Abstract
A variety of mathematical models is used to describe and simulate the multitude of natural processes examined in life sciences. In this paper we present a scalable and adjustable foundation for the simulation of natural systems. Based on neighborhood relations in graphs and the complex interactions in cellular automata, the model uses recurrence relations to simulate changes on a mesoscopic scale. This implicit definition allows for the manipulation of every aspect of the model even during simulation. The definition of value rules
Introduction
The simulation of complex physical, chemical, and biological phenomena has become an important tool for the research in life sciences. Models that mimic the behavior of natural systems support the understanding and analysis of laboratory experiments [1]. In systems and cell biology such models are valuable for inferring the life-sustaining chemical transformations in cells [2], the self-organization of the cytoskeleton synthesis [3], and phenotypes of whole organisms [4]. Quantitative models are able to give actual predictions about the quantities in a system. Most models are generally systems of ordinary or partial differential equations (PDE). Such systems are composed of multiple equations, where each equation represents the concentration of a chemical compound. It is of high importance to model these equations as accurately as possible to correctly capture the phenomena observed on the biological level. Even small differences in the initial conditions may produce large discrepancies between the simulation and the experimental outcome [5]. Hence, each experimental model has to be addressed differently, and many parameters have to be measured and set to describe the phenomenon observed under specific conditions. While helpful for a small group of scientists, those models can rarely be adapted for problems that are the result of a similar underlying phenomenons [6]. Scientists can only observe consequences of physical and chemical laws that are entangled in complex systems such as cells and organisms. So it is no surprise, that most models do not describe the “real shape” of the problem and only characterize the guise of the phenomenon. For an illustration of the problem, consider a medical doctor who tries to cure an illness. Only by approaching and treating the underlying causes, he can hope to cure the patient. Treating the symptoms is only viable in the most severe cases. Similarly, if we are able to characterize the processes in a cell and know their interactions, it is possible to modify the system and obtain the behavior that is required. The creation of new fundamental laws is a large step in the development of systems biology [7]. Modeling those fundamental processes and combining them as system-theoretic concepts is still one of the greatest challenges in the field of systems biology [8, 9].
One technique to explore the general traits of interacting systems and spatio-temporal patterns are cellular automata (CA). Originated in the 1940s, CA are simulations discrete in time, space, and state. By applying a simple set of rules – which depend on automata elements, so-called cells, and their respective associations between each other – at each time-step, remarkably complex behaviors emerge from such automata [10, 11]. This discrete character to CA is tailored for the computation using traditional computers that operate at finite precision by design. Dynamic Cellular Automata, as an agent-based adaptation of CA, have been shown to describe a multitude of different cellular or biochemical processes [12]. Although CA have been used in many fields of biology [13], they played a minor role in recent years, because powerful tools such as ECell [14] and VCell [15] started to dominate the research field. Both tools and many others alike are based on PDEs that are composed and specified by the user of the software. With all reactions and processes defined, a numerical solution of the system of PDE is generated and calculated. The user is able to inspect the results and adjust parameters to fine-tune the model to fit the results gained via laboratoryexperiments.
Space is one of the key components when modeling the processes in cells. In the last decade simulations in systems biology have been moving from temporal-only models to spatio-temporal models [16]. An established practice for simulations with a spatial component is to partition the simulation space into smaller compartments that are able to exchange components or information. This holds true for CA, which are traditionally defined to handle spatial neighborhood relations on two- or three dimensional geometric tilings. The neighbors of each cell are defined by adjacent edges or corners.
Another approach to model natural systems are stochastic simulation algorithms (SSA) such as Chemical Master Equations (CME), which can be necessary when biological phenomena depend on stochastic fluctuations [17]. CME are able to model diffusion using particle position distributions – which are either compartment-based or reliant on Smoluchowski equations – and chemical reactions, based on probabilities of molecular reactions during a defined time period. Assuming a spatially homogeneous system the changes in concentration can be simulated.
PDEs, SSA and CA represent different techniques to explore the behavior of dynamic systems, all designed to define the change that occurs in events or time steps. In PDEs this change is explicitly defined from the starting point and for all following steps in time. Therefore it is possible to poll the state of the system by solving a single equation. In comparison, using CA, numerical methods, or SSE the next state of the system is implicitly defined by its current state, thus adjusting parameters or even states can be accomplished readily at every time step, but every state in between the current and the required time step has to be calculated [18, 19]. This leads to a consideration of the advantages and disadvantages of implicit definitions in opposition to explicit definitions. The updating procedure of explicit systems depends on the value of the previous time step, and may even be analytically formulated as a closed form solution. Therefore the determination of the updates is straightforward and computationally inexpensive. As a compromise, explicit methods are only conditionally stable, meaning errors at any stage of the computation or divergence in the starting conditions are amplified and not attenuated as the simulation progresses. Hence, explicit solutions require very small time steps, especially when there are major changes in a short time period. Implicit systems can include values of the current and previous time steps. The calculations performed in each time step are generally more difficult, resulting in a higher time complexity of the simulation. Nevertheless, implicit schemes are stable and can tolerate larger time steps than explicitly defined systems [20]. This also includes a robustness to unexpected changes. Especially in biological environments the ability to respond and adjust to spontaneous disturbances, such as molecular signals from other cells or changes in temperature is vital in order to create a realistic and robust model [21].
In 2001
Methods
The concept of
Modeling cellular graph automata
The definition of
For the adaption of
A
The position of a node in a space of dimension
CGA are composed of nodes, and unlike CA, their neighborhood relations are based upon the connections represented by edges. Let
While CA operate in a predefined tiling, GAC act depending on their neighbors, defined by
The cardinality of the multiset is the sum of all multiplicity functions and equal to the degree
As an example consider a node
The transition of a node from one state
This deterministic transition function determines the next state of a node, by the number of neighboring nodes in a certain state. If the number of nodes in state
With these definitions the state of a node can be set. Considering the initial representation of nodes as a segment of space inside of a biological system, possible states could be comprised of the environment or compartment a node is enclosing. In this context, the state of a node may be any organelle, such as chloroplast, endoplasmic reticulum, or Golgi apparatus. The states or objects can be allowed to move through the cell with rules adapted from the String definition in [26] or remain static. In order to handle continuous values, such as concentrations of species, the previous definition is augmented to
The description
The Definition 2 of GAC also applies to the multiset
The domain
The set of value rules

A section of a cellular graph automaton illustrating nodes, edges, and Voronoi regions. A set of species
The idea behind this definition is the creation of multiple value rules that represent natural phenomena inside of the cell. It is possible to define multiple value rules that modify the same value in a value set. For example, consider the diffusion of a species into neighboring compartments in addition to the conversion into another species during a chemical reaction. Both events would modify the concentration of the same species. In this case the sum of the changes for each species and value function will be applied to the node.
The size of the time step is uniform and can be parametrized as described in Section 2.3. This enables the simulation of multiple time scales depending on the phenomenon that is to be addressed. Additionally, the size of the time step provides the possibility to choose between a more accurate prediction (using numerous small time steps) or a faster simulation (using fewer large time steps). A schematic representation of the sequence of operations that are needed to setup and perform a simulation is depicted in Fig. 2.
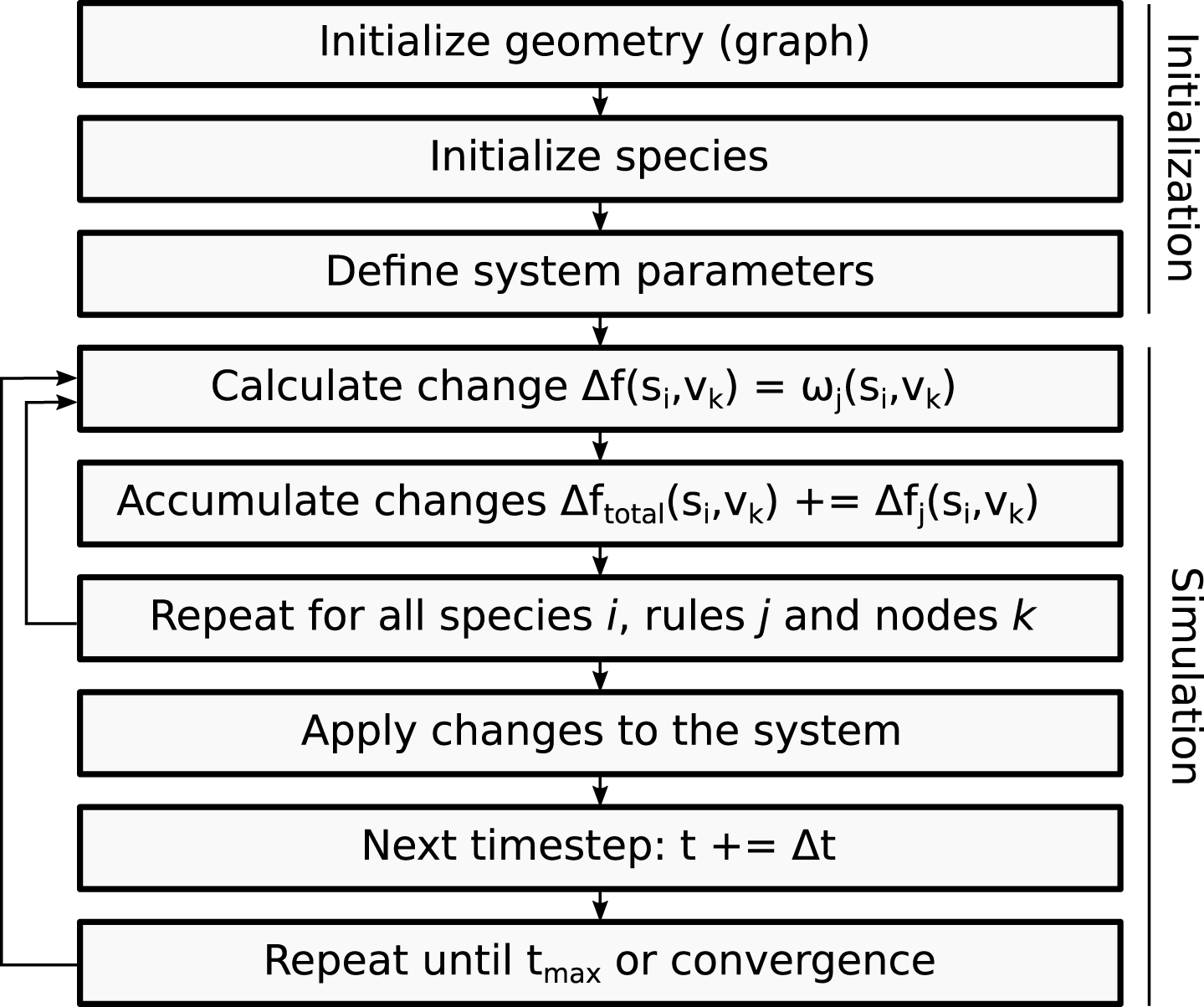
The flowchart describes the general sequence of steps during the initialization process and actual simulation of CGA.
To evaluate the proposed model, the process of diffusion has been implemented as a value rule
Diffusion is a natural physical process in which molecules disperse from an area of high concentration proportional to a concentration gradient. This phenomenon is describes the random motion of all particles in solutions, called Brownian motion. Fick proposed the Second Law of Diffusion, in which the flux of matter in a liquid is proportional to the gradient of its concentration
This equation needs to be transformed into a recurrence relation, which then enables the implementation into CGA. Using the finite differences method as shown in [20], the infinitesimal small steps in time ∂
By calculating the limit with Δ
Since the time scale of CGA is uniform and homogenous, it is possible to assume Δ
Now it is possible to translate the equation to a value rule usable in CGA. The new concentration of a species
As mentioned in the definition of CGA, the spatial and time scales of CGA are dimensionless and uniform. The scaling factors that result from the true dimensionality of the system (time step size and node distance) have to be included in the value or transition rules, in order to affect the model system.
The diffusion coefficient
The distance between two nodes Δ

The first step to setup a simulation with cellular graph automata is to define the desired geometry of the system that one wants to simulate. Afterwards, the system is partitioned by covering it with a net-like graph. Each node is responsible for a subsection of space. Further adjustment methods can be applied to truncate and rearrange nodes, so that the true geometry is represented accurately. After the CGA has been adjusted to cover the simulation space, chemical species can be added to the system. Species and their attributes can be retrieved from databases such as ChEBI [35] or Pubchem [34]. For every node in the system the concentration can be set individually, a gradient qualitatively indicates nodes with a high concentration of a particular species in darker color and low concentrations in light color.
The viability of the simulated diffusion process, calculated by the implementation of the suggested model, is assessed in comparison to a numerical solution of the two-dimensional representation of the diffusion equation [30]. To compare the different approaches, the half-life
The sample system to be simulated was constructed as a square with a side length of 2500 nm. This square is partitioned into two rectangles with a width of 1250 nm (half of the system) and a height of 2500 nm. One of the rectangles is set up to have a starting concentration of 1.0 mol·l-1 and the other rectangle is set to a concentration of 0.0 mol·l-1. The system’s initial setup and progression is depicted in Fig. 4. Multiple species have been considered to assess the viability of the system. We have chosen some of the smallest molecules that may be of interest in cellular biology. Those species should be the most critical in the system because large quantities are moving between nodes. The diffusivity covers ranges from 4.40·10-5cm2·s-1 for hydrogen gas up to 6.40·10-6cm2·s-1 for ethane-1.2-diol (seeTable 1) [28].
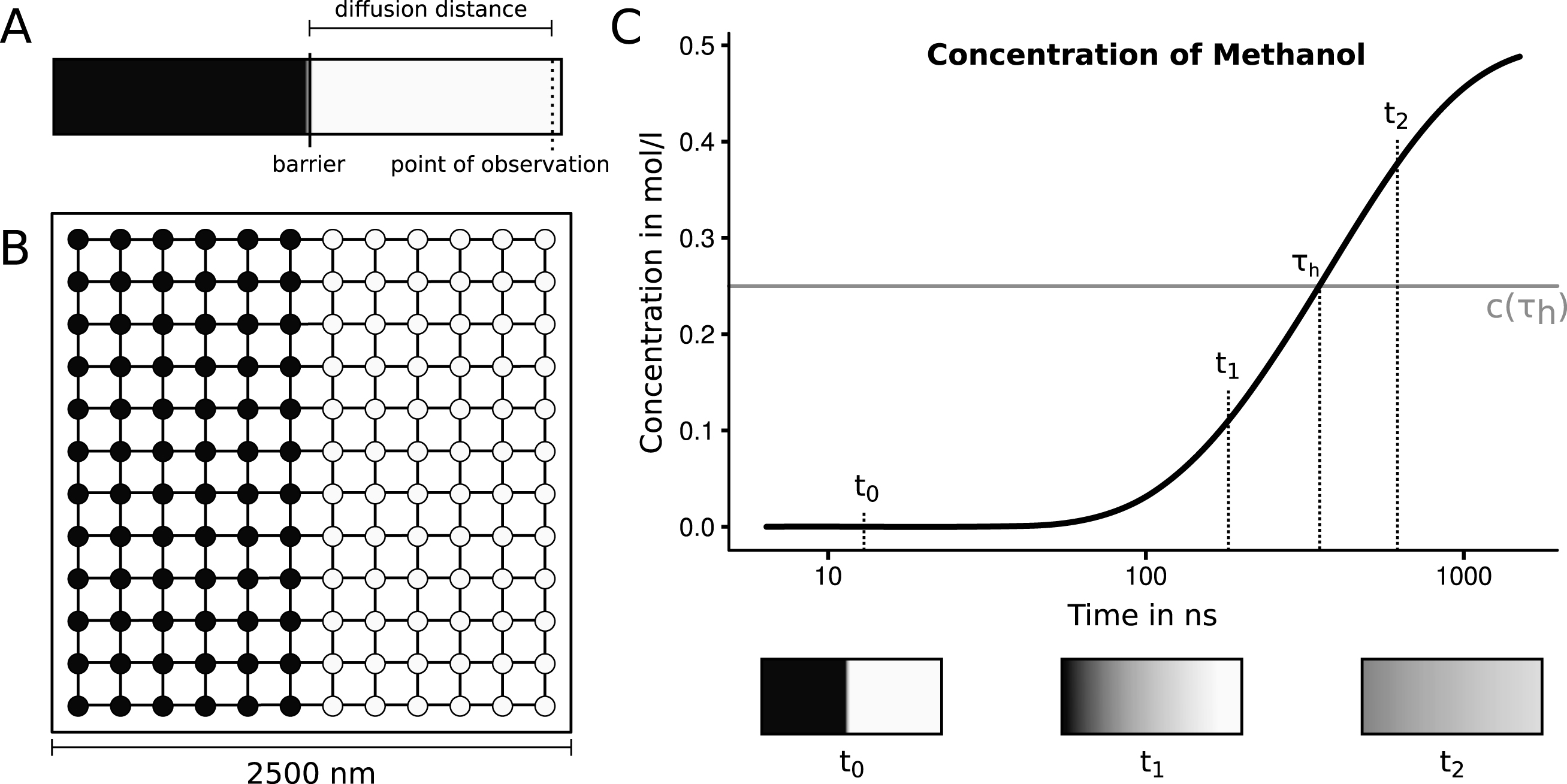
A: The initial setup of the proposed experiment. A tube is filled with two solutions, separated by a central barrier. One half (indicated with light color) contains a high concentration of the proposed substance, the other half does not contain the solved substance. The distance between the barrier and the point of observation is the diffusion distance, for which the half life will be measured. To start the experiment the barrier is removed. Now the time is measured until half of the equilibrium concentration is reached at the point of observation. B: The initial setup of the rectangular graph automaton with twelve nodes side length. The nodes contain concentrations of 1.0 mol·l-1 (indicated with dark color) and 0.0 mol·l-1 (indicated with light color). C: A schematic depiction of the progression of the diffusion process of methanol at the point of observation. The half life is measured when the concentration reaches half of the equilibrium. This concentration of 0.25 mol·l-1 is indicated with a grey line. The coloring shown in the three boxes illustrate the state of the system at three points in time based on the experimental progression.
Comparison of the simulation results
The graph automata simulation has been performed with 5 ns time steps and 40,000 nodes (200 x 200). The values for the diffusivity have been experimentally determined and are presented in [28].
Initially, a numerical approach was used to simulate the diffusion process in a closed system. This computation was performed using Octave [32]. The simulation uses an explicit finite difference method with a first order upwind in time and a second order central difference in space scheme [30, 33]. To specify the square shape of the simulation space, four reflecting Neumann boundaries were set. After specifying the systems boundaries, the kinematic viscosity of the system is set to exhibit the properties of the desired species
Additionally, using the definition presented in this paper, a CGA is designed to simulate the same rectangular system (see Fig. 4B). The number of nodes at either side of the grid

Simulations for six different substances. The number of nodes (side length of the square graph) ranges from 10 to 70 (implicitly changing the node distance Δ
A mathematical concept was developed that allows the simulation of multiple chemical species, subject to natural phenomena described by so-called value rules
After the underlying geometry of the simulation is defined, chemical components can be added to each node of the system (see Fig. 3). Different attributes can be assigned to the chemical components that may be subject to change during the progression of the simulation. The diffusion requires the concentration and the diffusivity of each species. The concentration is node specific and will be changed during the simulation depending on the concentration of the surrounding cells, whereas the diffusivity is species specific and remains constant over time. The two global parameters that determine the scaling of the system are species and node independent. Those are the node distance Δ
The simulation system and model were implemented in order to verify their applicability. We developed the application programming interface
The results of an exemplary system (see Fig. 5) indicate, that the simulation using CGA with an increasing number of nodes and a decreasing time step size converge to the half life calculated with the numerical simulation. However, the graphs also show that the system is unstable when a large number of nodes (equivalent to small node distances Δ
that traces the arches that can be seen in Fig. 5. Here, let
As depicted in the flowchart in Fig. 5 for every time step every node needs to be addressed at least once. The complexity of each time step is
PDEs are an established strategy to simulate a variety of natural systems. In systems that have predictable behaviors and outcomes PDEs flourish to their full potential. In biology however, one often encounters systems that are very complex and influenced by multiple aspects. In these cases PDEs reach the limits of their applicability. Using implicit systems such as CA or CGA, it may be possible to simulate different systems that cannot be tackled efficiently using PDE.
With the proposed definition of CGA we have shown that it is possible to simulate diffusion of different chemical species to a high degree of accuracy. Diffusion is one of the most fundamental processes that many biological systems rely on. The system shown here is able to easily adjust to different time and space scales with little adjustments to the configuration of the system. Using the underlying structure of graphs it is possible to define different topologies. It is possible to add plenty of species to a simulation. Furthermore, the user is able to simulate and observe the system in order to improve the understanding of its current behavior. It is even possible to alter the concentration of a species at a specific node, or the structure of the graph at any point during the simulation, due to the implicit definition. Finally, not much mathematical understanding that is needed to set the system up (see setup process in Appendix A). As shown in Fig. 2, the user has to initialize the graph first, than choose the required species and finally define the systems parameters Δ
Computer simulations are able to act as a bridge between theory and experiment [37]. In the past the gap between theory and experiment for molecular behavior has been overcome by molecular dynamic simulations [38, 39]. However there is still a lot to be done to bring theory and experiment together when it comes to the dynamics of cellular processes. First steps were done by many researchers over the past centuries. Nevertheless, the gap between generalizable simulations and experimental outcome still needs to be closed. We hope that the different fields of science will work together to make a similar approach to “Cell Dynamic Simulations"possible.
In the near future, research will be directed towards the creation of value rules for different chemical reaction types and membrane transport. The setup process for simulations will be further simplified by providing possibilities to extract information from relevant databases and by offering visual feedback using the graphical user interface. Additionally the SiNGA application programming interface (available on GitHub
Author contributions
All authors designed the research. C.L. designed and implemented the model and simulation strategy, and evaluated the performance. C.L. and F.H. wrote the paper.
Footnotes
Appendix A. Simulation setup
A node is neighbored to a maximum of four neighbors, the node distance has been set to 0.9 mm, and the maximal difference in concentration is 0.4 mol·l-1. Now the following three time steps will be considered: 1 s, 50 s and 100 s. To scale the experimentally determined diffusivity
Appendix B. Implementing value rules
The implementation of additional value rules is dependent on their requirement for neighboring values. If the value rule does not require information from neighboring values, its implementation is straight forward. As an example a simple case of chemical kinetics shall be demonstrated:
The general case of a first order reaction with two chemical species
The reaction rate of a first-order reaction
The differentials will be be transformed to discrete changes, to transition to CGA.
Further, the concentration will be rewritten to
Because of the uniformity of the time steps Δ
Therefore the recurrence relation for the new concentration of a species
In this equation
Note that the rate of change is always calculated in dependence of the substrate concentration of the reaction
Acknowledgments
This work was supported by the European Social Fund (ESF) and the Free State of Saxony, Germany. We would like to thank our colleagues, especially Sebastian Bittrich and Florian Kaiser, for thought-provoking impulses and fruitful discussions. Additional we thank Hannah Siewerts for proofreading and feedback on the manuscript.
