Abstract
Introduction
Head and neck squamous cell carcinoma (HNSCC) is a malignant tumor originating from the mucous epithelium of the oral cavity, pharynx, and larynx.1–3 It is the most common pathological type of head and neck tumor, accounting for 90% of head and neck malignant neoplasms. 2 Most of the early symptoms are not obvious and are easily overlooked by patients; thus, HNSCC can develop into serious complications that affect speech, breathing, and swallowing. 1 The occurrence of HNSCC is mainly associated with alcoholism and excessive smoking, and the occurrence of oropharyngeal cancer, especially tonsil cancer, is associated with high-risk HPV infection.1,4 The occurrence of nasopharyngeal carcinoma is also associated with Epstein-Barr virus infection. 3 Treatment options for head and neck cancer include surgery, chemoradiotherapy, targeted drug therapy, and immunotherapy, and most patients with HNSCC require multimodal therapy. 1 In terms of targeted therapy, anti-EGFR cetuximab can improve the survival rate of patients with recurrence or metastasis. However, besides the anti-EGFR drug, there is currently no clearly effective targeted drug treatment available. 5
Ubiquitin-specific peptidases (USPs) encode members of the ubiquitin-specific protease family, which is the largest family of deubiquitinating enzymes. 6 In addition to being a member of this family of more than 50 USPs, USP39 is also an important component of the spliceosomal complex 25S U4/U6.U5 tri-snRNP, which plays an important role in RNA splicing.7,8 In recent years, carcinogenic effects of USP39 have been reported in many cancer types. It has been reported that USP39 is downregulated in lung cancer. USP39 directly deubiquitinates checkpoint kinase 2 (CHK2) to stabilize its expression, while the knockdown of USP39 significantly reduces CHK2 expression. This downregulation of CHK2 expression leads to decreased apoptosis in lung cancer cells and renders the cancer cells resistant to radiotherapy and chemotherapy. 9 USP39 was also reported to be highly expressed in hepatocellular carcinoma (HCC) tissues and was found to be associated with poor prognosis. Furthermore, USP39 was shown to facilitate the progression of HCC by inhibiting the degradation of ZEB1 (a zinc-finger E-box binding homeobox 1) through deubiquitination. 10 Another study has indicated that USP39 was significantly overexpressed in HCC tumor tissues compared to adjacent normal tissues and promoted tumor growth in HCC patients by effectively splicing FoxM1. 11 According to the research findings, USP39 protein exhibited a markedly elevated expression level in both cervical squamous cell carcinoma, in stark contrast to its expression in normal tissues. 12 Another study showed that the splicing factor USP39 was highly expressed in high-grade serous ovarian carcinoma, and promoted ovarian cancer by increasing the splicing efficiency of high mobility group A2. 13 Previous studies have shown that USP39 is highly expressed in HNSCC, but the underlying mechanism of USP39 in HNSCC remains unclear. 14
The signal transducer and activator of transcription (STAT) is a signal transducer molecule and a transcription activator factor. 15 STAT1 is 1 of 7 STAT family members (STAT1, STAT2, STAT3, STAT4, STAT5a, STAT5b, STAT6). 16 In cancer cells, STAT protein transduces signals of abnormal expression of various cytokines and growth factors in tumor cells to subsequently regulate the expression of many genes at the transcriptional level. 17 STAT1 generally acts as a tumor suppressor, but it can also promote tumorigenesis. For example, high expression of STAT1 can inhibit the proliferation and stem cell characteristics of paclitaxel-resistant ovarian cancer cells and improve the overall survival of patients with epithelial ovarian cancer. 18 Anderson et al. 19 showed a role for STAT1 in promoting proliferation and inhibiting apoptosis in HNSCC. STAT1 expression is elevated in malignant pleural mesothelioma and may serve as a target to restore chemotherapy sensitivity. 20
In the present study, we demonstrated that USP39 knockdown inhibited the viability and migration of head and neck squamous cell carcinoma cells. USP39 might regulate STAT1 in such progress. These findings indicate that USP39 is required to regulate HNSCC in proliferation and migration. Therefore, targeting USP39 is a promising therapeutic approach to inhibit head and neck cancer progression.
Materials and Methods
Human HNSCC Tumor Samples
The primary tumors and their adjacent nontumor samples were obtained from patients with HNSCC at Lu’an Hospital Affiliated with Anhui Medical University (Lu’an City, China). Each patient underwent surgical resection without preoperative systemic chemotherapy. Written informed consent was obtained from each participating patient before the specimens were collected. The collected tissues were promptly stored at −80°C in a freezer for subsequent experiments. This retrospective study was conducted in accordance with the requirements of the Ethics Committee of Lu’an Hospital Affiliated with Anhui Medical University (Lu’an City, China, Approval No: 2020LL003). The procedures were performed consistent with the Declaration of Helsinki and Good Clinical Practice.
Liquid Chromatography-Tandem Mass Spectrometry
To identify differentially expressed proteins (DEPs) between the groups, a liquid chromatography-tandem mass spectrometry (LC-MS/MS) system was utilized. 21 Tissue protein was extracted with the mammalian total protein extraction kit (#AP0601-50, Bangfei Bioscience &Technology Co., Ltd, China). The labeled sample was mixed with 40 µL of a 0.1% formic acid aqueous solution. The digested polypeptides in the mixture were then separated using a high-performance liquid chromatography system (Ultimate 3000, Thermo Fisher Scientific, USA). The mobile phase consisted of phase A, which was 0.1% formic acid in water, and phase B, which was 0.1% formic acid in acetonitrile. The B phase gradient parameters were set as follows: 0 to 50 min, increased to 26%; 50 to 70 min, increased to 38%; 70 to 71 min, increased to 100%; and 71 to 78 min, remained at 100%. Subsequently, the peptides were detected using an Orbitrap Fusion Lumos Mass Spectrometer (Thermo Fisher Scientific, USA).
Proteome Discoverer software (Version 2.4, Thermo Fisher Scientific, USA) was used to analyze the data generated from LC-MS/MS analyses. Peptide identification was carried out by comparing the data against a human proteomic database that includes sequences from UniProt (https://www.uniprot.org/).
Gene Ontology and Kyoto Encyclopedia of Genes and Genomes Enrichment Analyses
The GO database is a biological database used for storing, organizing, and disseminating annotation information about genes and proteins.22,23 We acquired the ontologies by utilizing DEPs from the Database for Annotation, Visualization and Integrated Discovery (DAVID) through the DAVID web interface (https://david.ncifcrf.gov/, accessed on August 18, 2023).24,25 We specifically employed the Functional Annotation function, focusing exclusively on biological processes, cellular components, and molecular function ontologies. Each ontology displays the top 10 terms based on protein counts.
Kyoto encyclopedia of genes and genomes (KEGG) is a comprehensive bioinformatics database that provides rich gene and protein annotation data along with associated information on signaling pathways.26,27 KEGG analysis and visualization were conducted using the R package clusterProfiler (Version 3.16). 28
Immunohistochemistry
Immunohistochemistry was performed using frozen sections. 29 Frozen tissues were cut into 5- to 8-μm slices. After being cryosectioned, the tissue sections were placed in 4% paraformaldehyde, and antigen retrieval was carried out in citrate buffer. Subsequently, sections were incubated with 3% hydrogen peroxide (Catalog No. 7722-84-1, Sigma-Aldrich, Germany) to block endogenous peroxidase activity and nonspecific antigens. Primary anti-USP39 antibody (dilution of 1:100; Catalog No. DF12180, Affinity Biosciences, USA) or anti-STAT1 antibody (1:100; Catalog No. AF6300, Affinity Biosciences, USA) was added, and sections were incubated overnight at 4 °C. After being washed with phosphate-buffered saline (PBS), sections were incubated with an antirabbit secondary antibody for 1 h at room temperature and subsequently incubated with horseradish peroxidase. Then, sections were developed using 3,3′-diaminobenzidine and counterstained with hematoxylin.
Cell Culture and siRNA Transfection
Two commonly used cell lines for studying the proliferation and migration of HNSCC, namely SCC25 and CAL27, 30 were obtained from the American Type Culture Collection (Catalog No. CRL-1628 and CRL-2095, ATCC, USA) and cultured in cell culture dishes using Dulbecco's Modified Eagle Medium (DMEM) (Catalog No. 12491015, Thermo Fisher Scientific, USA) supplemented with 10% fetal bovine serum (Catalog No. A5669401, Thermo Fisher Scientific, USA) and 100 U/mL penicillin/streptomycin. All cells were maintained in a humidified incubator at 37°C with 5% CO2. Lipofectamine 3000 (Catalog No. L3000075, Thermo Fisher Scientific, USA) reagent was obtained from Invitrogen. Specific small interfering RNA (siRNA) targeting human USP39 (5′-GAGAAGGAAUAUAAGACUUTT-3′, 5′-AAGUCUUAUAUUCCUUCUCTT-3′), STAT1 (5′-CAGCAUAACAUAAGGAAAATT-3′, 5′-UUUUCCUUAUGUUAUGCUGTT-3′), and scrambled siRNAs (5′-ACGCGUAACGCGGGAAUUU-3′, 5′-UUCUCCGAACGUGUCACGUTT-3′) were designed and synthesized by Sangon Biotech Co. Ltd, China.
Quantitative Real-Time PCR
Total RNAs were extracted following the manufacturer's instructions using the RNA Extraction Kit (Catalog No. 9767, TaKaRa Bio, Japan). The extracted RNA was then reverse-transcribed using the PrimeScript RT reagent kit (Catalog No. RR047Q, TaKaRa Bio, Japan). We conducted quantitative real-time polymerase chain reaction (qPCR) using a Bio-Rad CFX96 system (Bio-rad, USA) with TB Green Premix Ex Taq II (Catalog No. RR820A, TaKaRa Bio, Japan). The mRNA levels of each gene were normalized to β-actin. The primers used for qPCR were as follows: sense primer for β-actin: 5′-ATCCACGAAACTACCTTCAACTCCAT-3′, antisense primer for β-actin: 5′-CATACTCCTGCTTGCTGATCCACATC-3′; sense primer for USP39: 5′-GGTTTGAAGTCTCACGCCTAC-3′, antisense primer for USP39: 5′-GGCAGTAAAACTTGAGGGTGT-3′; and sense primer for STAT1: 5′-CAGCTTGACTCA AAATTCCTGGA-3′, antisense primer for STAT1: 5′-TGAAGATTACGCTTGCTTTTCCT-3′. The primers for qPCR were obtained from Sangon Biotech Co. Ltd, China.
Cell Counting Kit-8 Assay
Cell viability was determined by cell counting kit-8 (CCK-8) assay. Cells were seeded in a 96-well plate, and 20 µl of CCK-8 reagent (#C0037, Beyotime, China) was added to each well. The absorbance value at a wavelength of 450 nm was measured using an enzyme microplate reader after 48 h. 31
Wound Healing Assay
HNSCC cells were seeded in a 6-well plate and incubated in a 5% CO2 incubator at 37 °C until they reached confluence. One day before the experiment, the cells were rinsed with PBS and switched to a low-serum medium (0.1% serum in DMEM). To create a wound, a straight scratch was made using a 200 μL pipette tip. The pipette tip was held at an angle of less than approximately 30° to control the width of the wound and allow both edges to be imaged together. Afterward, the HNSCC cells were rinsed with PBS to remove any floating cells and then cultured in a normal medium supplemented with 10% fetal bovine serum. Images were captured at the same position in each well using an inverted microscope (TE300, Nikon, Japan) at 0 h and 24 h after the wound was created while maintaining a temperature of 37 °C. The percent wound closure was calculated by measuring the difference in wound area between the initial time point and the indicated final time points.
Statistical Analysis
GraphPad Prism software (Version 9.0) was used for the analysis of statistical significance and data visualization. All numerical data were presented as means ± SEM from at least three independent experiments. Comparisons between groups were analyzed with Student's t-test. Effect sizes and confidence interval calculations were performed using the R package MBESS (Version 4.8.1). A two-sided P ≥ .05 was considered not statistically significant.
Results
DEPs Are Enriched in the Spliceosome Pathway
We collected HNSCC tumor and adjacent tissues from 4 patients (identification numbers 1-4) with hypopharyngeal carcinoma for LC-MS/MS identification of DEPs. Patient characteristics are shown in Table 1. A total of 590 proteins were differentially expressed in tumor tissues versus adjacent tissues (Figure 1). To gain a deeper understanding of the function of DEPs, we performed the GO and KEGG enrichment analysis. The GO functional annotations of the DEPs were categorized into 3 components, biological process, cellular component, and molecular function (Figure 2A-C). Among the enriched KEGG pathways, the number of DEPs enriched in the spliceosome pathway was highest (Figure 2D). The DEPs enriched in the spliceosome pathway are shown in Table 2.

Protein profile of head and neck squamous cell carcinoma (HNSCC) and adjacent tissues. (A) Heat map showing the clusters of the differentially expressed proteins (DEPs) identified by liquid chromatography-tandem mass spectrometry between HNSCC tumor tissue and adjacent tissues. The red blocks represent upregulated proteins. The green blocks represent downregulated proteins. H1 to H4 represent HNSCC tumor tissues, and C1-C4 represent adjacent tissues from patients identified as 1 to 4. (B) Volcano plot of DEPs. Red dots represent upregulated proteins, and green dots represent downregulated proteins. Black dots represent proteins with no significant difference in expression.

Gene ontology (GO) and Kyoto encyclopedia of genes and genomes (KEGG) enrichment analysis of proteins differentially expressed between head and neck squamous cell carcinoma tumor tissue and adjacent tissue. GO term enrichment of the proteins in the categories of (A) biological process, (B) molecular function, and (C) cellular component. (D) KEGG enrichment map of differentially expressed proteins (DEPs). The rich factor is calculated as the ratio of the number of differentially expressed genes annotated in a specific pathway term to the total number of genes annotated in that pathway term. The higher the rich factor, the more significant the pathway enrichment. The larger the size of the dot, the greater the number of DEPs enriched in that pathway. The red dot represents significant enrichment of differentially expressed genes in this GO term and the blue dot represents less significant enrichment of differentially expressed genes in this GO term.
Clinical Diagnoses and Demographic Characteristics of Participants.

Thirty-Four Differentially Expressed Splicing Factors.
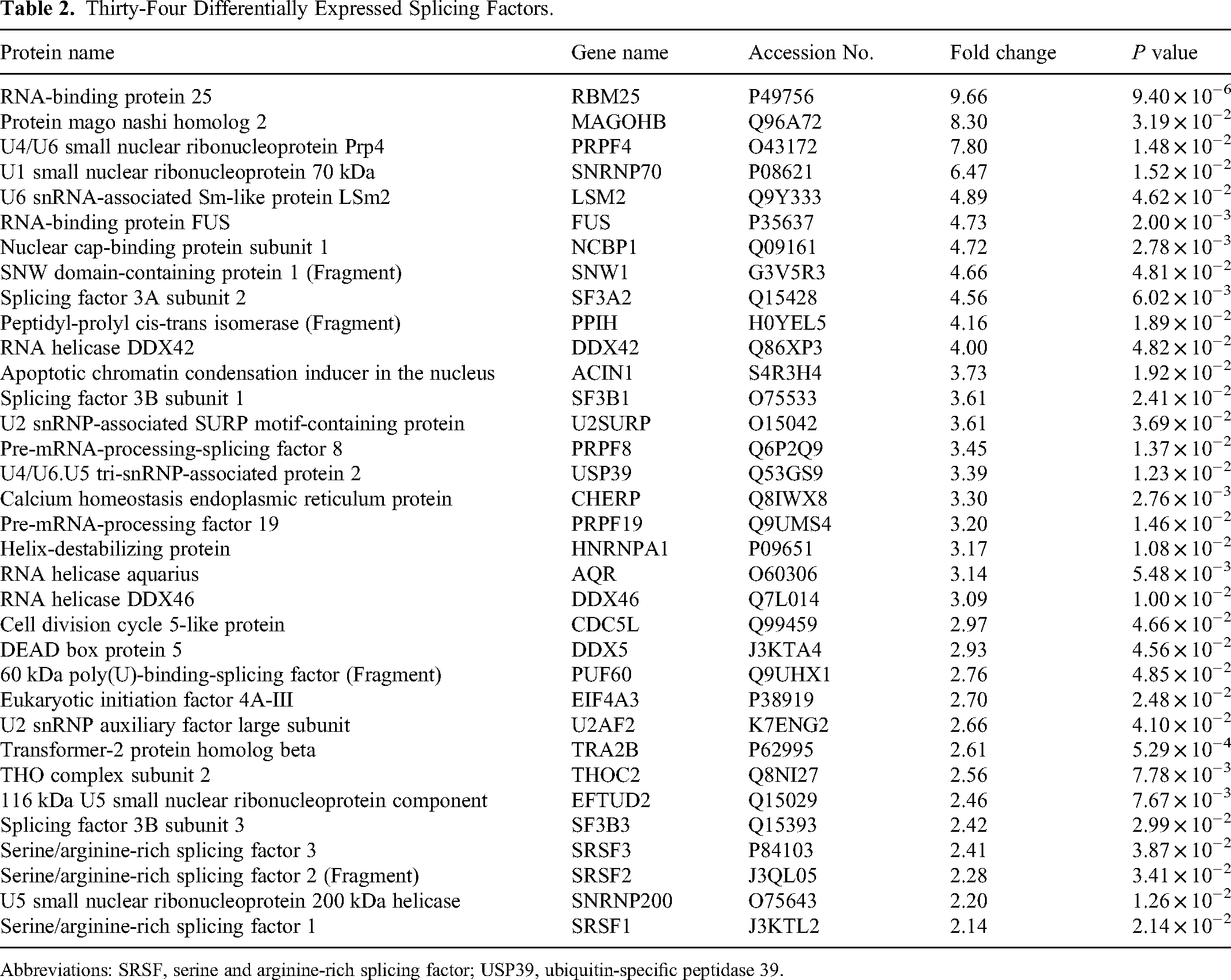
Abbreviations: SRSF, serine and arginine-rich splicing factor; USP39, ubiquitin-specific peptidase 39.
USP39 and STAT1 Expression Are Increased in HNSCC
Among all the differentially expressed splicing factors, USP39 is an important factor and received general concern. To assess the expression of USP39 in HNSCC, we collected HNSCC tumor and adjacent tissues from an additional seven patients (patient identification numbers 5-15) with HNSCC (Table 1). Immunohistochemical comparison of USP39 expression levels in adjacent tissue versus tumor tissue showed that USP39 expression was higher in HNSCC than in the adjacent tissue (Figure 3A). The results of LC-MS/MS and of immunohistochemical analyses (Figure 3B) showed that the expression level of STAT1 in tumor tissues was also significantly higher than that in adjacent tissues.

USP39 and STAT1 expression levels in head and neck squamous cell carcinoma (HNSCC) tumor and adjacent tissues. (A) Representative immunohistochemical images and statistical summary data for USP39 levels in HNSCC tumor and adjacent tissues (AT). The effect size between the two groups is 1.39348 (95% CI: 0.44, 2.32). (B) Representative immunohistochemical images and statistical summary data for STAT1 levels in HNSCC tumor and adjacent tissue. The effect size between the two groups is 1.21 (95% CI: 0.28, 2.11). Data are shown as the mean ± SEM; n = 11. *P < .05, **P < .01 versus adjacent tissues.
Knockdown of USP39 Inhibited HNSCC Cell Viability and Migration
Because our results indicated that the expression of USP39 was increased in HNSCC tissue, to explore the influence of USP39 on the occurrence and development of HNSCC, we used siRNA to silence USP39 and thus inhibit its expression in CAL27 and SCC25 cells. Compared with that in cells transfected with scrambled control siRNA, the expression of USP39 was significantly reduced after transfection of siRNA USP39 (Figure 4A). The results of the CCK-8 assay showed that USP39 knockdown significantly reduced the cell viability of these HNSCC cells compared with the control group (Figure 4B). The ability of cells with the USP39 knockdown to migrate was lower than that of the control group, as assessed using a wound healing assay (Figure 4C-E). These results indicated that USP39 knockdown inhibited the viability and migration of HNSCC cells.
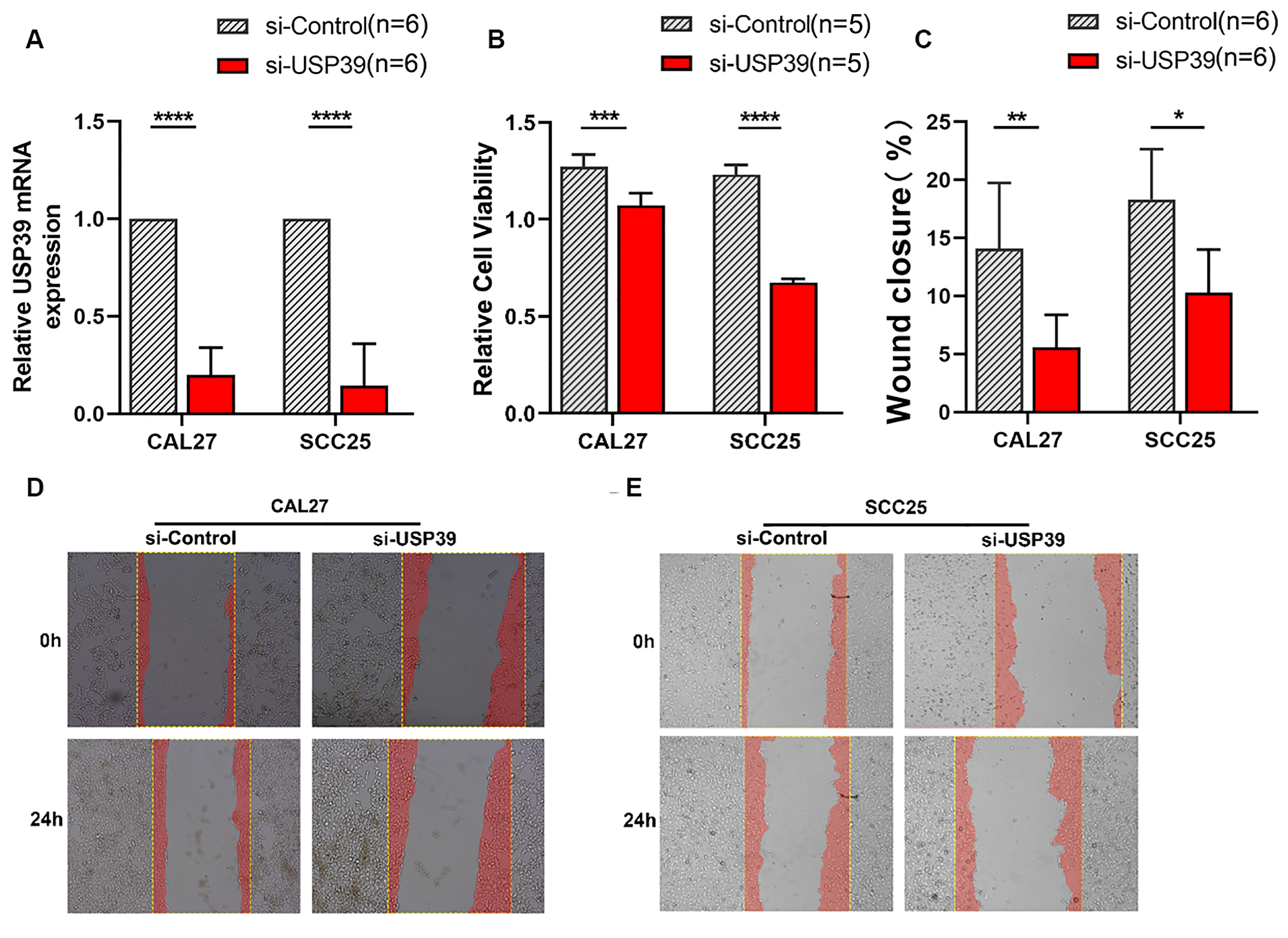
Viability and migration of head and neck squamous cell carcinoma cells with USP39 knockdown. (A) Relative USP39 mRNA in CAL27-si-Control, CAL27-si-USP39, SCC25-si-control, and SCC25-si-USP39 cells were detected by qPCR. The effect sizes and confidence intervals for CAL27 and SCC25 measurements were 8.11 (95% CI: 4.42, 11.77) and 5.66 (95% CI: 2.94, 8.32), respectively. (B) The effect of USP39 on the viability of SCC25 and CAL27 cells transfected with scrambled siRNA (si-control) or with USP39 siRNA (si-USP39) evaluated using the CCK-8 assay. The effect sizes and confidence intervals for CAL27 and SCC25 measurements were 3.21 (95% CI: 1.19, 5.15) and 14.88 (95% CI: 7.65, 22.13), respectively. (C-E) Representative images and migration of CAL27 and SCC25 cells transfected with scrambled siRNA (si-control) and USP39 siRNA (si-USP39) evaluated by the wound healing assay. The effect sizes and confidence intervals for CAL27 and SCC25 measurements were 1.91 (95% CI: 0.48, 3.28) and 1.99 (95% CI: 0.39, 3.52), respectively. Data are shown as the mean ± SEM; *P < .01, **P < .001, ***P < .001, ****P < .0001 for the indicated comparisons.
Knockdown of STAT1 Inhibits HNSCC Cell Viability and Migration
Because the results of our LC-MS/MS experiments indicated that the expression level of STAT1 was higher in HNSCC tumor tissue than in adjacent tissues, we used siRNA knockdown to lower the expression of STAT1 in CAL27 and SCC25 cells to explore the influence of decreased STAT1 expression in HNSCC (Figure 5A). The results of the CCK-8 assay showed that the viability of HNSCC cells was reduced in the STAT1 knockdown group compared with the control group (Figure 5B). The results of the wound healing assay showed that inhibition of STAT1 expression significantly reduced the migration of CAL27 and SCC25 cells (Figure 5C-E). These results suggest that STAT1 knockdown inhibits the viability and migration of HNSCC cells.

Viability and migration of head and neck squamous cell carcinoma cells with STAT1 knockdown. (A) Relative STAT1 mRNA in CAL27-si-control, CAL27-si-STAT1, SCC25-si-control, and SCC25-si-STAT1 cells were detected by qPCR. The effect sizes and confidence intervals for CAL27 and SCC25 measurements were 3.25 (95% CI: 1.42, 5.03) and 7.64 (95% CI: 4.14, 11.11), respectively. (B) Viability of SCC25 and CAL27 cells transfected with scrambled siRNA (si-control) and STAT1 siRNA (si-STAT1) evaluated by the CCK-8 assay. The effect sizes and confidence intervals for CAL27 and SCC25 measurements were 2.83 (95% CI: 1.13, 4.46) and 14.51 (95% CI: 8.68, 20.11), respectively. (C-E) Representative images and migration of CAL27 and SCC25 cells transfected with scrambled siRNA (si-control) and STAT1 siRNA (si-STAT1) evaluated by the wound healing assay. The effect sizes and confidence intervals for CAL27 and SCC25 measurements were 2.43 (95% CI: 0.85, 3.94) and 2.69 (95% CI: 1.03, 4.28), respectively. Data are shown as the mean ± SEM; n = 6. **P < .05, ***P < .001, ****P < .0001 for the indicated comparisons.
Knockdown of USP39 Inhibits STAT1 Expression
In order to study the regulatory relationship between USP39 and STAT1, we knocked down USP39 expression in CAL27 and SCC25 cells by transfecting them with specific or scrambled siRNA and performed real-time qPCR. The results showed that compared with the control group, USP39 knockdown significantly reduced STAT1 mRNA levels (Figure 6). These results indicate that USP39 knockdown may inhibit STAT1 expression.

Effect of USP39 knockdown on expression levels of STAT1 mRNA in head and neck squamous cell carcinoma. The mRNA expression levels of STAT1 in SCC25 and CAL27 cells transfected with scrambled siRNA (si-control) or USP39 siRNA (si-USP39) examined by qPCR. The effect sizes and confidence intervals for CAL27 and SCC25 measurements were 1.75 (95% CI: 0.36, 3.08) and 7.64 (95% CI: 4.14, 11.11), respectively. Data are shown as the mean ± SEM; n = 6. *P < .05, ***P < .001 for the indicated comparisons.
Discussion
In recent years, study progress in HNSCC has been rapid, and the molecular mechanisms of HNSCC are increasingly being revealed. 32 This is the first study, to our knowledge, to provide evidence that USP39 may promote the development of HNSCC by regulating STAT1 expression. Using LC-MS/MS to analyze the protein changes between HNSCC tumor and adjacent tissues of patients with HNSCC, we obtained 590 DEPs. Through GO and KEGG pathway analyses, 34 different proteins were found to be enriched in the spliceosome pathway. Increasingly more attention has been paid in recent years to the role of splicing factors in the occurrence and development of cancer. Some small molecule splicing modulators are currently in the clinical trial stage, providing new options for the treatment of cancer. 33 In the field of HNSCC research, a very recent study shows that serine and arginine-rich splicing factor 3 (SRSF3) can regulate the variable splicing of Ki67 to promote the development of HNSCC. 34 Our mass spectrum analysis also showed increased expression of SRSF3 in HNSCC tumor tissue. In addition to SRSF3, another commonly observed splicing factor in cancer research, USP39, is also upregulated in HNSCC tumor tissue.
USP39 which is reported upregulated in human head and neck cancers has carcinogenic effects in ovarian cancer, glioma, esophageal squamous cell carcinoma, renal cell carcinoma, and HCC, suggesting a key role in treatment.12,13,35–37 It has also been reported that the expression of USP39 is increased in nasopharyngeal carcinoma tissues, the overexpression of USP39 can promote the proliferation of nasopharyngeal carcinoma cells, and the expression of USP39 is regulated by the LINC00520/miR-26b-3p axis. 38 Those findings are consistent with our immunohistochemical results showing that USP39 expression was elevated in HNSCC tumor tissue. In further functional experiments, we found that knockdown of USP39 inhibited the viability and migration of HNSCC cells. Therefore, USP39 may function as a novel oncogene in HNSCC and might represent a potential therapeutic target.
It has been reported that USP39 also acts as a deubiquitination enzyme to maintain the continuous expression of STAT1 and thus exert immune effects. 39 We also found a regulatory effect of USP39 on STAT1 in HNSCC cells. When we knocked down USP39, STAT1 expression was also decreased. In experiments further exploring the effects of STAT1 in HNSCC, we found that STAT1 expression was upregulated in HNSCC tumor tissue compared with adjacent tissues. Knocking down STAT1 inhibited the viability and migration of HNSCC cells. These results suggest that STAT1 may play a carcinogenic role in HNSCC. The results of a study by Knitz et al 40 also suggest that STAT1 may play a tumor-promoting role in HNSCC. Subsequent studies will be required to verify whether the inhibition of STAT1 is responsible for the inhibition of gastric cancer cell proliferation by USP39 siRNA.
The article also has some limitations, as the role of USP39 has not been validated in animal models. In addition, although this study initially discovered that USP39 can influence tumor progression by regulating STAT1, whether the regulatory effect of USP39 on STAT1 is through alternative splicing or deubiquitination requires further investigation.
Conclusion
In summary, our research indicates that USP39 is significantly upregulated in human HNSCC tissues, and knocking down USP39 can inhibit cancer cell viability and migration. According to our research findings, USP39 may be considered a promising candidate for the future treatment of HNSCC.
Footnotes
Abbreviations
Human Subjects Research
The Ethics Committee of Lu’an Hospital Affiliated with Anhui Medical University approved this study (Approval No: 2020LL003) on November 20th, 2020.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article. This work was supported by grants from the Natural Science Foundation of Lu’an People's Hospital (Grant Nos. 2020kykt05 and 2023kykt03); the Lu’an Science and Technology Bureau Foundation (Grant No. 2020laskt03 and 2023lakj003); Innovation and Entrepreneurship Training Program of Anhui Provincial College Students (Grant No. S202310366050); and the Youth Science Foundation of Anhui Medical University (Grant No. 2022xkj101).
