Abstract
BACKGROUND:
Epstein-Barr virus (EBV)-associated gastric cancer (EBVaGC) is a common malignant tumor associated with EBV infection. Insulin-like growth factor 2 (IGF2) is an imprinted gene and a key protein that regulates growth, especially during normal fetal development. Loss of imprinting (LOI), is a common epigenetic anomaly in a variety of human cancers. However, the promoter methylation, imprinting status and function of IGF2 gene in GC are unclear.
OBJECTIVE:
To explore the role of IGF2 in the occurrence and development of gastric cancer.
METHODS:
The biological function of IGF2 in gastric cancer was investigated by Transwell, wound healing, CCK-8 and flow cytometry assays. IGF2 imprinting status and gene promoter methylation in gastric cancer tissues were detected by PCR-RFLP and BGS.
RESULTS:
The results showed that the expression of IGF2 was higher in GC tissues than adjacent tissues. IGF2 gene promoter methylation and LOI were significantly higher in EBVaGC tissues than in EBV-negative gastric cancer (EBVnGC) tissues. The high expression of IGF2 in gastric cancer can promote the migration and proliferation of gastric cancer cells.
CONCLUSION:
Our data suggest that IGF2 is involved in the occurrence and development of gastric cancer. Targeting IGF2 may be a potential therapeutic target for gastric cancer.
Keywords
Introduction
Gastric cancer (GC) is often diagnosed at an advanced stage and has a high mortality rate, with 784,000 deaths worldwide in 2018, making it the third most common cause of cancer death worldwide [1]. EBV-associated GC (EBVaGC) is a unique subtype of GC and a common malignancy associated with EBV infection [2]. It has been confirmed that EBV infection induces abnormal methylation of CpG islands and thus causes abnormal changes in host gene expression, which is an important mechanism of EBV involvement in EBVaGC, and also an important mechanism of initiation of malignant transformation of tumor in the early stage of tumor [3]. Therefore, only with a clearer understanding of its tumor biology will it be possible to make substantial progress in the treatment and prevention of gastric cancer [4].
Genomic imprinting, also known as gamete imprinting or parental imprinting, is a special phenomenon that does not follow Mendelian inheritance law and only relies on single parents to pass on some genetic traits. The role of genetic imprinting in human genetic diseases and tumorigenesis has attracted more and more attention. IGF2 is the first confirmed endogenous imprinted gene, and was derived only from the paternal allele [5]. In recent years, with the continuous development of molecular biotechnology, the regulation mechanism of IGF2 imprinting has been further understood, such as enhancer competition theory, chromatin configuration theory, transcriptional cycle theory, etc [6]. IGF2 loss of imprinting (LOI) leads to increased expression of IGF2, and can promote cancer development. For example, IGF2 LOI enhances the pluripotency of colorectal cancer stem cells by promoting tumor autophagy, and IGF2 LOI can promote the development of prostate cancer [7, 8]. Although IGF2 LOI was previously reported to be present in 34.5% of GC tumor tissues [9], the incidence of IGF2 LOI in EBVaGC and its relationship to IGF2 expression have not been studied until now.
In recent years, epigenetic abnormalities, including LOI and DNA methylation, have been identified as important mechanisms in the development of many human cancers. Epigenetic abnormalities of host cells caused by viral infection are the focus of research on oncogenic mechanism of tumor viruses. The main objective of this study was to analyze the promoter methylation and imprinting status of IGF2 in EBVaGC and EBVnGC. And the role of IGF2 gene in the occurrence and development of gastric cancer was elucidated to provide theoretical and experimental basis.
Materials and methods
Cell culture and tissue samples
AGS-EBV, C666-1, Daudi and Raji cell lines were EBV positive cell lines, among which AGS-EBV and C666-1 were donated by Professor Sun Lunquan from Xiangya Medical College, Central South University. SGC7901, HGC-27, BGC823, AGS, MKN-45, CNE and Jurkat cell lines were all EBV negative cell lines, among which SGC7901 and HGC-27 cell lines were preserved in our laboratory. BGC823 and AGS cell lines were donated by Prof. Shao Chunkui, Department of Pathology, the Third Affiliated Hospital of Sun Yat-sen University; MKN-45 cell lines were purchased from Peking Union Cell Bank; CNE cell lines were donated by Prof. Zhu Wei, Department of Pathology, Guangdong Medical College; and Jurkat cell lines were donated by Department of Bioinformatics, Tottori University Health Science Center, Japan. GT38, GT39, Daudi and Raji cell lines (EBV positive) were provided by T. Sairenji (Tottori University, Japan). The SNU719 cell line (EBV positive) was provided by Tao Qian (Chinese University of Hong Kong). MGC803 was provided by Jiang Lei (Peking University). All cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% fetal bovine serum, 1% penicillin-streptomycin at 37
Transfection with IGF2 siRNA and plasmid
Specific siRNA against IGF2 and Negative control (NC) were designed and synthesized by GenePharma (Shanghai, China). IGF2 gene was cloned into pcDNA3.1 containing FLAG tag and has identification reports. The sequences are as follows: siIGF2: 5’-GCUC AGAAAUUGGCUUUAATT-3’, 5’-UUAAAGCCAAU UUCUGAGCTT-3’; Negative control (NC) 5’-UUCUC CGAACGUGUCACGUTT-3’, 5’-ACGUGACACGUU CGGAGAATT-3’. Transfection of 50 nM siRNA was performed using Lipofectamine 2000 Reagent (Invitrogen, Thermo Fisher Scientific, Germany). The cells were gathered after 48 h to assay relative gene expression.
Cell proliferation assay
Cells were seeded at a density of 4
Cell cycle assay
Cells were seeded into six-well plates and incubated overnight. After treatment for 48 h, the cells were digested by trypsin and washed gently with PBS. The collected cells were fixed in 66% ethanol at 4
Cell apoptosis detection
GC cells were seeded in 6-well plates and were transfected with plasmid, siRNA, or NC for 48h cells and then washed with cold PBS. Apoptosis was detected using an Annexin V-FITC Apoptosis Detection Kit (BD Biosciences, USA) according to the manufacturer’s instructions.
RNA extraction and quantitative reverse transcription PCR (qRT-PCR)
Total RNA was extracted from the cultured cells using the TRIzol reagent (BioFlux, Beijing) and then reverse transcribed into cDNA according to the manufacturer’s protocol. We tested the absorbance values of the proposed RNAs, all of which were between A260/A280
Western blot analysis
All cells were lysed using radioimmunoprecipitation assay (RIPA) buffer containing a protease inhibitor, phenylmethanesulfonyl-fluoride (PMSF), and a phosphatase inhibitor mixture. These samples were transferred to polyvinylidene difluoride (PVDF) membranes (Millipore, USA) using SDS-PAGE gel with a 10% mass concentration, blocked with bovine serum albumin (BSA) with a 5% mass concentration at room temperature for 2 h, incubated with the primary antibody overnight at 4
Transwell migration assay
The migration ability of the cells was measured using the transwell inserts with 8
Wound healing assay
The treated cells were inoculated into the six-well plate, and when the cells had grown to fusion, a high pressure toothpick was used to scrape the fused cell layer to form a linear wound. The cells were washed twice to remove shed cells and debris. Then observe and measure the size of the wound at the specified time.
Bisufite genomic sequencing (BGS)
BGS specific primers were used to amplify the promoter region of IGF2 coding gene, and TA cloning was combined with DNA sequencing technology to randomly select amplified bands for sequencing. Six clones were selected by sequencing, and the methylation rate was calculated according to the proportion of CG sites in the total CG sites of the six TA clones. The 46 CG sites in the amplified DNA fragments were drawn as dot plots to visually display the methylation state of IGF2, in which the black circle indicated that the CG site was methylated, and the blank circle indicated that the CG site was not methylated.
PCR-restriction fragment length polymorphism (PCR-RFLP)
Genomic DNA was amplified by polymerase chain reaction (PCR), and Apa I digestion was performed on the amplified products. The forward and reverse specific primers used to amplify exon 9 of IGF2 gene were as follows: 5’-CTTGGACTTTGAGTCAAATTGG-3’ and 5’-GGTCGTGCCAATTACATTTCA-3’. The 25
IGF2 protein expression in GC and normal adjacent tissues of gastric cancer samples
IGF2 protein expression in GC and normal adjacent tissues of gastric cancer samples
GC, gastric cancer; GC tissues including EBV-associated gastric cancer and EBV-negative gastric cancer tissues.
Specimens with DNA genotype AB (heterozygote) were selected for reverse transcription into cDNA. Then PCR was run on cDNA with the same primers used to determine informative status. The RT-PCR products were also digested by Apa I and electrophoresed on 2% agarose gel. Two bands of 160 bp and 38 bp were observed as alleles A, and only 198 bp bands were observed as alleles B, both of which were imprinting maintain. If 198 bp, 160 bp, 38 bp three bands were observed, it was a biallelic expression state (AB type), that is, loss of imprinting.
Immunohistochemistry
Paraffin embedded and fresh gastric tissues were collected from the Affiliated Hospital of Qingdao University and Qingdao Municipal Hospital. EBVaGC tissues were identified by in situ hybridization for EBV-encoded small RNA1 (EBER1). Sections were incubated for 1h at room temperature with anti-IGF2 antibody (abcam, 1:200). Antigen-antibody complexes were visualized using a DAB chromogenic kit (Zsbio, China). The specimens were randomized, coded, and then analysed by two independent pathologists. The study was conducted in accordance with the ethical standards and the principles of the Declaration of Helsinki and was approved by the Ethical Committee of the Medical College of Qingdao University. Written informed consent was obtained from each participant before the start of the study.
Statistical analysis
All data are expressed as the means standard error of the mean (SEM). The experimental data were analyzed using Student’s
Results
Role of insulin-like growth factor 2 (IGF2) in the development of gastric cancer. A. (A) IGF2 positive expression in EBV-associated gastric cancer (EBVaGC) tissues, the arrow indicates the specific cytoplasmic expression of IGF2 in gastric cancer cells; (B) IGF2 negative expression in gastric tumor tissues. B. The transfection efficiency of IGF2 overexpressing plasmids and siIGF2 in AGS cells. C. Effect of IGF2 and siIGF2 on the migration ability of AGS cells as detected by Transwell assay 48 h after transfection (40x). D. Wound healing assays were used to assess cell migration. E. The apoptosis rate was detected by flow cytometry in AGS cells. F. The cell cycle was observed using flow cytometry in AGS cells. G. After transfection for 24 h, 48 h, 72 h and 96 h, 10 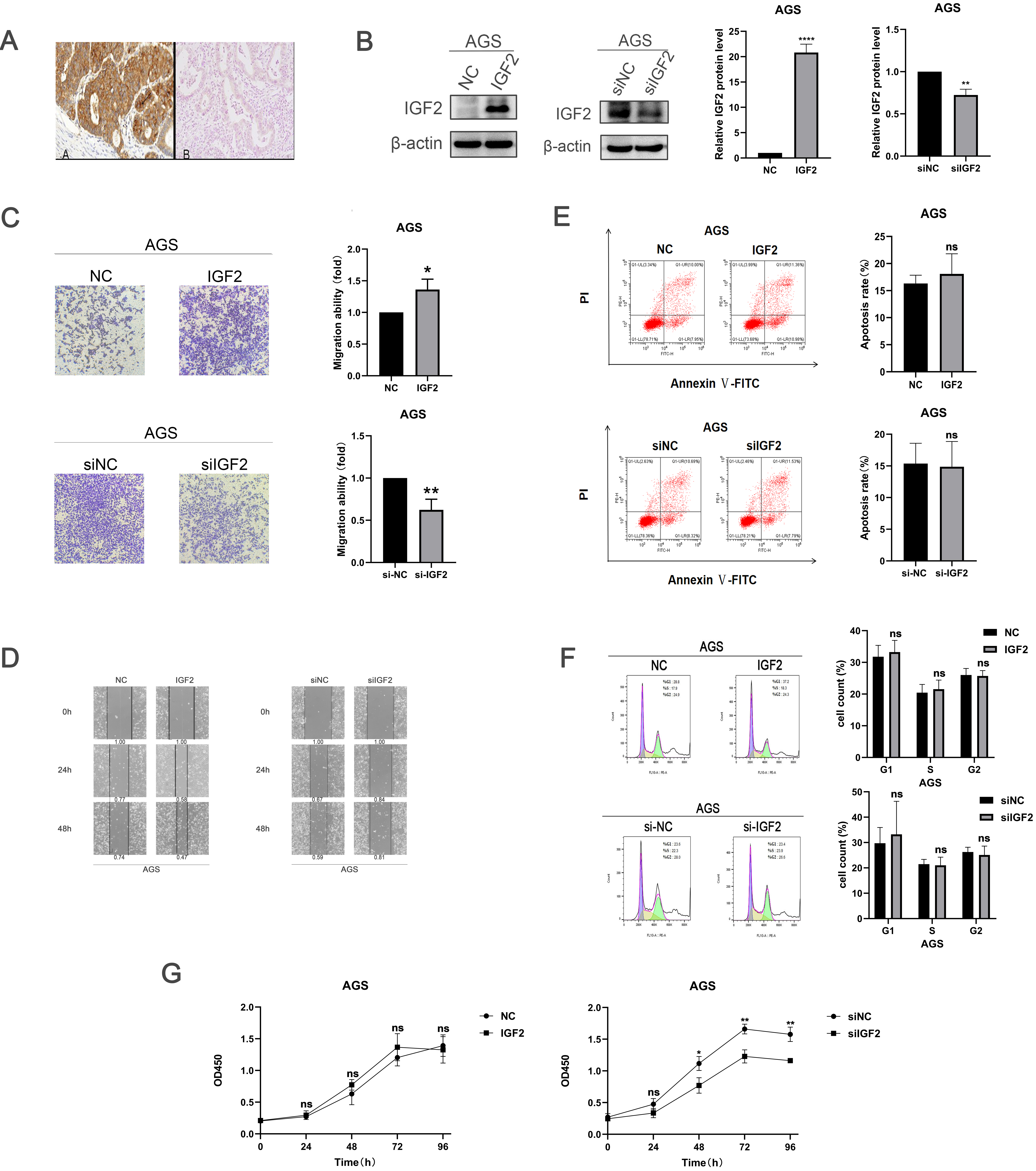
IGF2 protein expression in GC and normal gastric tissues of gastric cancer samples
GC, gastric cancer; GC tissues including EBV-associated gastric cancer and EBV-negative gastric cancer tissues.
Role of insulin-like growth factor 2 (IGF2) in the development of gastric cancer. A. The transfection efficiency of IGF2 overexpressing plasmids in MGC803 cells. B. Effect of IGF2 on the migration ability of MGC803 cells as detected by Transwell assay 48 h after transfection (40x). C. Wound healing assays were used to assess cell migration. D. The apoptosis rate was detected by flow cytometry in MGC803 cells. E. The cell cycle was observed using flow cytometry in MGC803 cells. F. After transfection for 24 h, 48 h, 72 h and 96 h, 10 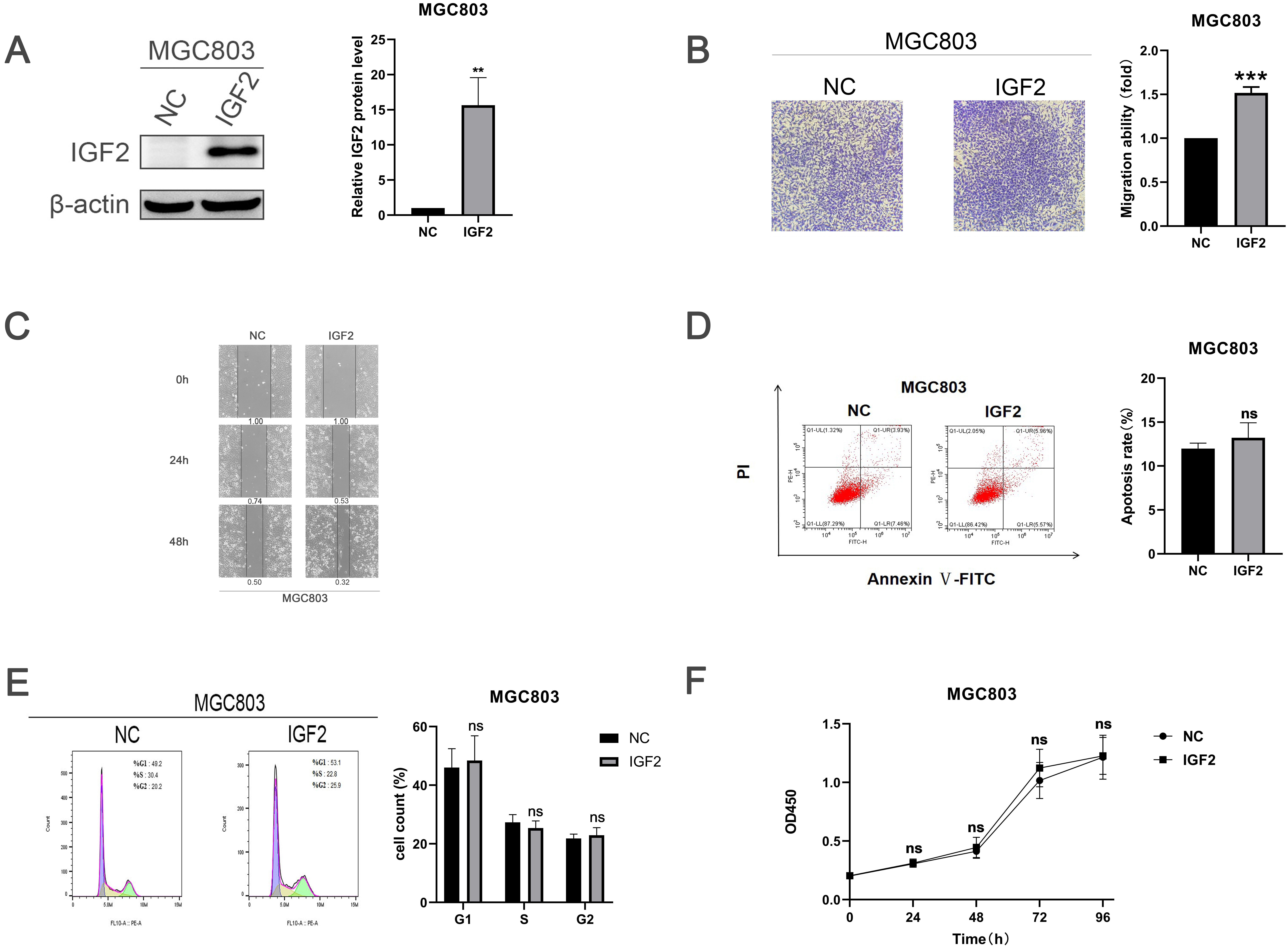
Firstly, in order to understand the role of IGF2 in the development of gastric cancer, the expression of IGF2 in gastric cancer tissues and adjacent normal tissues was detected by immunohistochemistry. We found that the expression of IGF2 in gastric cancer tissues was significantly higher than that in adjacent tissues (Fig. 1A) (Table 1). We further examined the expression of IGF2 in normal gastric tissues and found that it was significantly lower than that in gastric cancer tissues (Table 2). Besides, results have shown that there was no significant difference in the expression of IGF2 protein in EBVaGC and EBVnGC tissues (supplementary material, Table S1).
IGF2 promotes the migration and proliferation of gastric cancer cells
To explore the function of IGF2 in GC cells, we overexpressed IGF2 by transfecting plasmid and suppressed IGF2 expression using siRNA (Fig. 1B). Using Transwell, we found that the overexpression of IGF2 could significantly promote the migration of AGS cells and inhibition of IGF2 could significantly down-regulate the migration ability of AGS cells (Fig. 1C). Wound healing experiments with AGS cells also showed similar results (Fig. 1D). Overexpression and inhibition of IGF2 in AGS cells showed no significant effect on apoptosis and cell cycle by flow cytometry (Fig. 1E and F). Although CCK-8 assays showed that overexpression of IGF2 had no significant effect on cell proliferation, we found that the cell proliferation ability was decreased after interference with IGF2 compared with the control group (Fig. 1G). For further verification, we overexpressed IGF2 in MGC803 cells and obtained similar results as in AGS cells (Fig. 2).
The molecular mechanism of IGF2 regulation. A. IGF2 overexpression upregulates p-ERK and ZEB1 in AGS cells by Western blot. B. Silencing IGF2 down-regulates p-ERK and ZEB1 in AGS cells through Western blot. C. Western blot showed that IGF2 overexpression upregulates p-ERK and ZEB1 in MGC803 cells.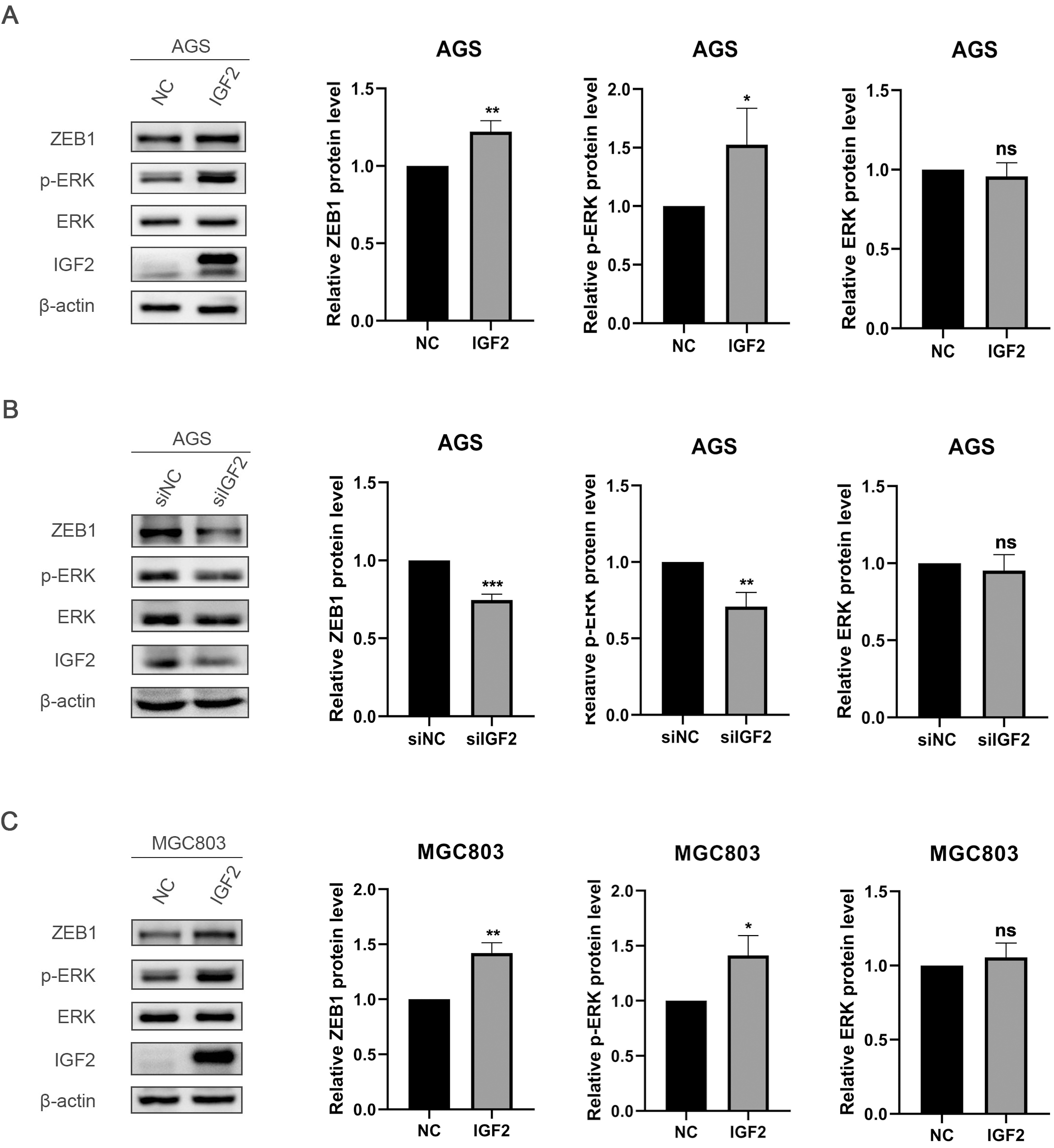
To further explore the molecular mechanism of IGF2 regulation, we overexpressed IGF2 in AGS cells and MGC803 cells and found that overexpression of IGF2 could activate the ERK pathway and up-regulate ZEB1 (zinc finger E-box binding homeobox 1) (Fig. 3A and C). Down-regulation of p-ERK and ZEB1 was found after inhibiting the expression of IGF2 in AGS cell lines (Fig. 3B).
In order to further investigate the higher expression of IGF2 in gastric cancer tissues than in adjacent tissues, we detected the imprinting status and promoter methylation of IGF2.
The subtype of insulin-like growth factor 2 (IGF2) gene, imprinting status and methylation status analysis in gastric cancer tissues. A. The subtype of IGF2 gene in gastric cancer tissues. Lane M: DNA marker DL2000; lane 1, 3: homozygotes (231 bp and 61 bp, both alleles with ApaI sites) and lane 2, 4 homozygotes (292 bp, both alleles without ApaI sites); lane 5 and 6: heterozygous specimens (292 bp, 231 bp and 61 bp). B. IGF2 imprinting status in gastric cancer tissues. Lane M: DNA marker DL2000; lane 1 and 6 (198 bp,168 bp and 30 bp): LOI showed both alleles expression; lane 2–4 (198 bp): normal imprinting status showed only one allele expression; lane 5 (168 bp and 30 bp): normal imprinting status showed only one allele expression. C. The methylation status of IGF2 promoter in EBV-associated gastric cancer (EBVaGC) and EBV-negative gastric cancer (EBVnGC) tissues. 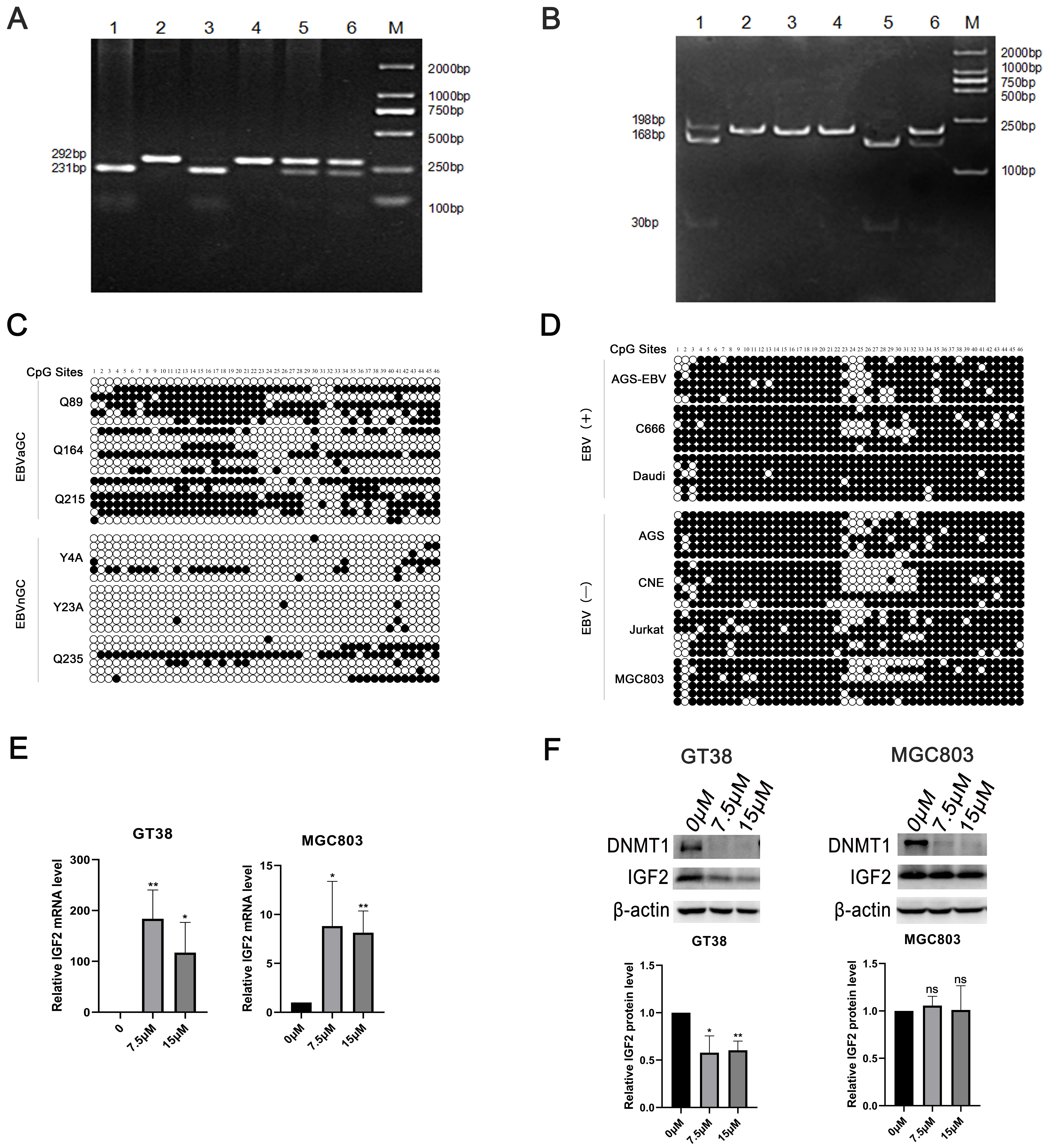
DNA IGF2 gene heterozygous screening in gastric cancer tissue The results of PCR-RFLP showed that among the 80 EBVaGC tissue samples, 40 were heterozygous. Among the 43 EBVaGC adjacent tissue samples, 22 were heterozygous. Among the 92 EBVnGC tissue samples, 56 were heterozygous. Among the 57 EBVnGC adjacent tissue samples, 34 were heterozygous. We can see that there is no significant difference in the proportion of screened heterozygotes in each group. And there was no significant difference in heterozygotes between cancer and adjacent tissues. The classification results are shown in Table 3. The gel imaging results are shown in Fig. 4A. DNA IGF2 gene heterozygous screening in cell lines The subtype of IGF2 in gastric cancer and normal adjacent tissues EBV, Epstein-Barr virus; GC, gastric cancer; EBVaGC, EBV-associated gastric cancer; EBVnGC, EBV-negative gastric cancer.
The subtype of IGF2 in cell lines
EBV, Epstein-Barr virus.
The PCR-RFLP method detected IGF2 genotypes in 5 EBV-positive cell lines and 8 EBV-negative cell lines, and the results showed that the EBV-positive cell lines AGS-EBV, C666-1, OB, Raji, Daudi and EBV-negative cell lines AGS, SGC7901, BGC-823, CNE, Jurkat, HGC-27 and MKN-45 were all homozygous. Only the EBV-negative cell line Ramos is heterozygous. The specific results are shown in Table 4.
IGF2 imprinting status in EBVaGC and EBVnGC tissues
EBV, Epstein-Barr virus; GC, gastric cancer; EBVaGC, EBV-associated gastric cancer; EBVnGC, EBV-negative gastric cancer; LOI, loss of imprinting.
The above-mentioned IGF2 gene heterozygous specimens screened at the DNA level were selected, their RNA was extracted, reverse-transcribed into cDNA, and their imprinting status at the RNA level was detected by PCR-RFLP technology. Among the 40 EBVaGC tissue samples with heterozygous DNA, 31 cases of qRT-PCR amplification were successful. The results of imprinting typing showed that 19 cases were AB biallele expressions, that is, the loss of imprinting. 12 cases were allele type B, that is, the maintain of imprinting. Among the 56 EBVnGC tissue samples with heterozygous DNA, 45 cases of qRT-PCR amplification were successful. The results of imprinting typing showed that 15 cases were AB biallele expression type, that is, the loss of imprinting. 12 cases were allele type A, and 18 cases were allele type B, that is, maintain of imprinting. The results of vertical electrophoresis of some samples are shown in Fig. 4B. Statistical analysis showed that there were significant differences in the distribution of the two imprinting expression states in EBVnGC and EBVaGC (
The methylation ratio of IGF2 promoter in gastric cancer tissues
The methylation ratio of IGF2 promoter in gastric cancer tissues
Footnotes
Conflict of interest
The authors declare no conflict of interest.
