increased abundance of the genus Lactobacillus and family Ruminococcaceae has been reported in patients with PD compared to healthy controls in several meta-analysis studies [48, 68]. Furthermore, after controlling age and disease duration, increased fecal levels of Lactobacillus correlated with both worse motor and cognitive function in PD patients [68]. The Lactobacillus strain is commonly considered beneficial bacteria. Whether they could influence PD or are simply well adapted to thrive in a pro-inflammatory gut environment remains to be elucidated. These results indicate that the gut microenvironment in PD mice may favor some specific microbes starting at an early age throughout the whole disease progression. In addition, humans and mice may share some common PD-associated gut microbes. It has been shown that Ruminococcus gnavus is associated with human Crohn’s disease by producing an inflammatory polysaccharide [69]. In this study, we found that Ruminococcus torques, which could contribute to intestinal propionate formation [70], is enriched in PD patients. With another PD enriched propionate formation microbe Eubacterium hallii [71], our metabolic pathway analysis showed that the propionate fermentation pathway is significantly higher in PD patients. Together, our data is in line with the previous report showing that the short-chain fatty acid is elevated in serum from the PD patients than in controls [42]. Many species under the genus Clostridium were observed in human and mouse gut flora and are considered probiotic. However, a particular species Clostridium difficile has been shown to cause diarrhea. Therefore, upregulation of specific gut microbes such as Ruminococcus, Lactobacillus, or Clostridium may influence the disease progression of PD through inflammation. Together, we showed that the dysbiosis of PD mice might occur early and start to accelerate the PD progression at a certain stage after the age-related accumulation of pro-inflammation microbes.
The strength of this study is to investigate the changes in gut microbiota, locomotor function, and neuropathology with time-sequential multiple measurements in different PD mouse models. The results were further compared to the fecal microbiota data obtained from a cohort of PD patients with age, sex, and diet habit matched controls. Our study has some limitations. First, the whole data set was obtained from many cohorts of mice. We could not collect fecal samples from the same animal from 2 to 10 months old. Therefore, we did not compare gut microbiota difference between month in the same animal. However, the littermate control pairs were strictly used in this study. Second, the PD rodent model we used in this study was those carrying the SNCA p.A53T and LRRK2 p.G2019S mutation. Although the genetic forms of PD may not represent the majority of sporadic PD patients, they could still provide suitable mammalian models for recapitulating clinical and pathological changes in human PD patients. Finally, age has been demonstrated to have an impact in gut microbiota. Elderly people have different gut microbiota compared with young adults and this difference can be attributed to several factors, including diet habits, lifestyle, medications, and co-morbidities. The genetic background of the host and the host-microbiome interactions also contribute to the complex evolution of gut microbiome from birth to aging, especially in the chronic disease status, such as PD. Future work integrated with polyomic data, including gut microbiota metagenomic analysis in PD patients with known genetic risk factors for disease combined with food frequency questionnaire in a time-series follow-up design is needed to confirm the early dysbiosis in disease progression of PD.
In summary, our results from different PD mice models demonstrated altered gut microbiota communities and dampened microbiota oscillation ahead of dopaminergic neuronal degeneration and locomotor dysfunction. Further studies to identify the initiation factors of PD dysbiosis at an early age may help understand the disease pathogenesis at the initial stage of PD. Our study highlights the possibility of a potential surrogate gut microbiota marker for the early detection of PD, which may shed light on the future early intervention before the onset of motor symptoms in PD patients.
Footnotes
ACKNOWLEDGMENTS
We thank the Technology Commons, College of Life Science at National Taiwan University for technical assistance with NGS, Computer and Information Networking Center, National Taiwan University for the support of high-performance computing facilities. We are also grateful for the technical assistance provided by the First and Second Core Labs in the National Taiwan University College of Medicine, Taipei, Taiwan.
This work was supported by the Taiwan Ministry of Science and Technology for the grant MOST 109-2636-B-002-005 (to S.-K. C.) and grant MOST 109-2320-B-002-073 (to C.-H. L.). We are also grateful for the funding support from National Health Research Institute (NHRI-EX109-10716NC) and National Taiwan University (NTU-110-A-CC-5400-64841).
CONFLICT OF INTEREST
All authors report no conflicts of interest.
DATA AVAILABILITY STATEMENT
Source data are provided with this paper. The datasets generated during and analyzed during the current study are available in the public database of European Nucleotide Archive.
