Abstract
BACKGROUND:
Mitosis-associated genes are dysregulated in many types of cancers and play important roles in disease progression and chemotherapy resistance. However, their expression and functions in chemotherapy-resistant Acute Myeloid Leukemia (AML) are still largely undetermined.
OBJECTIVE:
This study aims to explore the roles of spindle assembly checkpoint (SAC) genes CENPE, CENPF, and DLGAP5 in chemotherapy-resistant AML.
METHODS:
RNA-sequencing (RNA-seq) was performed in patients with chemotherapy-resistant AML and chemotherapy-sensitive AML. AML mRNA data from 151 patients with recurrence were downloaded from TCGA. Integrated analysis of the differentially expressed genes (DEGs), GO and KEGG pathways. CENPE, CENPF, or DLGAP5 knockdown cell lines were used to analyse proliferation, apoptosis and cell cycle alterations.
RESULTS:
A total of 87 DEGs (48 upregulated and 39 downregulated) were obtained through gene analysis of R/R-AML and a total of 329 DEGs (202 upregulated and 127 downregulated) were obtained in refractory S-AML. Upregulated DEGs were mainly enriched in cell cycle (GO: 0007049, hsa04110) and mitotic cell cycle (GO: 0000278) processes and pathway. Venn diagram analysis identified the most upregulated DEGs (including CENPE, CENPF, and DLGAP5) in chemoresistant AML. The expression of CENPE, CENPF and DLGAP5 in R-AML (TCGA) was significantly higher than that of primary AML (GEO). The proliferation of K562 cells after CENPE and DLGAP5 knockdown was significantly decreased (
CONCLUSIONS:
In conclusion, the mitotic cell cycle-associated genes CENPE, CENPF, and DLGAP5 were upregulated in chemotherapy-resistant AML patients and might be useful for predicting poor prognosis.
Introduction
Acute Myelogenous Leukemia (AML) is a group of highly heterogeneous hematologic malignant diseases that arise from the malignant transformation of hematopoietic progenitor cells at different stages. Complete remission (CR) rates in adult AML can reach 50%–80%, however, long-term survival is only approximately 20%–30%, and relapse and drug resistance are the main causes of death in AML patients. Relapsed/refractory Acute Myelogenous Leukemia (R/R-AML) and secondary AML (S-AML) are high-risk subsets of AML with adverse prognoses that often show resistance to traditional chemotherapy and are the main causes of death induced by AML [1]. Treatment of R/R-AML and S-AML has been a challenge for hematologists over the past decades. Despite numerous emerging clinical studies, outcomes for AML are consistently unsatisfactory, with 5-year overall survival rates of 10% [2]. The mechanisms leading to therapy resistance and relapse are not fully understood [3]. Therefore, a comprehensive understanding of the molecular biological characteristics of patients with R/R-AML and S-AML is of great significance for exploring potential drug resistance mechanisms. It helps to discover key therapeutic targets and develop new and effective treatment methods.
AML patients normally present with uncontrolled proliferation of myeloid progenitor cells and mitotic errors over the cell cycle result in aberrant chromosomes and contribute to genetic instability, which plays critical roles in the onset and progression of AML [4]. Mitotic slippage contributes to genomic instability [5], and the mitotic process is supervised by the spindle assembly checkpoint (SAC). The study revealed that AML cells with reduced BUBR1 (regulated by CENPE and CENPF) were able to progress through mitosis even in the presence of structural DNA aberrations [6]. As a result, such AML cells can skip mitotic arrest, resulting in premature sister chromatid separation and subsequently reduced sensitivity to antibiotic treatment [7]. In our preliminary bioinformatics study of RNA sequencing in clinical patients with chemoresistant AML, we identified three mitotic cell cycle-related genes, CENPE, CENPF, and DLGAP5, that may be associated with chemoresistance. Centromeric protein E (CENPE) is a kinesin-like motor protein and is crucial for microtubule kinetochore capture, which is required for chromosome alignment over prometaphase [8]. CENPE is abnormally expressed in a variety of solid tumors, and is closely related to tumor occurrence and development [9]. High CENPE expression in lung cancer and breast cancer tissues can promote cancer cell proliferation and predict poor prognosis [10, 11, 12]. Furthermore, CENPE regulated by the chromatin promoter and FOXM1 increases the proliferation of prostate cancer and lung adenocarcinoma, respectively [10, 13]. CENPE inhibitor PF-2771 can suppress the growth of breast cancer cell xenografts [12]. Previous studies have shown that CENPE has the potential to promote the occurrence and development of AML. Most recent studies showed that knockdown of CENPE expression in THP-1 cells resulted in a reduction in cell numbers, cell proliferation, and reversal of cytarabine (Ara-C) chemoresistance [14, 15]. And the role of leukemogenesis development can by regulated by RNA binding protein Lin28A [15]. Thus, it is of great importance to explore the role and mechanisms of CENPE in chemotherapy-resistance AML.
Mitotic protein centromeric protein F (CENPF) was originally found in the serum of patients with autoimmune diseases [16]. CENPF is a protein associated with chromosomal segregation during mitosis [17, 18]. CENPF accumulates in the G2/M phase and is rapidly degraded after mitosis [19]. Knockdown of CENPF can inhibit chromosome assembly during mitosis, affect chromosome segregation, and block cell cycle progression [20, 21]. CENPF is abnormally expressed in a variety of cancer types and participates in cancer onset and development. In prostate cancer, silencing CENPF inhibited the proliferation, invasion and migration of prostate cancer cells [17]. In addition, CENPF interference can prevent the proliferation of liver cancer cells and induce cell cycle arrest in a similar way [22]. Thus, upregulated CENPF has been identified as a cell proliferation marker for a series of solid tumors and lymphoma [23, 24]. However, the role of CENPF in the occurrence and development of AML has rarely been studied.
Clinical parameters of AML patients
Clinical parameters of AML patients
Discs large homologue-associated protein 5 (DLGAP5) is mainly involved in regulating the formation and movement of the spindle in mitosis, cell cycle progression, cell adhesion and signal transduction [25, 26], which plays fundamental roles in spindle assembly, kinetochore fiber stabilization, and chromosome segregation during mitosis [27, 28]. Recent studies have found that DLGAP5 is significantly highly expressed in various kinds of solid tumors, including breast cancer, colorectal cancer, and non-small-cell lung cancer, and is closely related to an unsatisfactory prognosis [28, 29, 30]. DLGAP5 knockout can significantly prevent proliferation and induce cell cycle arrest in breast cancer cells [29]. Abnormally expressed DLGAP5 can also participate in tumor progression. Yamamoto et al. suggested that DLGAP5, which is overexpressed in castration-resistant prostate cancer, can advance the growth and migration of prostate cancer cells [31]. More importantly, a previous study identified that high expression of DLGAP5 promoted AML cell clone formation [32]. DLGAP5 is highly expressed in bone marrow precursor cells but is decreased during the process of stem cell differentiation [33]. However, the role of DLGAP5 in chemotherapy resistance in AML is unclear.
As regulators of chromosome segregation, the roles of the mitotic proteins CENPE, CENPF, and DLGAP5 have rarely been studied in chemotherapy-resistant refractory and relapsed AML. In this study, RNA-sequencing (RNA-seq) was performed in R/R-AML, S-AML, de novo AML patients and healthy controls. We found that the CENPE, CENPF, and DLAGP5 genes were upregulated and further enriched in the mitotic cell cycle process in chemoresistant AML. We further downloaded mRNA data of Recurrence AML (R-AML) and primary AML from TCGA and GEO databases to analyze the expression of the 3 genes. In vitro knocking down CENPE, CENPF, and DLGAP5 inhibited cell growth, promoted cell apoptosis and induced cell cycle arrest in the M and G1/S phases. Taken together, our data indicate that the expression of the mitosis-associated genes CENPE, CENPF, and DLGAP5 is associated with unfavorable prognosis and chemotherapy resistance in AML patients.
Experimental subjects
The expression profiles of mRNA were analysed among 3 cases of R/R-AML (relapsed and resistance to cytarabine treatment), 3 cases of drug-resistant S-AML (MDS- or MPN-derived AML that failed to achieve CR after cytarabine and decitabine treatments), 4 cases of de novo AML (achieved CR after induction chemotherapy) and 3 cases of healthy controls. All samples were collected in Henan Provincial People’s Hospital. The Ethics Committee of Henan Provincial People’s Hospital approved this study. All de novo AML patients had not gone through chemotherapy or had infections when the samples were collected. Written consent was signed by all individuals or their parents/guardians following the Declaration of Helsinki. The diagnosis of de novo AML, R/R-AML and S-AML patients was established by applying FAB diagnostic criteria. The detailed information of the AML patients and healthy controls is delineated in Table 1.
shRNA primer sequences
shRNA primer sequences
Peripheral blood mononuclear cells (PBMCs) from all individuals were collected and separated by density centrifugation (Ficoll-Hypaque). Take 1 ml of fresh anticoagulated whole blood, mix 1:1 with saline and carefully add it to the liquid surface of 2 ml of Lymphocyte Separation Medium (Human, Solarbio). The second layer of cells was collected into a tube containing 4–5 ml of saline, mixed thoroughly, and centrifuged at 400 g for 20 minutes. The sediment was washed twice to obtain the desired PBMCs. PBMCs were enumerated and assessed for viability, using trypan blue and a Boeco haemocytometer (Hamburg, Germany). All specimens were obtained from EDTA peripheral blood in 4 hours, then preserved at
RNA-seq and bioinformatic analysis
A Nanodrop was applied to quantify the total RNA samples. Illumina kits were used to prepare the RNA-seq library, including procedures of poly-A mRNA purification, random fragment RNA, random mRNA priming to cDNA, second-strand cDNA synthesis, restriction enzyme digestion, sequencing adapter ligation and library PCR amplification. Ultimately, quantifying and qualifying the RNA-seq libraries, the sequencing was detected by an Illumina HiSeq 4000. DEGs were screened with adjusted
Downloaded TCGA and GEO RNA-seq data
RNA-seq dataset of R-AML was downloaded from TCGA (151 cases), and the primary AML dataset was downloaded from GEO (7 cases). The DEGs between R-AML and primary AML samples were identified based on screening criteria: fold change
Cell culture and transfection
The K562 and THP-1 (human myeloid leukemia) cell lines were purchased from the American Type Culture Collection (ATCC). Cells were cultivated in RPMI 1640 medium (Sigma Aldrich, USA) carrying 10% fetal bovine serum (Gibco, USA) and 1% 100 U/ml penicillin/streptomycin (37
Sequence of RT-PCR primers
Sequence of RT-PCR primers
CCK-8 method to detect the in vitro cell proliferation. In the CCK8 assay, 100
RT-PCR measurement
K562 and THP-1 cell lines with or without targeted gene knocked down were collected to extract total RNA. cDNA was synthesized using a Bio-Rad iScript cDNA Synthesis Kit (Bio-Rad Laboratories). RT-PCR was conducted with a SYBR Green reaction system (12
Statistical analysis
FPKM calculations at the gene level and transcript level were performed using R software Ballgown. PCA analysis, and correlation analysis based on gene expression levels. R language was applied to perform clustering, GO functional significance enrichment analysis, and pathway significance enrichment analysis for differentially expressed genes. The results of in vitro studies are presented as the mean
Result
Expression profiles of DEGs in chemoresistance R/R-AML and S-AML patients
In the present study, RNA-seq was performed in 3 R/R-AML, 3 drug-resistant S-AML, 4 de novo AML patients and 3 healthy controls, and the sequencing data were acquired by the Illumina Hiseq2500 platform. Scatter plots and hierarchical clustering were used to assess the gene expression variation among the samples (Fig. 1). DEGs with
Expression profiles of DEGs among R/R, S-AML and de novo AML patients. A. Scatter plots and hierarchical clustering in R/R-AML and de novo AML patients. B. Scatter plots and hierarchical clustering in S-AML and de novo AML patients.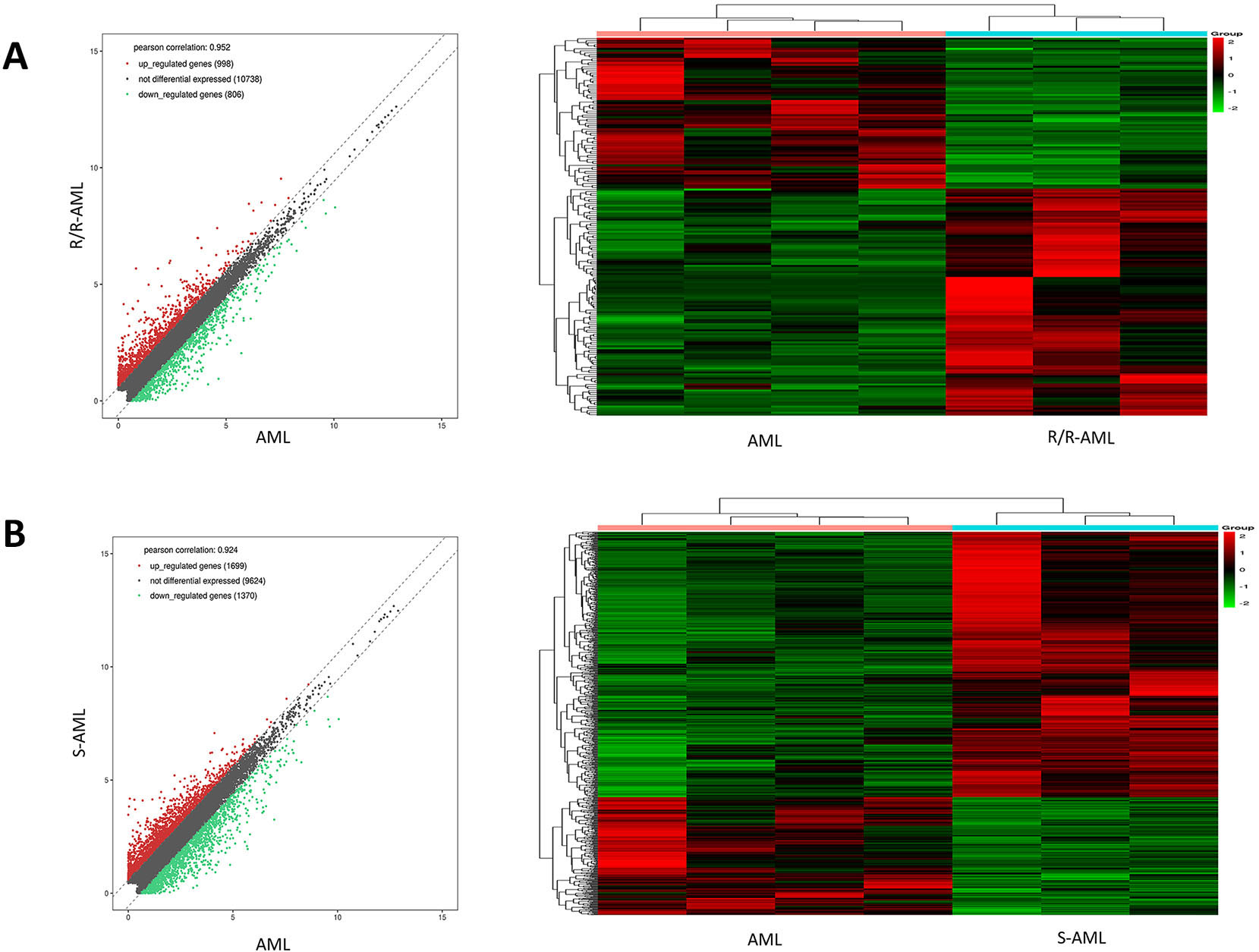
We performed GO and KEGG pathway analysis of DEGs involved in R/R-AML and S-AML patients. GO analysis was used to categorize and delineate the biological functions of different genes. The top 10 enriched GO terms in BP are shown in Fig. 2. Among the de novo AML samples, the upregulated DEGs of biological processes were mainly involved in mitotic cell cycle (GO: 0000278), cell division (GO: 0051301), organelle organization (GO: 0006996), cell cycle (GO: 0007049), and small molecule metabolism process (GO: 0044281). Conversely, downregulated genes were distributed in immune system process regulation (GO: 0002682), lymphocyte activation (GO: 0046649), T cell activation regulation (GO: 0050863), immune response regulation (GO: 0050776), T cell activation (GO: 0042110) (Fig. S2A). In KEGG pathway analysis, the upregulated DEGs were mostly enriched in the cell cycle (hsa04110), while the downregulated DEGs were enriched in the T cell receptor signaling pathway (hsa04660) (Fig. S2B). Among R/R-AML samples, upregulated DEGs were mainly involved in phosphatidylinositol 3 phosphate biosynthetic process (GO: 0036092), while downregulated DEGs were enriched in regulation of immune system process (GO: 0002682) (Fig. 2A). In the KEGG pathway analysis, the upregulated DEGs were enriched in transcriptional misregulation in cancer (hsa05202), while the downregulated DEGs were enriched in the hematopoietic cell lineage (hsa04640) (Fig. 2B). Among the S-AML samples, upregulated DEGs of biological processes were further enriched in the cell cycle (GO: 0007049) and mitotic cell cycle (GO: 0000278), while the downregulated DEGs were mainly enriched in the immune response (GO: 0006955) and immune system process (GO: 0002376) (Fig. 2C). In KEGG pathway analysis, the upregulated DEGs were further enriched in the cell cycle (hsa04110) while downregulated DEGs were enriched in the hematopoietic cell lineage (hsa04640) (Fig. 2D).
GO and KEGG pathway analyses of DEGs in chemoresistant R/R-AML and S-AML patients. A. Upregulated and downregulated DEGs enriched in the top 10 GO biological processes in R/R-AML. B. Upregulated and downregulated DEGs enriched top 10 KEGG pathways in R/R-AML. C. Upregulated and downregulated DEGs enriched in the top 10 GO biological processes in S-AML. D. Upregulated and downregulated DEGs enriched in the top 10 KEGG pathways in S-AML.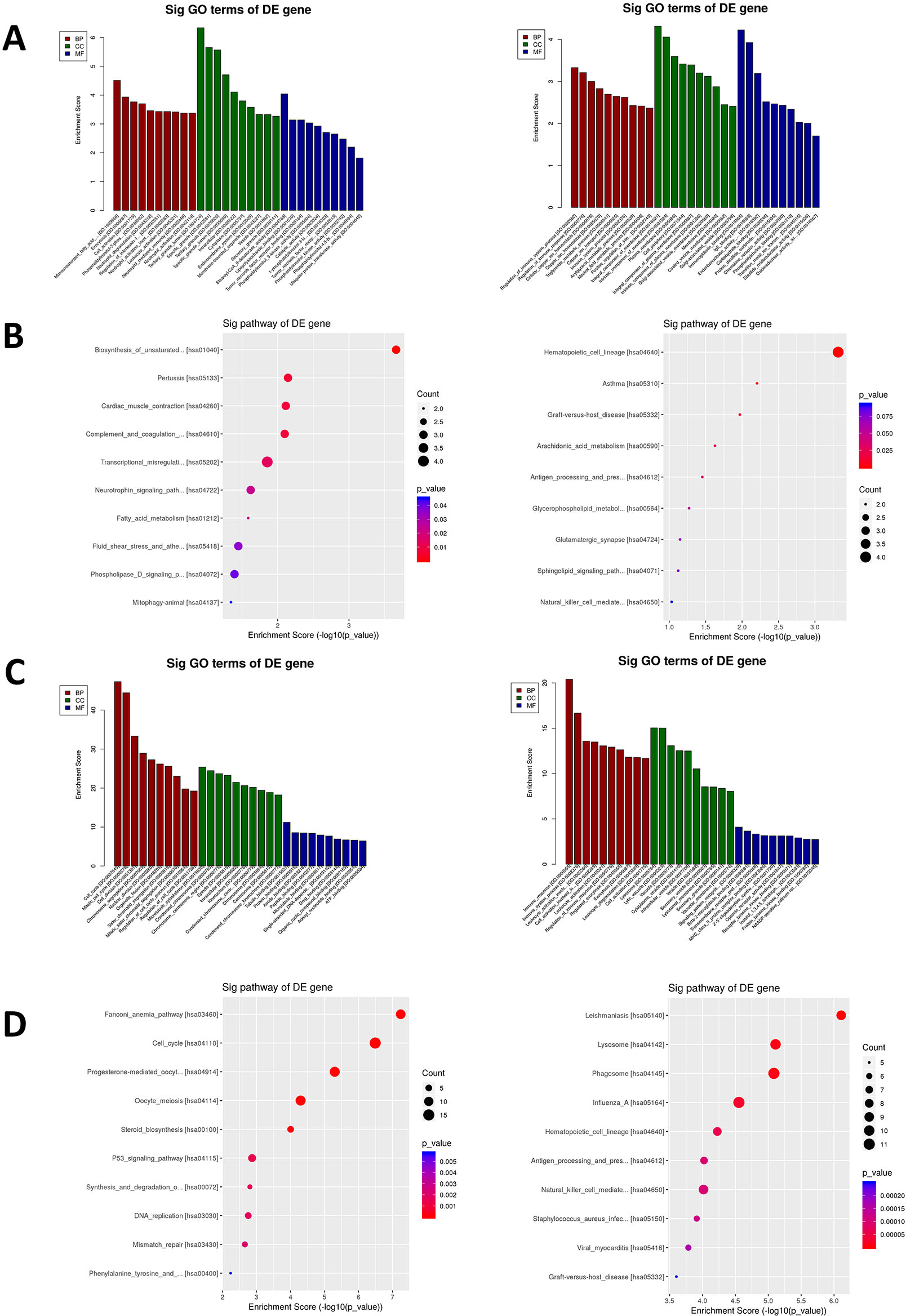
The expression profiles of 19 upregulated overlapping genes in chemotherapy resistant S-AML patients
Identification of target genes. A. Venn diagram and overlapping genes of upregulated DEGs. Upregulated DEGs were selected with a fold change 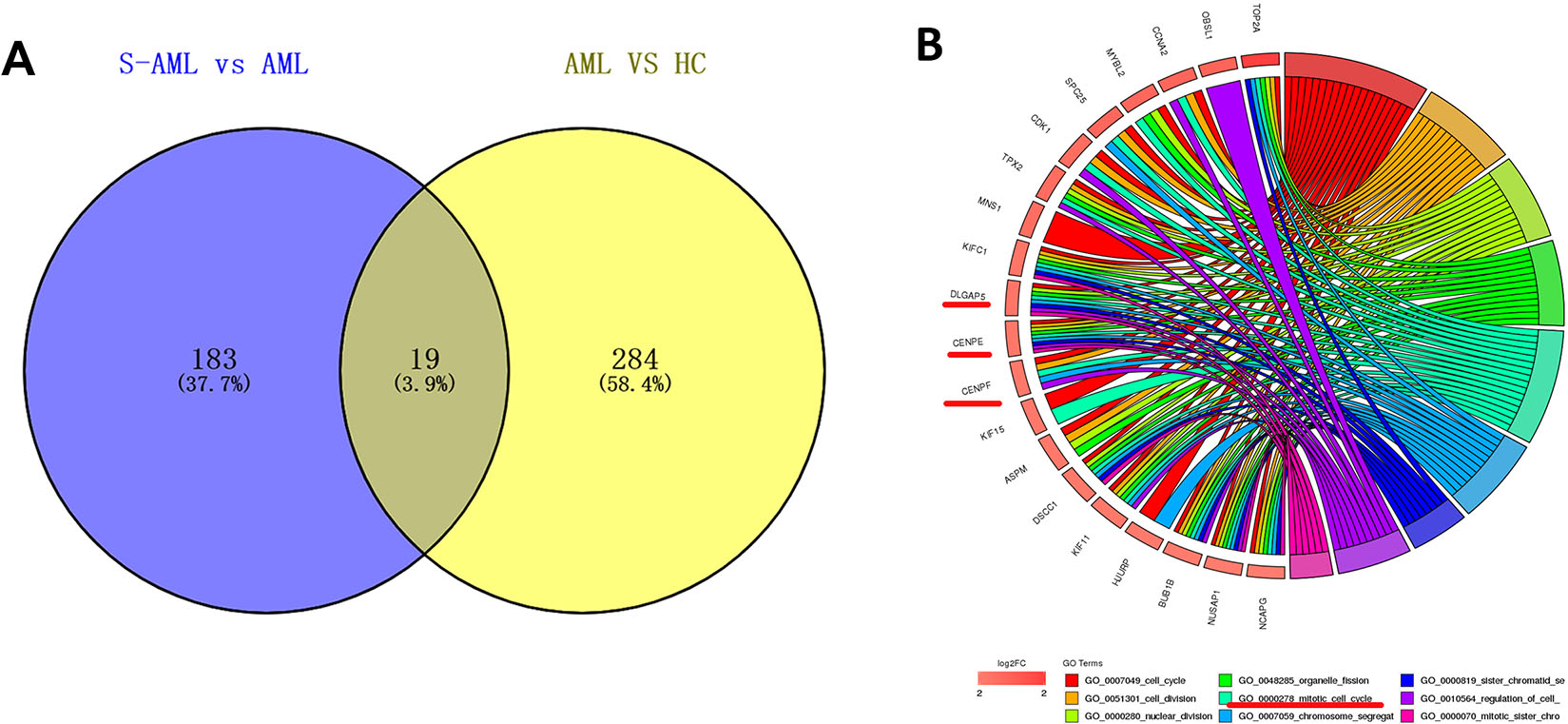
Clinical parameters of R-AML samples in TCGA database
Clinical characteristics of R-AML patients from the TCGA database. A. Survival curves for gender, age, prior treatment, and prior malignancy in R-AML (TCGA). B. Analysis of CENPE, CENPF, and DLGAP5 gene expression in R-AML (TCGA) and primary AML (GEO). C. Correlation between CENPE, CENPF, and DLGAP5 gene expression in AML. D. Survival rate of R-AML patients with different expression of CENPE, CENPF and DLGAP5.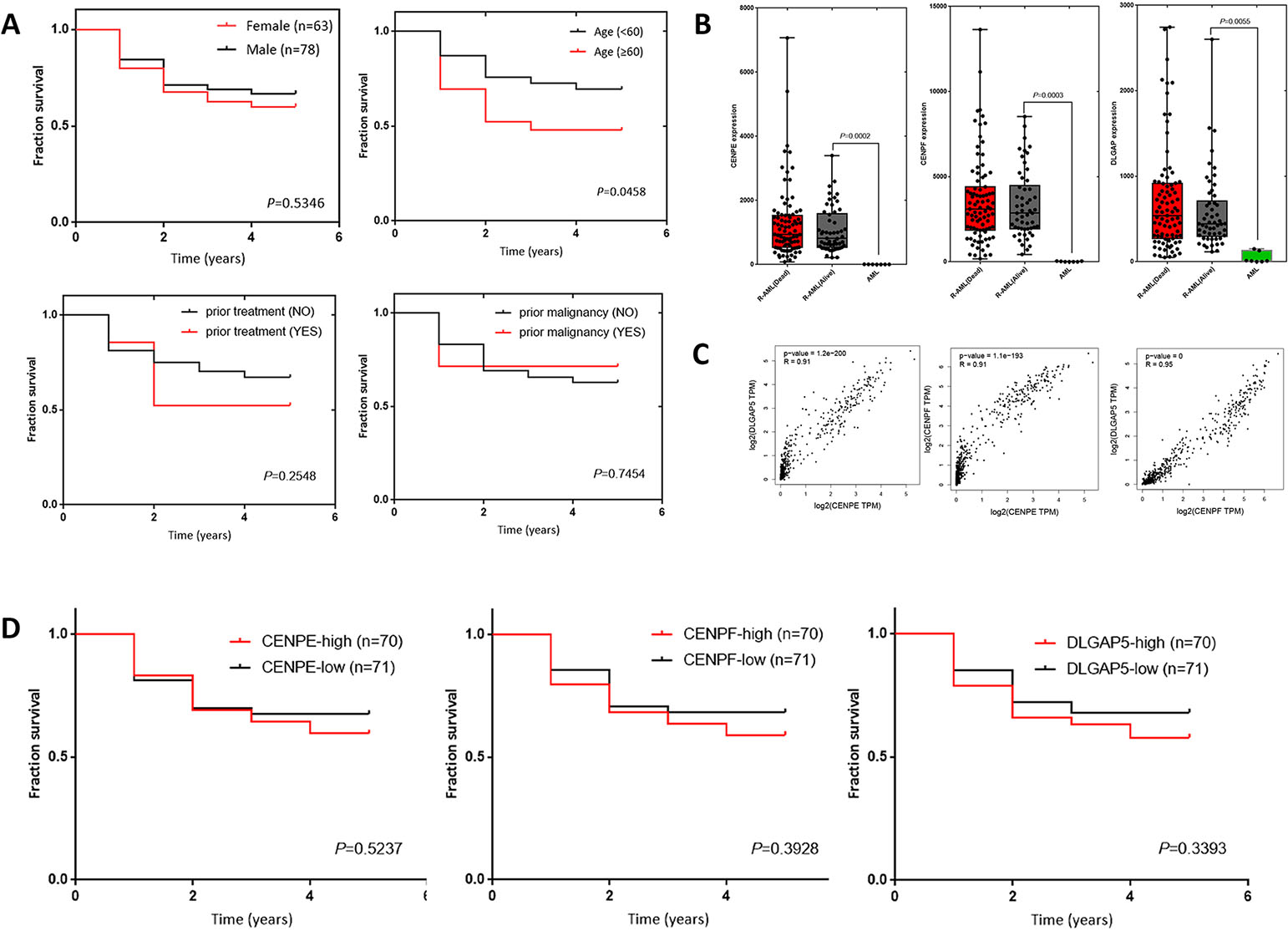
Functional analysis of CENPE, CENPF, and DLGAP5 by shRNA. A. Knockdown efficiency of CENPE, CENPF, and DLGAP5-shRNA. B. The CCK8 assay showed that cell proliferation was significantly suppressed after CENPE, CENPF, and DLGAP5 shRNA transfection.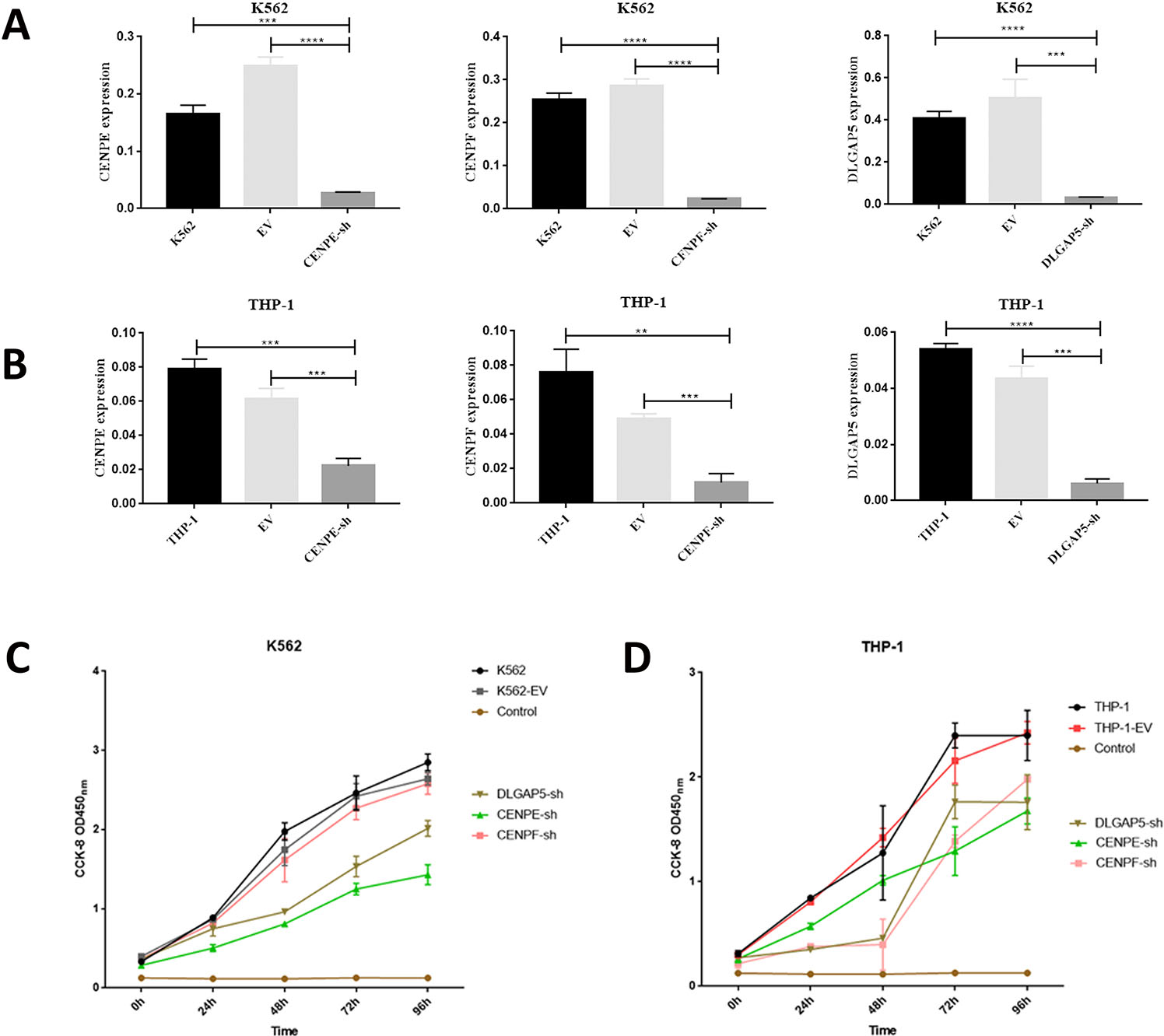
RT-qPCR analysis of the expression of cell apoptosis and cell cycle genes in leukemia cells. A. RT-PCR of the gene expression of the apoptosis genes caspase-9, BAX, bcl-2 and TP5 in K562 cells. B. RT-PCR of the gene expression of the cell cycle-related genes CDK1, CDK2, CDKN1A and CyclinD1 in K562 cells. C. RT-PCR of the gene expression of the apoptosis genes caspase-9, BAX, bcl-2 and TP5 in THP-1 cells. D. RT-PCR of the gene expression of the cell cycle-related genes CDK1, CDK2, CDKN1A and CyclinD1 in THP-1 cells.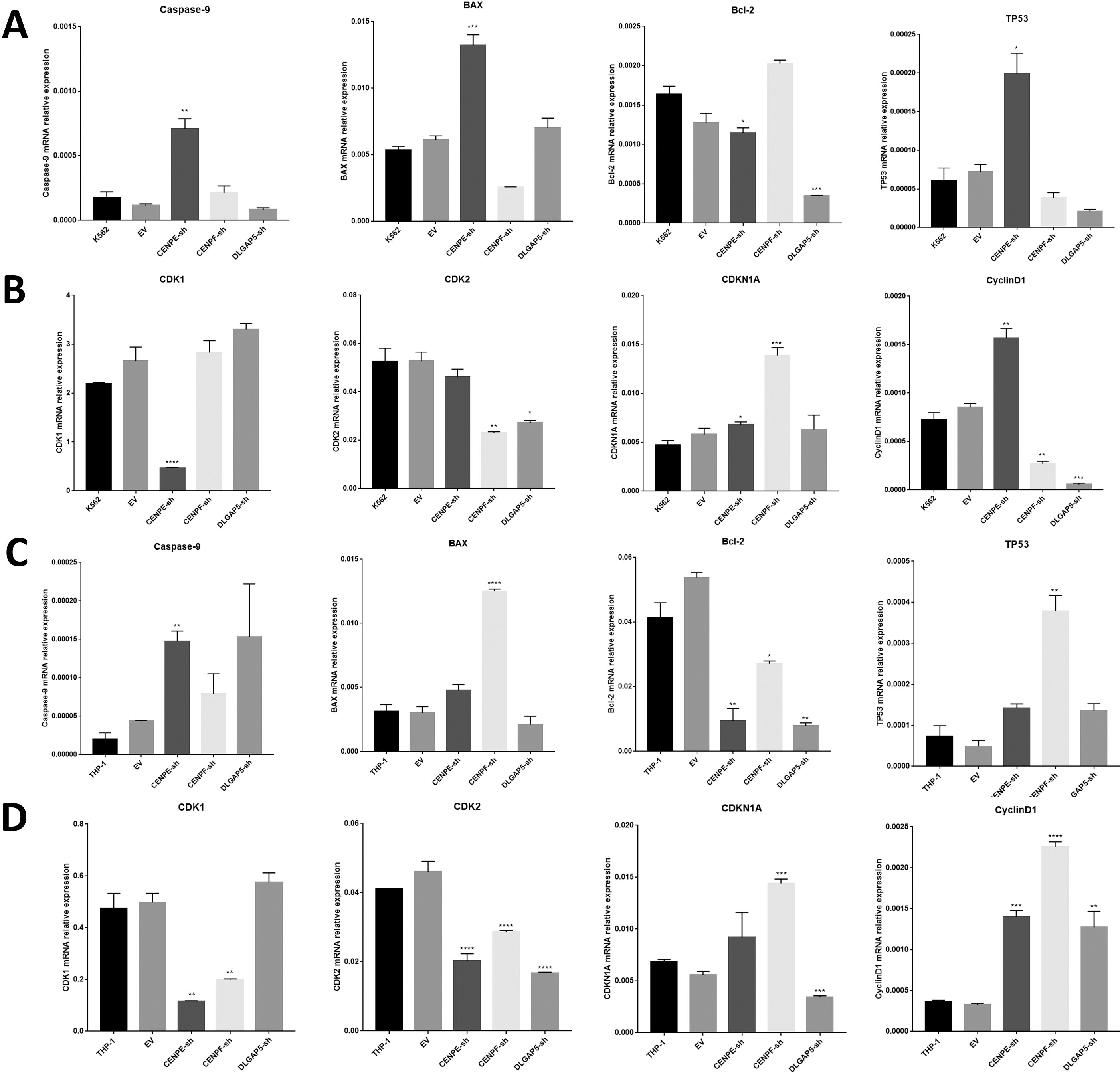
In conclusion, the GO and KEGG pathway analysis revealed that in R/R-AML and S-AML patients, the upregulated DEGs were mainly enriched in cell cycle related processes and pathways, while the downregulated DEGs were enriched in immune-associated pathways.
The Venn diagram analysis of the upregulated DEGs in “R/R-AML vs de novo AML patients” and “de novo AML vs Healthy control” revealed a total of 1 overlapping genes: FAR2; the venn diagram analysis of the upregulated DEGs in “S-AML vs de novo AML” and “de novo AML vs Healthy control” found 19 overlapping genes: DNMT3B, CCNA2, DLGAP5, CENPE, CENPF, KIF15, ASPM, KIF11, BUB1, NCAPG, CCNB2, HMMR, CEP55, GPSM1, UBE2T, CDCA8, SMC4, KIAA1524, PSMC3IP: Healthy control
CENPE, CENPF and DLGAP5 are highly overexpressed in R-AML patients
After deleting the 10 patients who lacked follow-up time, the clinical parameters of 141 R-AML patients from TCGA were analyzed (Table 5). Among them, 78 patients (55.3%) were male, 63 were female (44.7%), and the median age was 51 (21–88 years). Thirty-four patients received prior treatment, and 13 patients had a history of malignant tumors before diagnosis. Fifty-four patients remained alive (38.3%), and 87 patients died (61.7%) at the time of data update (2019). The 5-year survival of the 141 R-AML patients was analyzed by age, gender, with/without prior treatment and whether they had previously combined malignancies. By survival analyses, we found that among all these factors, age had the greatest prognostic significance (Fig. 4A) (Table 5). In patients aged
Knockdown of CENPE, CENPF, and DLGAP5 inhibited the growth of myeloid leukemia cells
To evaluate the biological function of CENPE, CENPF, and DLGAP5 in myeloid leukemia cells, we knocked down the expression of CENPE/CENPF/DLGAP5 in K562 and THP-1 cell lines by shRNAs. CENPE, CENPF, and DLGAP5 interference effectively downregulated gene expression at the mRNA level (
Knockdown of CENPE, CENPF, and DLGAP5 induced cell apoptosis and cell cycle arrest
The caspase-9 gene plays a central role in apoptosis [34]. Bcl-2 is an anti-apoptotic member, and BAX has a pro-apoptotic effect [35]. The TP53 gene is a tumor suppressor gene that can encode a tumor suppressor protein that leads to cell cycle arrest and apoptosis [34]. The transcription of the CDK1 gene usually indicates the M phase transition of the cell cycle, and transcription of the CDK2 gene prompts the G1/S phase transition of the cell cycle [36]. Cyclin D1 is a protein required for the progression of the G1/S transition and entering the S phase [37]. CDKN1A is a negative regulator of the cell cycle. It can bind to CDK or cyclin, causing the cell cycle to be blocked in the G1 phase [38].
To further investigate the effect of CENPE, CENPF, and DLGAP5 on the apoptosis and the cell cycle processes of leukemia cells, RT-PCR of cell apoptosis and cell cycle genes was performed. In K562 cells, CENPE knockdown significantly upregulated the expression of Caspase-9, BAX and TP53, while the expression of Bcl-2 decreased (
Taken together, after shRNA was used to interfere with the expression of CENPE, CENPF and DLGAP5 mRNA, the proliferation of leukemia cells was inhibited and apoptosis increased, mainly caused by the cell cycle M or G1/S phase block.
Discussion
The treatment of relapsed and/or refractory AML has been one of the great challenges in hematology for decades. The mechanism of drug resistance and relapse of AML has not been fully elucidated. AML patients usually show uncontrolled proliferation of myeloid progenitor cells. Genetic instability plays a significant role in the development and progression of AML. Mitotic flaws cause aberrant chromosomes and contribute to genetic instability [4], which leads to cell cycle arrest and drug resistance. Right mitotic control is a requirement to ensure that the genetic material is equally distributed into the two daughter cells [39]. The most recent studies have demonstrated that knockdown of CENPE expression in THP-1 cells caused a reduction in cell number and proliferation and reversed Ara-C resistance [14]. In this study, we found that in AML patients, mitosis cell cycle pathway and mitosis related genes were upregulated in chemoresistant AML. Based on the results of chemoresistance AML RNA-seq data, we explored the potential roles of mitotic cell cycle-associated genes CENPE, CENPF, and DLGAP5 in promoting leukemia cell growth and predicting poor prognosis and chemotherapy resistance in AML patients.
CENPE is a centromere-related protein that participates in the separation of chromatids, chromosomal arrangement, and the regulation of spindle checkpoints during mitosis [42, 43]. Silencing, deletion or abnormal expression of CENPE can lead to mitotic arrest, abnormal chromosome segregation, and even cell death [6, 8, 15, 18]. CENPF is a protein associated with chromosomal segregation during mitosis [16]. Knockdown of CENPF can inhibit the assembly of chromosome during mitosis, affect the segregation of chromosomes, and block cell cycle progression [19, 20]. CENPF is abnormally expressed in a variety of cancers and participates in cancer onset and development, including AML. Upregulated CENPF has been identified as a cell proliferation marker for a series of solid tumors and lymphoma [22, 23]. DLGAP5 is mainly involved in regulating the formation and movement of the spindle in mitosis [24, 25, 26]. Recent studies have shown that DLGAP5 is significantly highly expressed in various kinds of solid tumors [28, 29, 30]. More importantly, DLGAP5 expression is reported to be closely related to AML cell clone formation and stem cell differentiation [32, 33]. However, the role of mitosis-related genes CENPE, CENPF and DLGAP in chemotherapy resistance in AML is unclear. Thus, it is of great importance to explore the role and mechanisms of these genes in refractory and relapsed AML.
In recent years, RNA-seq technology has been widely used to screen DEGs related to the onset and evolution of various types of cancers, and to discover possible biomarkers and therapeutic targets [40, 41]. In our study, RNA-seq was carried out in chemotherapy resistant R/R-AML, S-AML, de novo AML patients and healthy controls. Furthermore, R-AML and primary AML RNA-seq datasets were downloaded from TCGA and GEO databases. Bioinformatic analysis was conducted to reveal the different gene expression profiles in R/R-AML, S-AML, R-AML compared with de novo/primary AML. GO and KEGG pathway analysis revealed that in R/R-AML and S-AML patients, we found that the upregulated DEGs were meaningfully enriched in processes and pathways related to the cell cycle. Venn diagram analysis of chemoresistant AML, de novo AML, and healthy controls identified the most upregulated DEGs (including CENPE, CENPF, and DLGAP5) in chemoresistance AML. Clinical data in enlarged samples obtained from TCGA and GEO database showed that the expression of CENPE, CENPF and DLGAP5 was significantly higher in R-AML. It is worth noting that the expression of CENPE/CENPF/DLGAP5 in R-AML patients who died was slightly higher. Further analysis of the correlation found that CENPE, CENPF and DLGAP5 all showed
We further explored the roles of CENPE, CENPF and DLGAP5 by using shRNA interference in K562 and THP-1 cells. After shRNA interference of the 3 genes, the proliferation of myeloid leukemia cells was inhibited, and cell apoptosis significantly increased. In K562 cells, the CCK-8 assay at 96 h showed that the proliferation of cells significantly decreased after CENPE and DLGAP5 knockdown. In THP-1 cells, the CCK-8 assay at 96 h showed that the proliferation of cells significantly decreased after CENPE, CENPF and DLGAP5 knocked down. Taken together, these results indicated that knockdown of CENPE, CENPF, and DLGAP5 could inhibit the growth of myeloid leukemia cells in vitro. After interfering with the expression of CENPE, CENPF and DLGAP5 mRNA, the proliferation of leukemia cells was inhibited. The apoptosis related genes Caspase-9, BAX, and TP53 were increased, while bcl-2 was decreased. Moreover, the CDK1, CDK2, and CDKN1A genes were downregulated after CENPE, CENPF, and DLGAP5 knockdown, mainly caused by cell cycle M or G1/S phase blockade. The conclusion of CENPE is consistent with the results of previous studies. In this study, we confirmed for the first time that CENPF interference can inhibit the proliferation of AML cells and participate in the regulation of AML progression. Furthermore, we confirmed through bioinformatic analysis and in vitro experiments that DLGAP5 interference can inhibit the proliferation activity of AML cells, and this result is also consistent with the above research results.
In conclusion, in this study, bioinformatics analysis and in vitro experiments were performed to explore the role of the mitotic cell cycle genes CENPE, CENPF, and DLGAP5 in refractory and/or relapsed AML. For the first time, we provide strong evidence that the expression of CENPE, CENPF, and DLGAP5 is associated with unfavorable prognosis and chemotherapy resistance in AML patients. Thus, direct targeting of mitotic-related genes might be a novel strategy for chemotherapy-resistant AML treatment.
Author contributions
Conception: Mingyue Shi, Kai Sun, Yuqing Chen.
Interpretation or analysis of data: Mingyue Shi, Honggang guo, Yanliang Bai.
Preparation of the manuscript: Mingyue Shi, Honggang guo, Yanliang Bai.
Revision for important intellectual content: Junwei Niu, Xiaona Niu.
Supervision: Yuqing Chen, Kai Sun.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203170.
sj-pdf-1-cbm-10.3233_CBM-203170.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-203170.pdf
Footnotes
Acknowledgments
This study was partially supported by the National Natural Science Foundation of China (No. 81971508, No. 81471589 and No. 81273259), the Health Bureau of Henan Province, P.R. China (No. 201201005) and the foundation and frontier research grant of Henan provincial science and technology bureau, P.R. China (No.142300410078). and Sansheng Chunya Funds for Young Scientists.
Conflict of interest
The authors report no conflicts of interest in this work.
