Abstract
BACKGROUND:
Breast cancer is the most worldwide commonly found malignancy among women. The evidence for lipidomic studies of breast cancer in the Chinese population is relatively limited.
OBJECTIVE:
Our current study aimed to identify peripheral lipids capable of distinguishing adults with and without malignant breast cancer in a Chinese population and to explore the potential lipid metabolism pathways implicated in breast cancer.
METHODS:
Lipidomics was performed with an Ultimate 3000 UHPLC system coupled with a Q-Exactive HF MS platform by using the serum of 71 female patients with malignant breast cancer and 92 age-matched (
RESULTS:
A total of 47 significantly different lipids were identified by applying the following criteria: false discovery rate-adjusted P < 0.05, variable importance in projection
CONCLUSIONS:
Using an untargeted LC-MS-based metabolic profiling approach, our study provides preliminary evidence that extensive dysregulations of OxPCs, PCs, SMs and TAGs were involved in the pathological processes of breast cancer. We provided clues for furtherly investigating the role of lipid alterations in the pathoetiology of breast cancer.
Introduction
Breast cancer is the most worldwide commonly found malignancy among women, which is characterized by complex interactions of genomic and physiological at the tumor site [1, 2]. Due to the lack of effective treatments, aggressive breast cancer is still the main cause of cancer-related deaths among females [3]. With the increasing incidence and mortality both in the clinical setting and in population-based screening during recent decades [4], the burden of breast cancer has grown rapidly [5].
Although the underlying molecular mechanisms of breast cancer have been explored by several publications, the etiology is still not completely understood and causal pathways have not been well delineated [6]. In recent years, with the continuous progress of detection technology, the exploration of the molecular mechanism of breast cancer has been further developed. Among them, metabolites or lipids are the final downstream product of metabolic reactions, which represent the level of small molecules (metabolites and lipids, less than 1200 Da) [7]. Lipidomic analyses, as the endpoint of the ‘omics’ cascade, could provide more global lipid status of a biological specimen compared to the genome, transcriptome, or proteome [8]. Recent evidence showed that abnormalities of lipids may influence the cellular processes, which could result in tumor development [9, 10]. Several studies demonstrated that the biomolecular processes in patients with cancers differ from normal individuals and a variety of cancers share common metabolic features. Thus, deregulated energy metabolism is considered one of the established hallmarks of cancer [11].
Several targeted lipid analyses have explored the association between specific lipids and breast cancer. For example, a genetic study reported that 16 lipid metabolism-related genes were associated with the process of lipid metabolism and fatty acid binding in breast cancer [12]. A vitro experimental study found that lipids palmitic acid (PA), ceramide (Cer), and docosahexaenoic acid (DHA) have anti-tumor efficacies and reduced the proliferative, migratory and invasive properties of breast cancer cells, which indicated that lipid metabolism targeting drugs have anti-tumor potential targeting breast cancer [13, 14]. Meanwhile, several untargeted lipidomic approaches have explored lipid profiles in patients with breast cancer [6, 10]. Previous untargeted lipidomic studies have found that many different lipids are in breast cancer and controls, and the differences between different breast cancer subtypes [15, 16, 17].
However, due to the diversity of detection instruments and technical methods, the results of different studies lack comparability. The evidence for lipidomic studies of breast cancer in the Chinese population is relatively limited. In this current study, an Ultimate 3000 UHPLC system coupled with a Q-Exactive HF MS platform, in conjunction with multivariate statistical analysis, was used to screen and determine potential serum lipidome among patients with breast cancer and compare it with the lipidome of healthy controls. Herein, we aimed to identify peripheral lipids capable of distinguishing adults with and without breast cancer in a Chinese population and to explore the potential lipid metabolism pathways implicated in breast cancer.
Methods
Ethical approval
The research was reviewed and approved by the Peking University Third Hospital in Beijing, China (Approval no. IRB-2018-413-02) and has been performed with the ethical standards as laid down in the 1989 Declaration of Helsinki and its later amendments or comparable ethical standards. All the participants were fully informed of the objectives of the study and signed the informed consent to before their inclusion.
Study population
The clinical data and serum samples of 71 female patients who were histopathological diagnoses of malignant breast cancer and were treated at Peking University Third Hospital in Beijing, China were retrospectively collected. The histopathological diagnoses of the malignant groups were based on ultrasound-guided breast biopsies by trained breast radiologists. The cell morphology was used to determine the malignant and benign cancers. The inclusion criteria of patients were as follows: (I) female breast cancer patients undergoing surgery and chemotherapy in Peking University Third Hospital; (II) aged 18–60 years; (III) with complete clinicopathologic data. At the same time, the clinical data and serum samples of 92 age-matched (
Serum samples preparation and lipid extraction
Fasting blood samples were collected in serum separation tubes at the time of pre-puncture coagulation function and pre-operative immune tests from the forearm vein. Serum samples were separated by centrifugation at 3000
LC-MS data acquisition
Global metabolomic analysis was performed on an Ultimate 3000 UHPLC system coupled with Q-Exactive HF MS (Thermo Scientific). Chromatographic separation was performed on a phase reversed-phase X-select CSH C18 column (2.1 mm
Raw data collected from the DDA-MS were processed on MS-DIAL software (
Statistical analysis
Metabolic analysis of serum samples from patients with breast cancer and healthy controls. (a) Overview of PLS-DA top 4 components. (b) 2D PLS-DA scores plot of top 2 components. Notes: The values of the abscissa and ordinate reflect the variability of the sample in different principal component directions, while the positive and negative signs represent the positional relationship of the sample in the principal component direction. The explained variances of each component are shown in the cells on the diagonal of Fig. 1(a) and brackets in Fig. 1(b).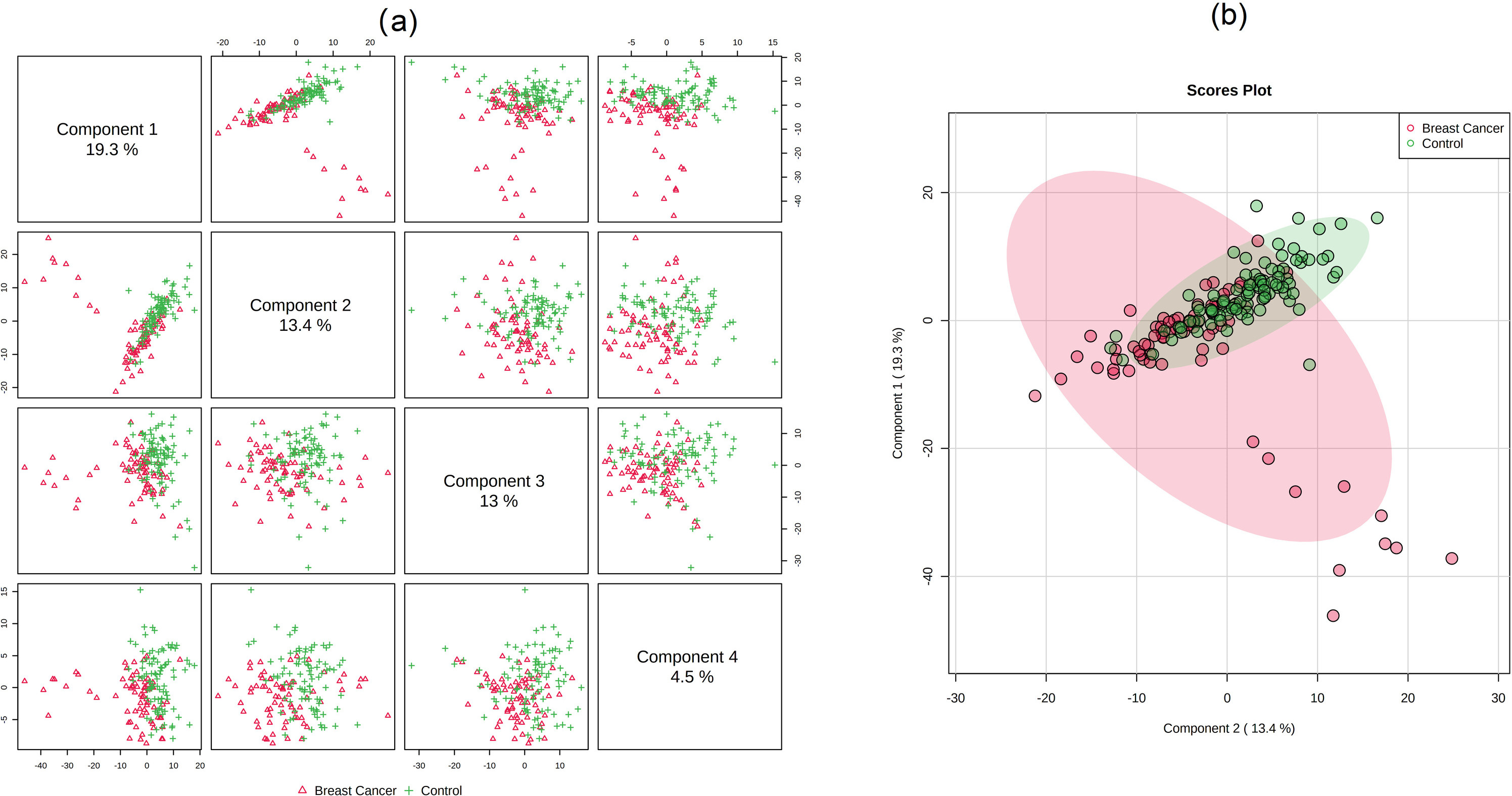
The data were uploaded to and processed by the specialized online software Metaboanalyst 5.0 (
Demographic and clinical characteristics of study subjects
Basic information of the included participants
Basic information of the included participants
ROC analysis revealed candidate lipid biomarkers for breast cancer diagnosis. (a), (b) and (c) ROC curves of PC 56:15, DGCC 50:14, and DGDG 12:0-20:1. The left side of each figure represents the distribution of the ROC curve for differentiating the breast cancer group from the health control group, and the right side is the difference between the features of the two groups. (d) Multivariate ROC curve constructed with 2–47 lipids based on the cross-validation (CV) performance. Each curve represents the potential of top 2, top 3, top 5, top 10, top 20, and total 47 features in differentiating the breast cancer group from the health control group. (e) The predicted class probabilities (average of the cross-validation) for each sample using the 5 features model of lipids. (f) The corresponding predictive accuracy of each PLS-DA model is constructed with different numbers of features. The predictive accuracy of 2 to 47 features is from 65.1% to 83.5%, respectively.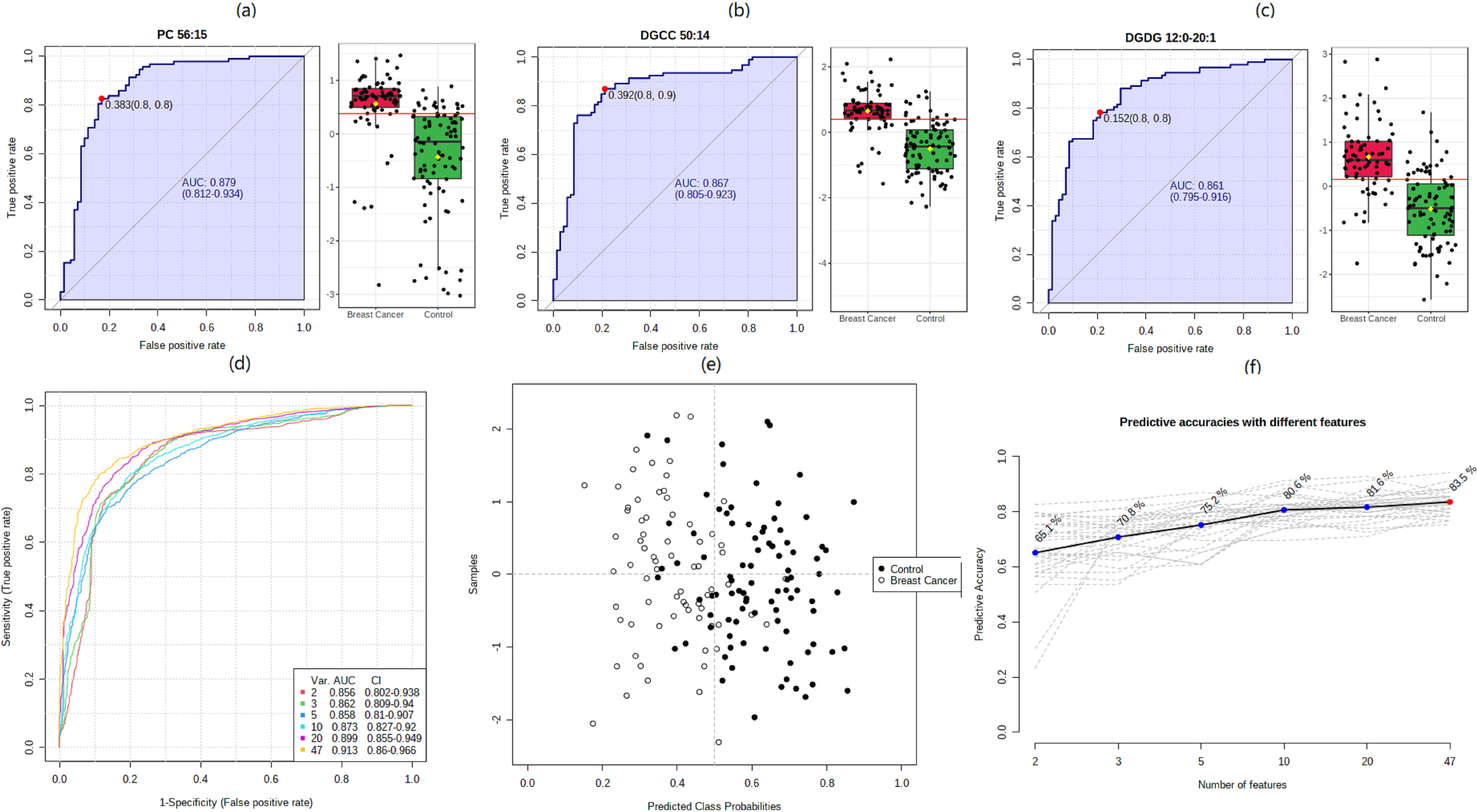
Differentiating lipids between breast cancer and healthy control groups identified from the lipidomic data
VIP, variable importance in the projection; FC, fold change.
A total of 71 patients with breast cancer and 92 healthy controls were included in the data analysis. No significant differences in menopause status, smoker, drinker, and BMI were observed between the two groups (
There were a total of 1122 lipids detected in positive and negative modes after peak deconvolution, alignment, and picking. We conducted univariate analysis after pre-processing the obtained LC-MS raw data. The independent-sample
We combined PCA and OPLS-DA to get a comprehensive view of identifying differences in lipidomic data between patients with breast cancer and healthy controls. According to our PCA scatter plot, the two comparison groups were separated. QC samples were gathered well, proving the reliability of the lipidomic data (Supplemental Fig. 1) The supervised OPLS-DA was applied to obtain a higher level of group separation and a better understanding of the variables responsible for classification. The results of OPLS-DA are shown in Fig. 1. And 470 lipids were identified to distinguish the two comparisons with VIP
In addition, we also conducted fold change analysis, 62 lipids were reported to be different with fold change
ROC analysis revealed candidate lipid biomarkers for breast cancer
To estimate the diagnostic value of the dysregulated lipids, ROC curves and binary logistic regression models were used to screen markers by examining the AUCs of the biomarkers in the breast cancer group and healthy control group (Fig. 2) According to the works of literature, the biomarkers can be employed as diagnostic biomarkers if the AUC is greater than 0.7. Our ROC analysis revealed 13 lipids with an area under the curve (AUC) greater than 0.7. Multivariate ROC curves constructed with 247 lipids are shown in Fig. 2d, AUCs of greater than 0.8 could be achieved, and the corresponding predictive accuracy of each PLS-DA model constructed with different numbers of features are shown in Fig. 2e and f.
Discussions
Using an untargeted LC-MS-based metabolic profiling approach, our study explored the distribution of lipids in patients with breast cancer, and several lipids, including but not limited to PCs, OxPCs, SMs, and TAGs, were found dysregulated between breast cancer and healthy controls, which were considered as potential abnormal biomarkers of the disease.
The most differential lipids reported in our study were PCs. Sixteen PCs were reported dysregulated eleven were up- and 5 were down-regulated in patients with breast cancer. Univariate ROC analysis indicated that the AUC of PC 56:15 was 0.879, which can be considered one of the good diagnostic biomarkers The association between PCs and breast cancer has been reported previously [6, 18]. A targeted lipidomics analysis reported that the total concentration of PCs and PEs of the patient with breast cancer was higher than controls but they decreased significantly following surgery [19] Similar to our finding, Buentzel et al.’s metabolomic profiling study also found several PCs, such as PC ae C40:6, PC aa C38:5, PC ae C34:2, PC ae C32:2, and PC ae C38:3, has good classification potential between breast cancer and healthy controls [20]. Moreover, another study reported that PC 32:0, PC 34:1, and PC 36:2 were overexpressed in HER2-positive breast cancer compared to adjacent normal tissue [21]. The distribution of PCs in breast cancer patients with different pathological characteristics deserves further investigation.
Various OxPC molecules are formed by the oxidation of PC molecules at unsaturated sites and have different biological activities from the parent molecules [22]. Recent evidence suggested that OxPC molecules can be considered end-product markers of oxidative stress and a novel class of bioactive lipids in human pathophysiology [23, 24, 25]. Our current study identified 9 OxPCs were significantly higher in patients with breast cancer than in healthy controls. However, we did not find much evidence to explain our current findings. A recently published study demonstrated that OxPCs could induce autophagic flux in cancer cells and increase the metastatic potential of cancer cells [26]. An OxPC named 1-palmitoyl-2-(5-oxovaleroyl)-sn-glycerol-3-phosphocholine (POVPC) was found to increase the migration and invasion of human hepatocellular carcinoma cells (HepG2) and human breast cancer cells (MCF7) by inducing epithelial-mesenchymal transition (EMT) [26]. Moreover, increased OxPCs were also reported in other diseases, such as neurodegeneration [27]. Thus, the abnormal alteration and potential pathways of PCs and OxPCs in breast cancer are worthy of further exploration.
It has been reported that SMs are the most abundant sphingolipid and important constituents of lipid rafts in normal cells [28], they are involved in the regulation of numerous signaling pathways [29]. There are conflicting reports on the association between SMs and malignancy. Reduced levels of SMs were observed in colon cancer tissues, A549 and U118 cancer cell lines [30, 31]. Consistent with our findings, Nagahashi and colleagues reported higher levels in the human breast cancer tissue compared to the normal breast tissue [32]. Bhadwal et al. did not find significant alteration of SM (18:2/22:0), SM (40:1) and SM (18:0/24:2) in breast tumor tissues [33]. The consistent increase of the six SMs found in our breast cancer samples is complementary to previous studies. In addition, based on the important roles of SMs in various signaling pathways, our findings are also worthy of further targeted validation
The abnormal changes in glycerolipids were also reported in breast cancer. Among them, 9 TAGs and 2 DAGs showed inconsistently significant changes. TAG is a necessary energetic substrate for cell membrane formation. DAG is a precursor to TAG [34]. Both of them are crucial to the metabolism of structural and signaling lipids [35]. Similar to the inconsistent alterations in our study, the changes of TAG in breast cancer also did not consistently in previous publications [36, 37, 38]. Interestingly, Dai et. al. found that triple-negative breast cancer patients with a high TAG level had shorter overall survival and disease-free survival [39]. Mika and colleagues found that colorectal cancer tissues had significantly lower TAG content than normal tissue, and attributed it to the result of enhanced oxidation [40]. Therefore, future research can focus on investigating the effect of different TAGs on the prognosis of breast cancer, potential pathway and mechanisms. Additionally, our lipidomics study also reported the difference between DGCC 50:14 and DGDG 12:0-20:1 between breast cancer and healthy controls, and both of them had good classification performance for the two groups. DGDG has been pointed out to have anti-inflammatory and cancer benefits [41]. However, the previous evidence is too limited, and our current findings of non-targeted lipidomics need to be furtherly verified.
Limitations
Some limitations should be noted in the present study. Firstly, this study is based on a single-institutional database, there may be selection bias in the study subjects. Secondly, due to the limited evidence, some of our findings have not found a reasonable explanation The detailed mechanisms and the biological significance of lipids in patients with breast cancer require further theoretical and experimental exploration. Thirdly, untargeted metabolomics does not enable the quantification of lipids. The putative lipid biomarkers identified in this study should be further validated using targeted metabolomics methods and verified in larger prospective studies.
Conclusions
In summary, using an untargeted LC-MS-based metabolic profiling approach, the present study provides preliminary evidence that extensive dysregulation of OxPCs, PCs, SMs and TAGs was involved in the pathological processes of breast cancer. Our current findings provided clues for furtherly investigating the role of lipid alterations in the pathoetiology of breast cancer, as well as several potential biomarker panels for breast cancer diagnosis or prognosis in clinical practice.
Funding
This work was sponsored by the National Natural Science Foundation of China (No. 81701067). The supporting foundation had no involvement in study design, the collection, analysis, or interpretation of data, writing of the report; or in the decision to submit the article for publication.
Availability of data and material
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Author contributions
Conception: Li N and Yan LL.
Interpretation or analysis of data: Cao B, and Yang SY.
Preparation of the manuscript: Cao B, Yan LL, and Li N.
Revision for important intellectual content: Yan LL and Yang SY.
Supervision: Li N and Yan LL.
Ethics approval
The study protocol was reviewed and approved by the research ethics committee of the Peking University Third Hospital in Beijing, China (IRB-2018-413-02).
Consent to participate
All subjects enrolled in this study were from this hospital and were provided written informed consent before their inclusion.
Consent for publication
All authors agreed to the publication of this manuscript.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-220462.
sj-docx-1-cbm-10.3233_CBM-220462.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-220462.docx
Footnotes
Conflict of interest
The authors declare that there is no conflict of interest.
