Abstract
OBJECTIVE:
Accumulating evidence indicates that circular RNAs (circRNAs) contribute to breast cancer (BC) development and progression. However, the role of circ_0058063 in BC and its underlying molecular processes remain unclear.
METHODS:
The expression of circ_0058063, miR-557, and DLGAP5 in BC tissues and cells was determined using real time quantitative PCR or western blotting. The functions of circ_0058063 in BC cells were detected using CCK-8, Transwell, caspase-3 activity, and xenograft tumor assays. The specific binding of circ_0058063/miR-557 and DLGAP5/miR-557 was verified using RNA immunoprecipitation (RIP) and dual-luciferase reporter assays.
RESULTS:
circ_0058063 expression was upregulated in BC tissues and cells. circ_0058063 knockdown inhibited proliferation and migration but promoted apoptosis in MCF-7 and MDA-MB-231 cells in vitro. In vivo studies further validated that the knockdown of circ_0058063 repressed tumor growth. Mechanistically, circ_0058063 directly sponged miR-557 and negatively regulated its expression. Additionally, miR-557 inhibition reversed the tumor-suppressive effects of the circ_0058063 knockdown on the survival of MDA-MB-231 and MCF-7 cells. Moreover, miR-557 directly targeted DLGAP5. DLGAP5 knockdown suppressed MCF-7 and MDA-MB-231 cell growth, and these effects were reversed by miR-557 downregulation.
CONCLUSION:
Our findings verify that circ_0058063 acts as a sponge for miR-557 to upregulate DLGAP5 expression. These findings suggest that the circ_0058063/miR-557/DLGAP5 axis is an important regulator of oncogenic function and may be a promising therapeutic target for BC.
Introduction
Among all non-skin cancers, breast cancer (BC) has become the most common since 2004 [1, 2]. Worldwide, approximately 30% of females are diagnosed with BC, with a mortality-to-incidence ratio of 15% [2]. Although the mortality rate has declined owing to improved techniques for early detection and effective systematic treatment, BC continues to be the most frequent type of cancer worldwide that causes death in women [3]. Early diagnosis and feasible targeted therapies are vital for patients with BC [4]. Therefore, the decline in BC mortality can be accelerated by exploring novel therapeutic targets and strategies.
In addition to being genetic, cancer is also epigenetic [5]. Approximately 10% of all BC cases are related to genetics [6]. Epigenetic mechanisms play vital roles in regulating cancer progression and metastasis [7]. Prognostic and diagnostic tools based on epigenetics will significantly improve the precision of oncology [7]. Circular RNAs (circRNAs), discovered in the 1970s, have been recognized as a novel class of RNA that are widely found in various species [8]. When pre-mRNA is backspliced without the 3’ tails and also 5’ caps, single-stranded circular transcripts (circRNAs) are produced [9]. circRNAs are resistant to degradation and their expression is tightly regulated by RNA-binding proteins [9]. Notably, circRNAs can serve as miRNA sponges, templates for protein translation, RNA-binding protein scaffolds, or transcriptional regulators [10]. Recent research has demonstrated that circRNAs are dynamically produced in cancer and are essential components of epigenetic regulatory pathways in tumor growth [11]. Therefore, as research on tumor circRNAs has increased, circRNAs may now be used as unique potential biomarkers for the detection and management of various cancers.
Accumulating evidence has shown that circRNAs are dysregulated and contribute to the development of various cancers [12, 13, 14]. CircRNAs, such as circ-RPPH1, circ 0000514, circBCBM1, and cirCHIPK3, have been found to have strong potential for controlling cell migration, tumor growth, proliferation, and metabolism in BC. Their expression is also associated with prognosis and tumor node metastasis (TNM) stages [15, 16, 17, 18]. However, the biological function of most circRNAs in BC remains largely unknown. According to previous reports, circ_0058063 is overexpressed and has an oncogenic function in a number of cancer types, suggesting that circ_0058063 may contribute to certain malignancies [19, 20]. However, its role in BC has not yet been elucidated.
In this study, circ_0058063 was the primary focus, and its biological properties as well as the underlying molecular processes in BC formation were investigated. Our findings suggest that targeting the circ_0058063/miR-557/DLGAP5 axis may be a novel strategy for the treatment of BC.
Materials and methods
The clinical characteristics of patients with breast cancer
The clinical characteristics of patients with breast cancer
TNBC, Triple-negative breast cancer.
Informed consent was obtained from 36 patients with BC who underwent surgery and the associated neighboring normal breast tissues were collected. No radiation or chemotherapy was administered to any patient before surgery. All tissues were placed in liquid nitrogen immediately and then stored at
Real-time PCR Primer synthesis list
Real-time PCR Primer synthesis list
The cell lines included a normal breast cell line (MCF-10A, cat, no. BNCC337734) and two BC cell lines (MDA-MB-231, cat. no. BNCC337894; MDA-MB-468, cat.no. BNCC339862) obtained from BeNa Culture Collection (Beijing, China). The remaining MCF-7 BC cells (cat. no. ZY-H061) were purchased from Ze-Ye Biotechnology Co., Ltd. (Shanghai, China). Dulbecco’s Modified Eagle’s Medium (DMEM; Invitrogen, USA) supplemented with 10% fetal bovine serum was used to cultivate all of the aforementioned cells (FBS; Gibco, USA). At 37
Cell transfection
Non-targeting siRNA (Si-NC), siRNA targeting circ_0058063 or DLGAP5 (Si-circ or Si-DLGAP5), miR-557 overexpression plasmid (miR-557 mimic:5’-GUUUGCACGGGUGGGCCUUGUCU-3’) or miR-NC, miR-557 knockdown plasmid (miR-557 inhibitor: 5’-AGACAAGGCCCACCCGUGCAAAC-3’) or inhibitor control (inhibitor-NC) were purchased from Ruibo (Shanghai, China). 24-well plates with 5
RT-qPCR
Total cellular RNA was extracted from lncRNA/mRNA using RNAiso Plus (TaKaRa Biotechnology, Japan). RNA was converted into cDNA using the First Strand cDNA Synthesis Kit (Thermo Scientific, USA). Using the Eco RT-PCR System, specific cDNAs was amplified with iTaqTM Universal SYBR
Subcellular fractionation location
Nuclear and cytoplasmic RNAs from MDA-MB-231 and MCF-7 cells were isolated using a Cytoplasmic and Nuclear RNA Purification Kit (Invitrogen, USA). RT-qPCR was performed to detect the cytoplasmic and nuclear circ_0058063 levels. GAPDH was used as a cytoplasmic control used was as a nuclear control used was U6.
RNase R treatment
RNase R was applied to circ_0058063 that was extracted from MCF-7 and MDA-MB-231 cells for 30 minutes at 37
CCK-8 assay
Using the Cell Counting Kit-8 test (Biyuntian, Shanghai, China), MCF-7 and MDA-MB-231 cell proliferation viability was examined. In 96-well plates, cells were seeded at a density of 1
Transwell assay
MCF-7 and MDA-MB-231 breast cell migration was assessed using Transwell assays [21]. A total of 5
Caspase 3 activity assay
Breast cancer cell apoptosis was investigated using a colorimetric caspase-3 activity assay kit (BioVision Research Products, CA, USA). MCF-7 and MDA-MB-231 cells that had not been transfected or treated were used as blank and unfavorable control samples, respectively. Cells were collected and lysed on ice for 15 min, and the supernatants were collected for further analysis after centrifugation (12,000
Xenograft tumor formation assay
All animal experiments were performed in accordance with the Guide for the Care and Use of Laboratory Animals and were approved by the Ethics Committee of our hospital. Six-week-old male nude mice were obtained from our hospital. First, MCF-7 cells stably downexpressing circ_0058063 (Sh-circ) or the corresponding control cells (Sh-NC) were constructed using a recombinant lentivirus expressing the shRNA for circ_0058063 or the control shRNA. The mice were then subcutaneously implanted with 5
Dual-luciferase reporter assays
The circRNA interactome (
RNA immunoprecipitation (RIP) assay
A complete RNA immunoprecipitation (RIP) lysis solution (Millipore, USA) with an RNase inhibitor was used to lyse 1
Western blot analysis
Total proteins were isolated using a kit made by KeyGEN (China), called Total Protein Extraction, and protein concentration was evaluated using a BCA kit (Thermo Scientific). RIPA Lysis Buffer (KeyGEN, China) was used to break down 30 g of protein, which was then resolved on a 10% polyacrylamide gel before being blotted onto a PVDF membrane. These membranes were blocked for 1 h in 5% skim milk at 37
circ_0058063 was markedly upregulated in BC. (A) circ_0058063 expression study using RT-qPCR in BC and normal tissues. (B) RT-qPCR study of the expression of circ_0058063 in BC cell lines (MCF-7, MDAMB468 and MDAMB231) and in normal BC cell line (MCF10A). 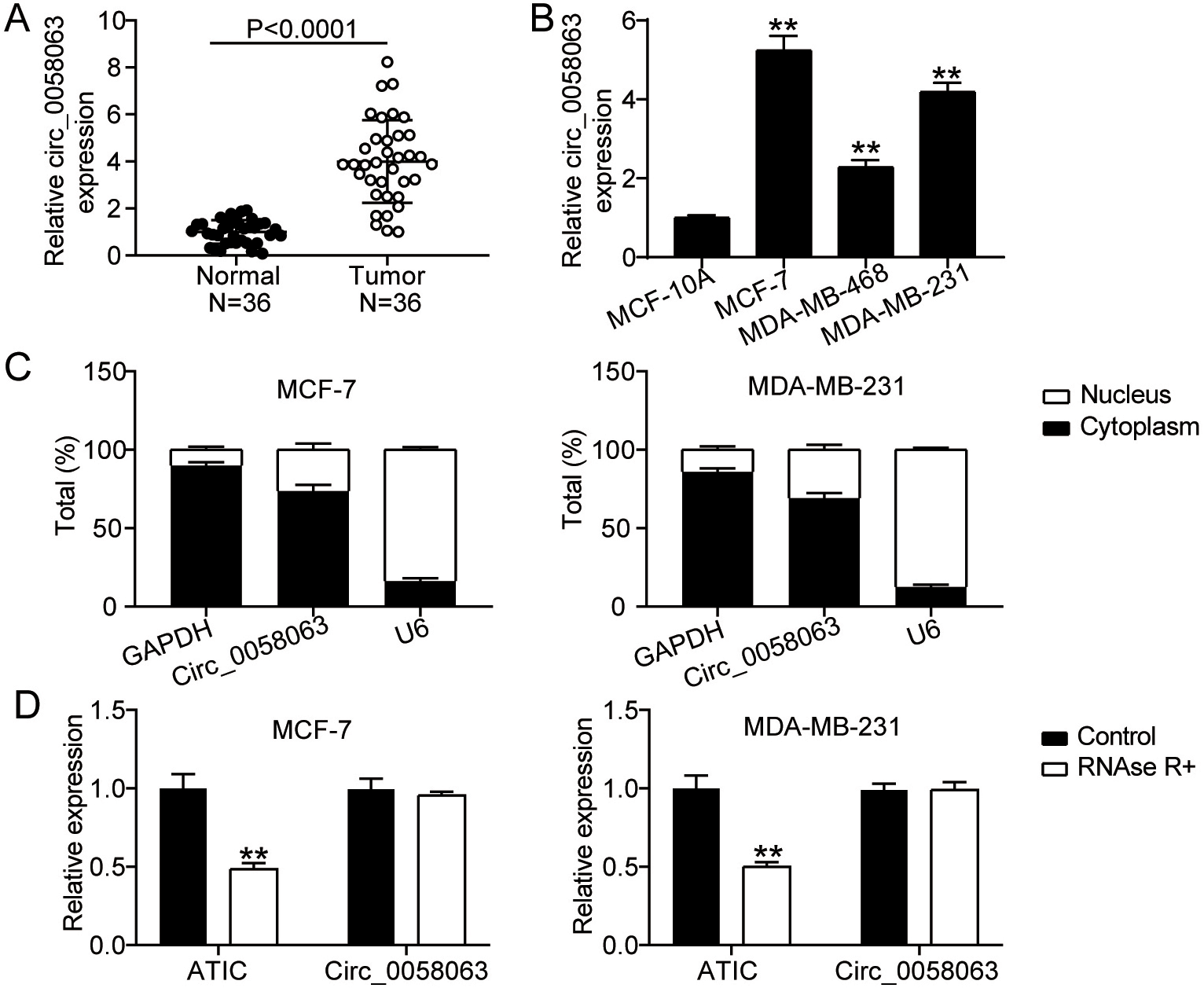
All data in the present study are presented as mean
Results
circ_0058063 was upregulated in BC cells and tissues
RT-qPCR was performed to determine the expression levels of circ_0058063 in BC. The results showed that circ_0058063 expression was considerably increased (four-fold) in cancer tissues compared to normal tissues (Fig. 1A). Additionally, the expression of circ_0058063 was examined in different BC cell lines. According to the RT-qPCR data, circ_0058063 was also clearly upregulated in many BC cell lines (MCF-7, MDA-MB-468, and MDA-MB-231) compared to healthy MCF-10A cells (Fig. 1B). We selected the MDA-MB-231 and MCF-7 cells with the highest circ_0058063 expression for further studies. Considering that cytoplasmic and nuclear circRNAs have different functions, we used a subcellular fractionation test to locate circ_0058063 gene [11]. The results indicated that the predominant locations of circ_0058063 in MCF-7 and MDA-MB-231 cell lines are in the cytoplasm (approximately 80 percent ) (Fig. 1C). These results imply that circ_0058063 may serve as a cytoplasmic circRNA and act as a miRNA sponge or an RNA-binding protein scaffold. Finally, to verify the circularity of circ_0058063, an RNase R digestion assay was performed because circRNAs are more resistant to RNase R than linear RNAs. These results indicated that circ_0058063 was resistant to RNase R digestion, whereas linear ATIC was degraded by RNase R digestion (Fig. 1D). These results confirm the presence and strong expression of circ_0058063.
circ_0058063 downregulation inhibited BC cell growth both in vitro and in vivo. (A) RT-qPCR study of circ_0058063 expression in MCF-7 and MDA-MB-231 cells after transfection of si-circ_0058063 (si-circ) or si-NC . 
To better understand its role in BC, the expression of circ_0058063 was inhibited in MCF-7 and MDA-MB-231 cells. RT-qPCR was performed to assess the expression level of circ_0058063, and the results indicated that it was down-regulated by approximately 60% in the si-circ group (Fig. 2A). CCK-8, Transwell migration, and caspase 3 activity assays were performed to investigate the effects of circ_0058063 on MCF-7 and MDA-MB-231 cell growth. The CCK-8 assay demonstrated that silencing circ_0058063 dramatically reduced the viability of MCF-7 and MDA-MB-231 cells (Fig. 2B). Moreover, Transwell assays showed that circ_0058063 silencing prevented the migration of MDA-MB-231 and MCF-7 cells (Fig. 2C). These experiments demonstrated that circ_0058063 promotes the proliferation and migration of BC cells. Interestingly, we also noted that circ_0058063 knockdown exhibited enhanced caspase 3 activity indicating that the inhibition of circ_0058063 promotes BC cell apoptosis (Fig. 2D). Finally, a mouse xenograft model with circ_0058063 knockdown in primary MCF-7 cells was constructed to assess the effect of circ_0058063 on BC cell proliferation in vivo. Consistent with the in vitro results, the knockdown of circ_0058063 led to a reduction in tumor weight and growth (Fig. 2E). These findings suggested that circ_0058063 regulates BC development in an oncogenic manner.
miR-557 was a target of circ_0058063. (A) The binding sites among miR-557 and circ_0058063 were anticipated by circRNA interactome. (B) The relationship between miR-557 and circ_0058063 was confirmed by dual-luciferase reporter assay in MDA-MB-231 as well as MCF-7 cells. 
Previous studies have reported that circ_0058063 serves as a cytoplasmic circRNA that acts as a sponge for miRNA. Therefore, to determine the probable targets of circ_0058063, we employed a circular RNA interactome (
circ_0058063 facilitated the migration and proliferation of BC cells while suppressed cell apoptosis by targeting miR-557. (A) RT-qPCR analysis of miR-557 expressions in MCF-7 and MDA-MB-231 cells after transfection of si-NC, si-circ, miR-557 inhibitor (inhibitor), inhibitor-NC or si-circ+inhibitor. (B) The proliferation viablity of these transfected MCF-7 and MDA-MB231 cells was assessed using the CCK-8 test. (C) These previously stated transfected MCF-7 and MDA-MB-231 cells underwent a transwell experiment to assess their migration. (D) Levels of caspase-3 of these above-mentioned transfected MDA-MB-231 as well as MCF-7 cells were examined using capase-3 activity assay. 
We tested whether miR-557 knockdown could undo the impact of circ_0058063 knockdown in BC cells to further support the hypothesis that circ_0058063 serves as a sponge for miR-557. According to the PCR data, miR-557 was considerably increased by the knockdown of circ_0058063, whereas it was strongly downregulated by the miR-557 inhibitor in both MDA-MB-231 and MCF-7 cells (Fig. 4A). Using functional assays, we found that miR-557 knockdown considerably aided MDA-MB-231 and MCF-7 cell growth (Fig. 4B). Although the knockdown of circ_0058063 inhibited BC cell proliferation, this effect was reversed when miR-557 expression was inhibited (Fig. 4B). Additionally, the Transwell assay in MDA-MB-231 and MCF-7 cells showed that the number of migrated cells was lower in the si-circ_0058063
DLGAP5 was a target of miR-557 (A) TargetScan made predictions about the miR-557 and DLGAP5 binding locations. (B) The relationship among miR-557 and DLGAP5 was confirmed by dual-luciferase reporter assay in MDA-MB-231 as well as MCF-7 cells. 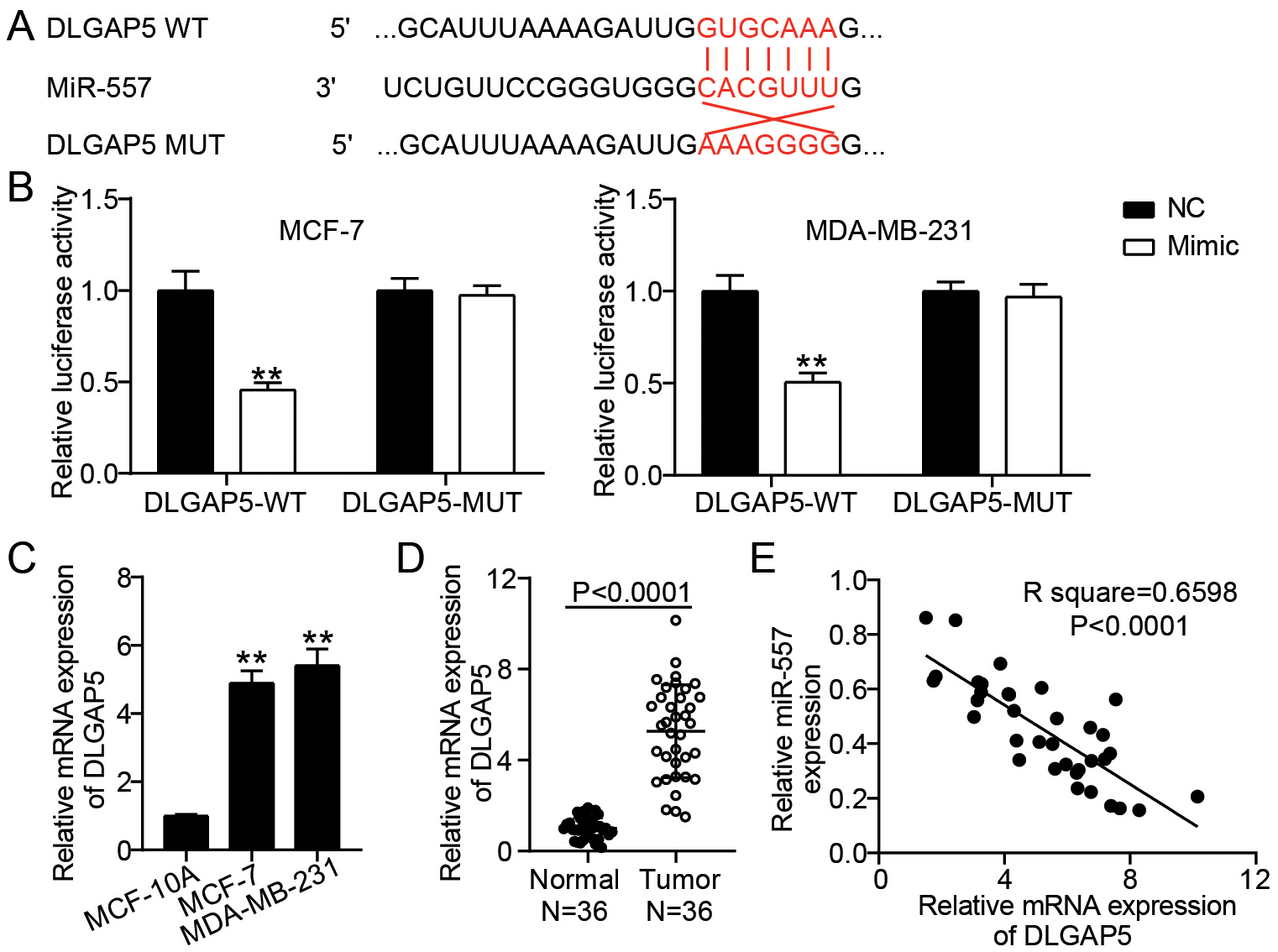
Next, we explored the downstream target genes of miR-557. TargetScan (
miR-557 repressed the proliferation and migration of BC cells while suppressed cell apoptosis by downregulating DLGAP5 expression. (A) Results of western blot study of DLGAP5 expressions in MDA-MB-231 as well as MCF-7 cells after transfection of si-NC, si-DLGAP5, miR-557 inhibitor (inhibitor), inhibitor-NC or si-DLGAP5+inhibitor. (B) To quantify the proliferation of these transfected MDA-MB-231 cells as well as MCF-7, the CCK-8 test was used. (C) To monitor the migration of these transfected MDA-MB-231 and MCF-7 cells, a transwell test was performed. (D) Employing a capase-3 activity test, caspase-3 levels in these transfected MDA-MB-231 as well as MCF-7 cells were evaluated. 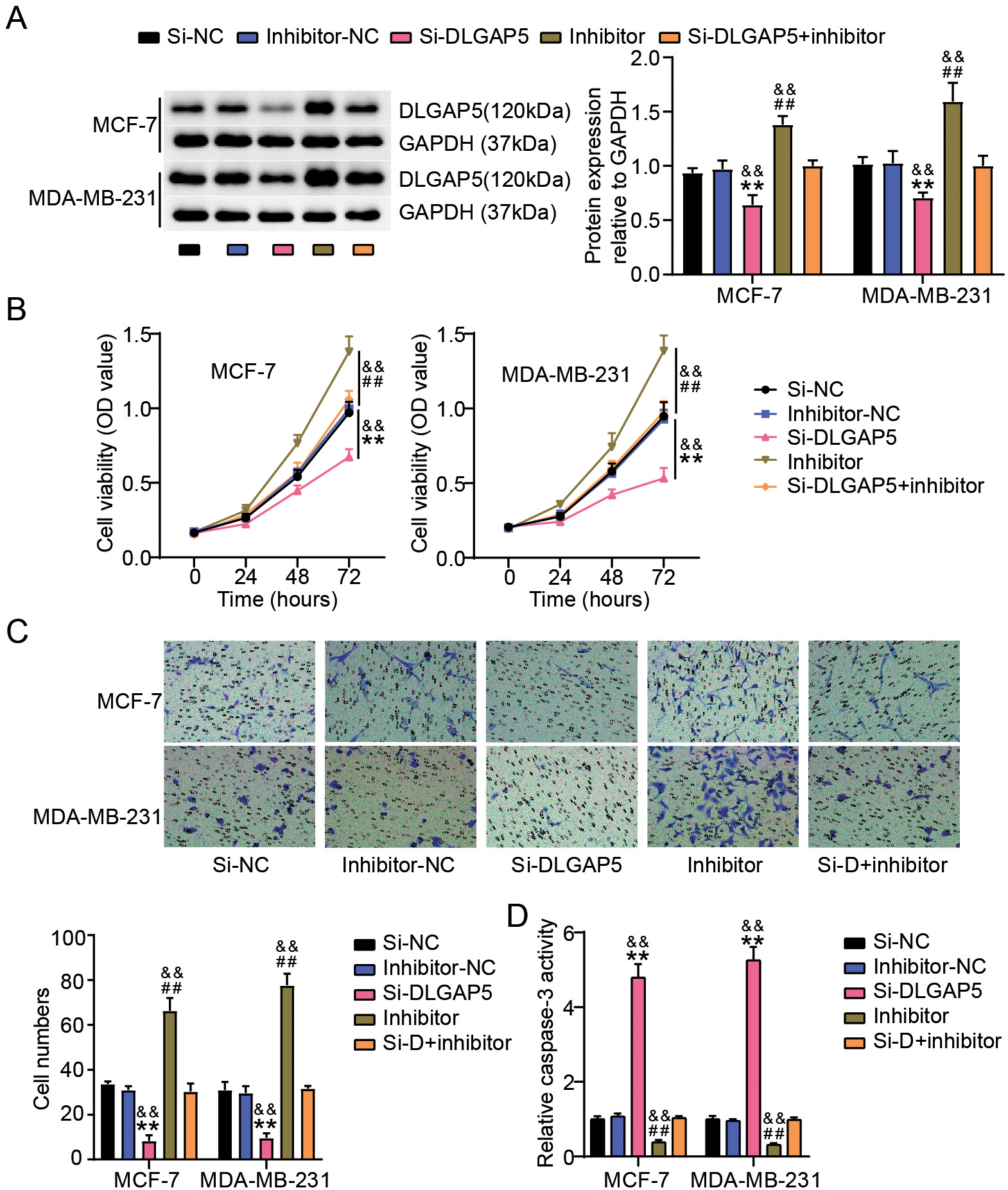
To confirm whether DLGAP5 acts as a target of miR-557, we evaluated whether DLGAP5 knockdown could reverse the effects of miR-557 silencing in BC cells. The results showed that transfection with si-DLGAP5 significantly reduced DLGAP5 expression, whereas miR-557 downregulation significantly upregulated the expression of DLGAP5 both in MDA-MB-231 and MCF-7 cells (Fig. 6A). Although DLGAP5 knockdown inhibited BC cell proliferation, this effect was reversed when miR-557 expression was inhibited (Fig. 6B). Additionally, the Transwell assay revealed fewer migrating cells in the si-DLGAP5
Discussion
The high incidence of BC constitutes a serious threat to the health of women, because it is the most prevalent cancer worldwide [1, 2]. Although significant efforts have been made to reduce mortality, the exact mechanisms underlying the initiation and progression of BC remain elusive [3, 22]. circ_0058063 expression is enhanced in BC. circ_0058063 knockdown prevents BC cells from proliferating and migrating, while inducing cell death. Mechanistically, circ_0058063 acted as a sponge for miR-557 to upregulate DLGAP5 expression, and knockdown of DLGAP5 significantly inhibited BC cell survival. Overall, circ_0058063 is a novel circRNA that regulates BC progression.
Recent studies have demonstrated an association between circRNAs and various cancers [12, 23]. CircRNAs have been implicated in the progression of BC through migration and regulation of cancer cell proliferation, invasion, and metastasis [23, 24, 25]. We identified circ_0058063 as a novel circRNA involved in the regulation of BC progression. Previous studies have reported that circ_0058063 is also upregulated in other types of cancers, including multiple myeloma, esophageal squamous cell carcinoma, and bladder cancer [19, 26, 27, 28]. In detail, circ_0058063 promotes cancer cell proliferation and invasion and inhibits apoptosis in multiple myeloma and bladder cancer, which is consistent with our results [19, 26, 27]. Our results in the present study demonstrated that circ_0058063 is abnormally highly expressed in BC cells. Moreover, circ_0058063 knockdown inhibited BC cell growth in vitro and in vivo. Mechanistically, circ_0058063 plays an essential role in regulating cellular metabolism and GLUT1 expression, which can increase glucose uptake in cancer cells to promote cell proliferation [27]. These findings suggest that circ_0058063 is a useful cancer biomarker for both diagnosis and treatment.
circRNAs regulate cancer progression through various biological effects and mechanisms [9, 29]. Many circRNAs serve as miRNAs or protein inhibitors (‘sponges’) that regulate protein functions [29]. Circ _0058063 acts as a molecular sponge for miRNAs, regulating the expression of its target genes to influence cell growth and death [19, 26]. For instance, miR-635 is targeted and negatively regulated by circ_0058063 in multiple myeloma (MM), and miR-635 knockdown reduces the effects of circ_0058063 on the migration, proliferation, and invasion of MM cells [26]. By sponging miR-486-3p, circ_0058063 promotes rapid growth of bladder cancer [28]. In this study, we demonstrate that miR-557 is a novel target of circ_0058063 in BC. miR-557 is an anti-tumor miRNA that plays a profound role in the formation and development of a number of malignancies, such as osteosarcoma, pancreatic cancer, and lung cancer [30, 31, 32, 33]. Overexpression of miR-557 targets SLC7A11, which is closely linked to the emergence of many malignant tumors and reduces the malignant potential of pancreatic cancer cells. miR-557 is downregulated in pancreatic cancer cells [31]. We also observed that miR-557 was downregulated in BC cells and that miR-557 knockdown reversed the inhibitory effect of circ_0058063 knockdown on BC proliferation and migration. These results also showed that miR-557 is a target of circ_0058063, which affects BC progression.
Subsequently, we demonstrated that DLGAP5 was a target of miR-557. Unlike other DLGAP family members, DLGAP5 is mainly associated with various types of cancers [34]. DLGAP5 promotes microtubule polymerization and bipolar spindle formation, which facilitates mitosis by passing through the G2/M checkpoint [35, 36]. Numerous malignancies, including hepatocellular carcinoma, lung cancer, ovarian cancer, gliomas, and pancreatic cancer, have been shown to have elevated levels of DLGAP5 [37, 38, 39]. According to previous studies, DLGAP5 is considerably elevated in BC tissues and is linked with tumor grade and lymph node status [40, 41]. Higher DLGAP5 expression is associated with poor prognosis [41]. Consistent with previous studies, we demonstrated that DLGAP5 was upregulated and participated in the regulation of BC cell proliferation. In mechanism, knockdown of DLGAP5 significantly downregulated the expression of cell cycle-related proteins, CDK1, Cyclin B1, induced G2/M phase arrest, and inhibited cell proliferation [37]. Furthermore, DLGAP5 upregulates the expression of Bcl-2 and downregulates Bax expression, which can inhibit cell apoptosis [19]. As DLGAP5 exhibits an isolated microtubule-associated protein 1A/1B light chain 3B-interacting region motif and is located in the mitochondria, recent studies have shown that DLGAP5 may play a role in mitophagy [41]. These studies showed that DLGAP5 plays a vital role in cancer progression and may serve as a candidate target for the diagnosis and treatment of BC. Furthermore, DLGAP5 was found to be a target of miR-557, which reversed the antitumor effects of miR-557 on BC cell growth. In summary, DLGAP5 facilitated BC progression and was targeted by miR-556-5p.
This study has some limitations. First, the correlation between aberrant circ_0058063 expression levels and the clinicopathological characteristics of patients with BC has not been fully analyzed. Second, whether there is a correlation between driver oncogene mutation type and circ_0058063 expression in BC remains to be explored. We intend to address these questions in future studies. In addition, owing to the key regulatory effect of abnormal circ_0058063 expression in BC, whether circ_0058063 with high and low expression is correlated with the prognosis of BC should be explored in the future by collecting more clinical samples.
Conclusion
Our study showed that miR-557 expression was decreased in BC, while circ_0058063 expression was increased. circ_0058063 knockdown significantly reduced BC cell proliferation and migration. Mechanistically, circ_0058063 targets miR-557, which downregulates DLGAP5. Our findings regarding the circ_0058063/miR-557/DLGAP5 axis may provide a potential target for BC therapy and early diagnosis.
Funding
There was no funding received.
Ethics approval
The Ethics Committee of Wuhan NO.1 Hospital (Wuhan, China) approved this study. The processing of the clinical tissue samples complied with the ethical principles of the Declaration of Helsinki. All of the patients completed an informed consent form.
The animal experiment conducted observed the ARRIVE guidelines and was authorized by the Ethics Committee of Wuhan NO.1 Hospital (Wuhan, China).
Consent to participate
All patients signed a written informed consent.
Consent for publication
The participants gave their consent for the study to be published.
Availability of data and material
This article contains all of the data that were created or examined during this investigation.
Authors’ contributions
Conception: CMT.
Interpretation or analysis of data: KJZ and CY.
Preparation of the manuscript: CMT and CY.
Revision for important intellectual content: KJZ.
Supervision: CY.
The article has been reviewed and approved by all authors.
Footnotes
Acknowledgments
None.
Conflict of interest
There were no conflicts of interest, according to the authors.
