Abstract
BACKGROUND:
Cervical cancer (CC) is a malignant tumor threatening women’s health. Replication factor C (RFC) 5 is significantly highly expressed in CC tissues, and the immune microenvironment plays a crucial role in tumor initiation, progression, and metastasis.
OBJECTIVE:
To determine the prognostic role of RFC5 in CC, analyze the immune genes significantly associated with RFC5, and establish a nomogram to evaluate the prognosis of patients with CC.
METHODS:
High RFC5 expression in patients with CC was analyzed and verified through TCGA GEO, TIMER2.0, and HPA databases. A risk score model was constructed using RFC5-related immune genes identified using R packages. Combining the risk score model and clinical information of patients with CC, a nomogram was constructed to evaluate the prognosis of patients with CC.
RESULTS:
Comprehensive analysis showed that the risk score was a prognostic factor for CC. The nomogram could predict the 3-year overall survival of patients with CC.
CONCLUSIONS:
RFC5 was validated as a biomarker for CC. The RFC5 related immune genes were used to establish a new prognostic model of CC.
Introduction
Cervical cancer (CC), the second most prevalent among gynecological tumors after ovarian cancer, seriously threatens women’s health [1]. In 2018, more than 800,000 cases of CC were reported, with approximately 30% of cases occurring in China and India [2]. The cause of CC development is believed to be persistent infection with high-risk human papillomavirus [3]. Vaccination against human papillomavirus can reduce the incidence of CC; however, in 2020, the global incidence of CC was 13.3 per 100,000 women, with 604,127 new cases [4]. CC is characterized by a 5-year overall survival (OS) of approximately 50–65%, a high recurrence rate, and a poor prognosis [5, 6, 7]. The prognosis of CC is not related to the patient age, T stage, or cancer grade [8, 9, 10]. Therefore, new prognostic models are essential for the detection, prognosis, and treatment of CC.
Replication factor C (RFC) is a DNA-binding protein composed of five subunits: RFC1, RFC2, RFC3, RFC4, and RFC5 [11, 12]. RFC, first extracted from human CC HeLa cells [13], is closely related to tumor cell proliferation, invasion, and metastasis [14]. RFC is involved in DNA replication [15], and RFC5 is necessary for RFC when offloading proliferating cell nuclear antigens from circular DNA [16]. Forkhead box M1 interacts with RFC5 to transcriptionally activate RFC in glioma [17]. RFC5 is involved in the process of DNA damage as well as cell cycle and cancer progression [18]. In acute myeloid leukemia, a correlation between RFC5 expression and immune cells was observed [19]; thus, RFC5 is considered a biomarker for acute myeloid leukemia [20].
RFC5 expression also significantly differed between CC and normal cervical tissues [21]. Studies have shown that CC has an active immune microenvironment [22, 23]. B cells show a certain predictive effect on the efficacy of CC immunotherapy [24]; fibroblasts associated with cancer have a certain impact on the prognosis of CC [25]. Immune cells are related to the expression of RFC in tumors [26].
This study aimed to establish a prediction model of RFC5-related immune genes for CC. RFC5 expression in CC tissues was first verified in public databases, and the specificity and prognostic value of RFC5 in CC diagnosis were analyzed using statistical methods. Immune cells and immune genes that interact with RFC5 in CC were then analyzed, and risk score models were constructed using these immune genes associated with CC prognosis. Finally, a nomogram was constructed to evaluate the prognosis of patients with CC.
Materials and methods
Data collection
The expression data of messenger RNA (mRNA) and clinical characteristics of CC cases were downloaded on May 9, 2022, from The Cancer Genome Atlas (TCGA;
Expression, tumor immune estimation resource (TIMER), and human protein atlas (HPA) analyses
RFC5 expression was analyzed in normal and CC tissues from TCGA using R packages BiocManager and limma. Next, RFC5 expression was verified in the gene expression profiling datasets GSE7410, GSE7803, GSE9750, and GSE63514 using the Wilcoxon rank sum test in R. TIMER2.0 (
Survival analysis and receiver operating curve (ROC) curves
The relationships between the different RFC5 expression groups and the OS rate of patients with CC were analyzed using the R packages BiocManager, limma, survival, and survminer to judge the prognostic value of RFC5 in CC. The normalized RNA sequencing (RNA-seq) data of 306 CC samples and 13 normal cervical samples were obtained from the University of California Santa Cruz XENA database (
Distribution of immune cells in different RFC5 expression groups
The RNA-seq data from TCGA in Fragments Per Kilobase per Million were log2 transformed, and normal sample data were removed. Next, a gene set variation analysis (GSVA) was performed in the GSVA R package to study the characteristics of the interaction between RFC5 expression and immune cells in CC [29]. Significance was set at
Relationship between RFC5 expression and immune genes
The expression of immune genes in tumors was analyzed in TISIDB (
Protein-protein interaction (PPI), Gene Ontology (GO), and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis
A PPI network between immunoinhibitors and immunostimulators significantly associated with RFC5 was built in the STRING database (
Screening the immune prognosis-related genes and establishing a prognostic model
Univariable Cox regression analysis was used to screen the immunoinhibitors and immunostimulators related to CC prognosis using the R packages limma and survival and
where
The risk scores of the CC samples in TCGA database were calculated, and the samples were divided into high- and low-risk groups according to the median risk score. Survival differences were compared between the high- and low-risk groups using R packages survival and survminer. The risk curve, survival status map, and heat map of immunoinhibitors and immunostimulators were drawn in R.
Independent prognostic analysis
The prognostic value of risk scores was evaluated using univariate and multivariate Cox regression analyses, and then the diagnostic value of risk scores was evaluated based on the time-dependent ROC curve using R packages survival, survminer, and timeROC.
Prediction and nomogram construction
A nomogram was established using the risk score and N and T stages to predict the 1-, 2-, and 3-year OS in patients with CC using R package rms according to the clinical characteristics of CC patients in TCGA. Next, a calibration curve was used to evaluate the accuracy of the nomogram predictions.
Statistical analysis
RFC5 expression between normal and cervical cancer tissues from The Cancer Genome Atlas database. 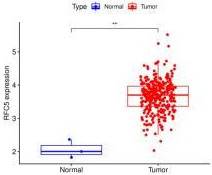
R packages BiocManager, limma, survival, survminer, and GSVA were used for differential expression, survival, and immune cell infiltration analyses in CC. The univariable and multivariate Cox regression analyses were used to screen the genes related to CC prognosis using R packages limma, glmnet, survival, and survminer. Statistical significance was set at
Expression of RFC5 in CC
Pan-cancer expression of RFC5, analyzed using the Tumor Immune Estimation Resource 2.0 database. 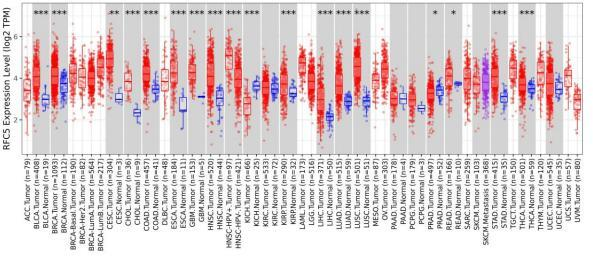
RFC5 expression in the Gene Expression Omnibus database. RFC5 expression in GSE7410 (A), GSE7803 (B), GSE9750 (C), and GSE63514 (D) datasets. 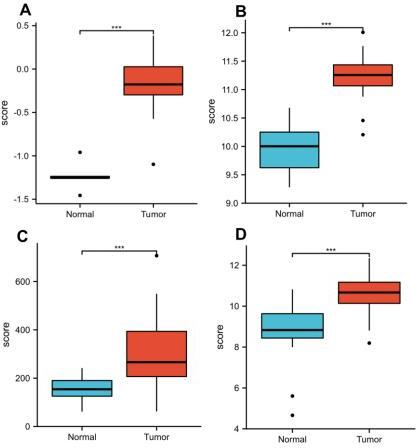
Protein levels of RFC5 in normal cervical and cervical cancer tissues from the Human Protein Atlas database using immunohistochemical staining. 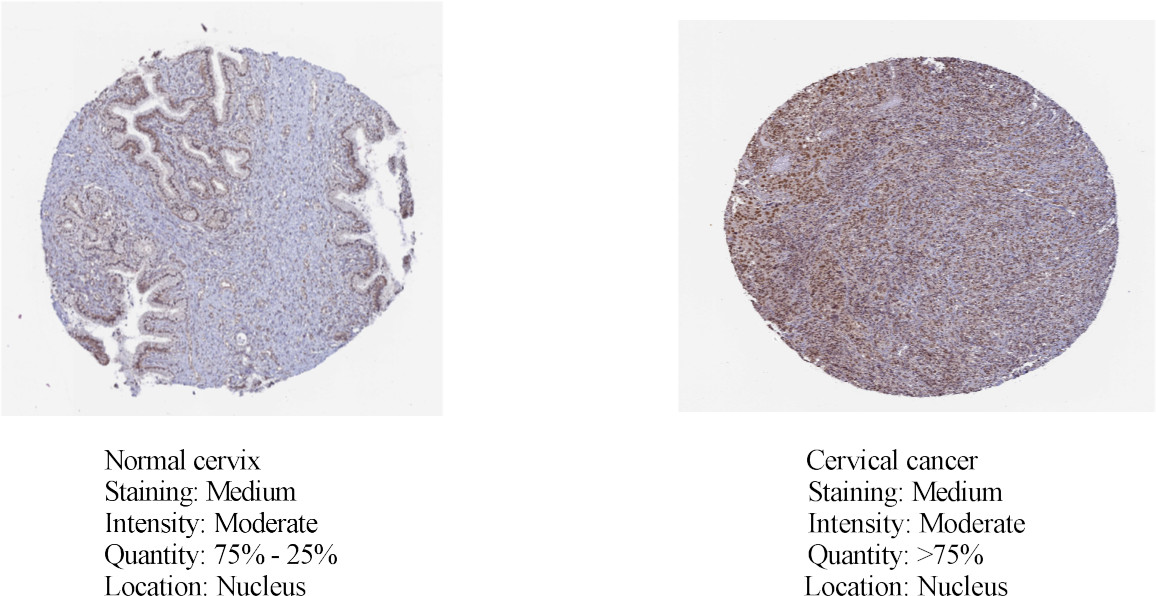
Verification of prognostic and diagnostic values of RFC5 in CC. Correlation between RFC5 expression and prognosis of CC (A). Receiver operating characteristic curve used to estimate the diagnostic value of RFC5 in CC. (B). CC: cervical cancer; AUC, area under the curve; HR, hazard ratio; TPR, true positive rate; FPR, false positive rate. 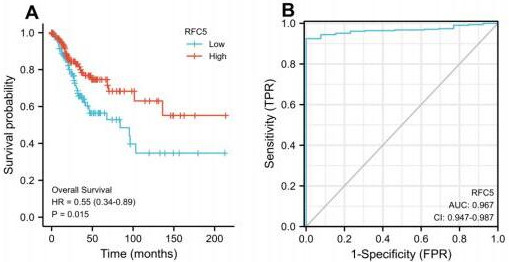
The clinical information and mRNA data of 306 patients with CC and 3 normal patients were obtained from TCGA database. The expression of RFC5 was significantly increased in CC (Fig. 1). RFC5 expression in various cancers was analyzed using the TIMER2.0 database; RFC5 showed significantly high expression in several cancers (Fig. 2). By analyzing the transcriptome data of the GSE7410, GSE7803, GSE9750, and GSE63514 datasets, the significantly high expression of RFC5 in CC was verified (Fig. 3A–D). The immunohistochemical results from the HPA database showed that RFC5 was located in the nucleus of normal cervical tissue and CC tissue (medium-intensity staining observed). However, the number of positive cells was significantly higher in CC tissue than in normal cervical tissue (Fig. 4).
RFC5 expression is associated with the prognosis of CC (
Distribution of immune cells
Relationship analysis of RFC5 expression and populations of cytotoxic T cells, DCs, iDCs, macrophages, neutrophils, CD56dim cells, pDCs, T helper cells, Tcm cells, Th1 cells, and Th2 cells. DCs: dendritic cells; iDCs: immature DCs; pDCs: plasmacytoid DCs; Tcm: central memory T cells; Th1: T helper 1 cells; NK, natural killer.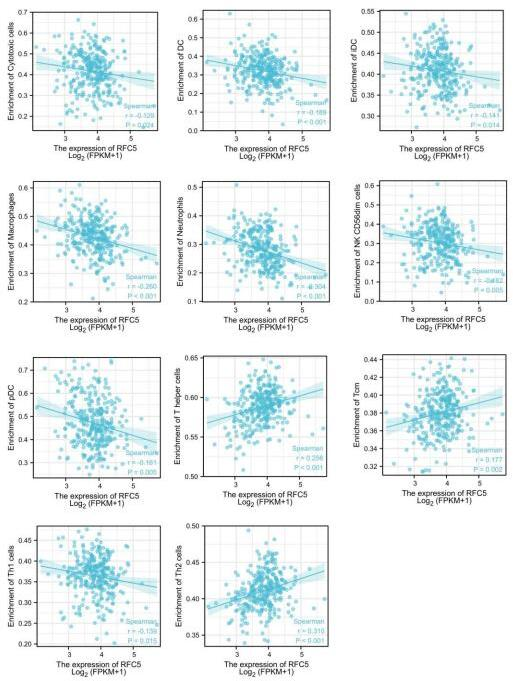
Relationship analysis of immune cells in RFC5 high- and low-expression groups; 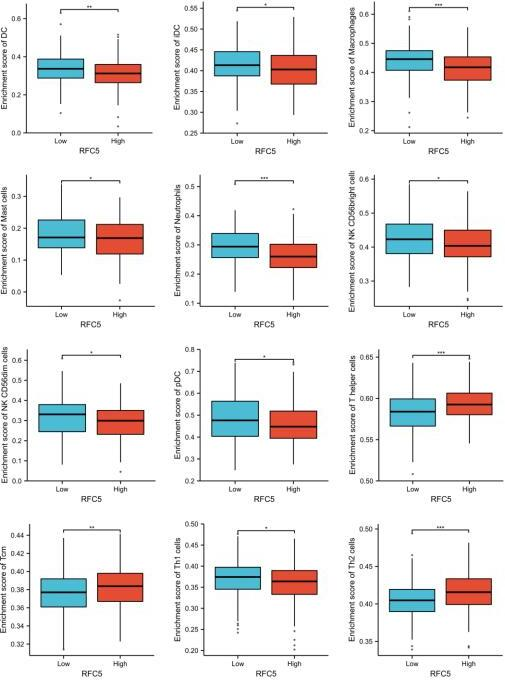
Survival analysis of patients with different populations of immune cells and RFC5 expression. Survival analysis of patients with CC with T cell gamma delta (A), myeloid dendritic cells (B), cancer-associated fibroblasts (C), and myeloid-derived suppressor cells (D). 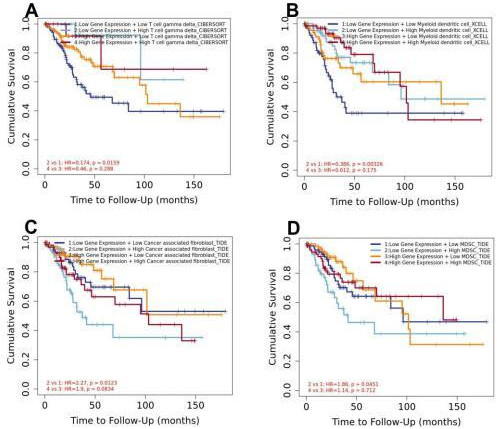
Analysis of immune cell distribution in CC showed that RFC5 expression was negatively correlated with the populations of cytotoxic cells, dendritic cells (DCs), immature DCs (iDCs), macrophages, neutrophils, natural killer (NK) CD56dim cells, plasmacytoid DCs (pDCs), and T helper (Th) 1 cells and positively correlated with those of central memory T (Tcm) cells, Th2 cells, and Th cells (Fig. 6).
Correlation analysis of RFC5 expression and immunoinhibitors.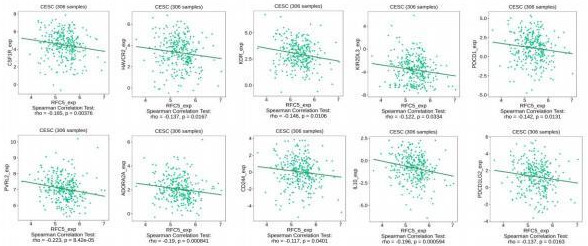
Correlation analysis of RFC5 expression and immunostimulators.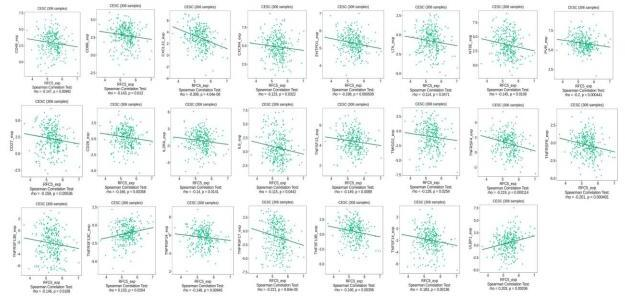
According to the median expression of RFC5 in CC samples in TCGA database, the samples were divided into high- and low-expression groups. A certain relationship was found to exist between RFC5 expression and immune cells. In the RFC5 high-expression group, Th, Tcm, and Th2 cell populations were significantly increased. The populations of macrophages, mast cells, DCs, iDCs, neutrophils, NK CD56bright cells, pDCs, NK CD56dim cells, and Th1 cells were significantly increased in the RFC5 low-expression group (Fig. 7). The impact of immune cell distribution on the prognosis of patients with CC was reassessed in the TIMER database. In the RFC5 low-expression group, high numbers of T cell gamma delta and myeloid DCs were positively correlated with CC prognosis, whereas high numbers of cancer-associated fibroblasts and myeloid-derived suppressor cells were negatively correlated with CC prognosis. Immune cell infiltration was not associated with CC prognosis in the RFC5 high-expression group (Fig. 8).
TISIDB analysis found that the expression of RFC5 was correlated with the immune genes of patients with CC. The immunoinhibitors such as CSF1R (rho
PPI network, GO and KEGG enrichment analysis
Univariate Cox regression analysis of immunoinhibitors and immunostimulators related to RFC5
Univariate Cox regression analysis of immunoinhibitors and immunostimulators related to RFC5
Interaction and functional analysis of the 10 immunoinhibitors and 23 immunostimulators. A PPI network of 10 immunoinhibitors and 23 immunostimulators was constructed in the STRING database (A). Gene Ontology (B) and Kyoto Encyclopedia of Genes and Genomes enrichment analysis (C) of 10 immunoinhibitors and 23 immunostimulators. PPI: protein–protein interaction; FDR, false discovery rate.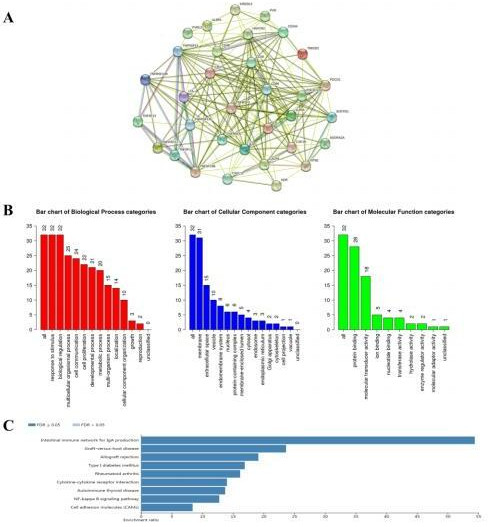
Multivariate independent prognostic analysis
Univariate (A) and multivariate (B) Cox regression analyses of immunoinhibitors and immunostimulators in patients with CC. 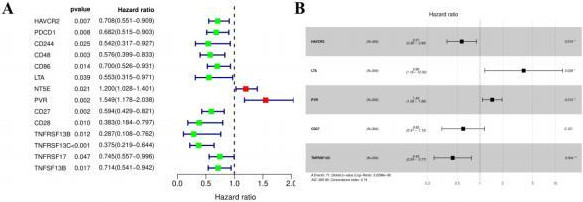
A PPI network of 10 immunoinhibitors and 23 immunostimulators was constructed, containing 33 nodes and 208 edges, in the STRING database (
Fourteen immunoinhibitors and immunostimulators were found to be related to the prognosis of CC through univariate Cox regression analysis (Table 1; Fig. 12A). Among them, NT5E and PVR, with a hazard ratio
Patients with CC were divided into low- and high-risk groups (A). Survival status of patients with CC with different risk scores (B). Expression of prognostic model genes in patients with CC with different risk scores (C). Survival analysis of patients with CC in low- and high-risk groups (D). CC: cervical cancer.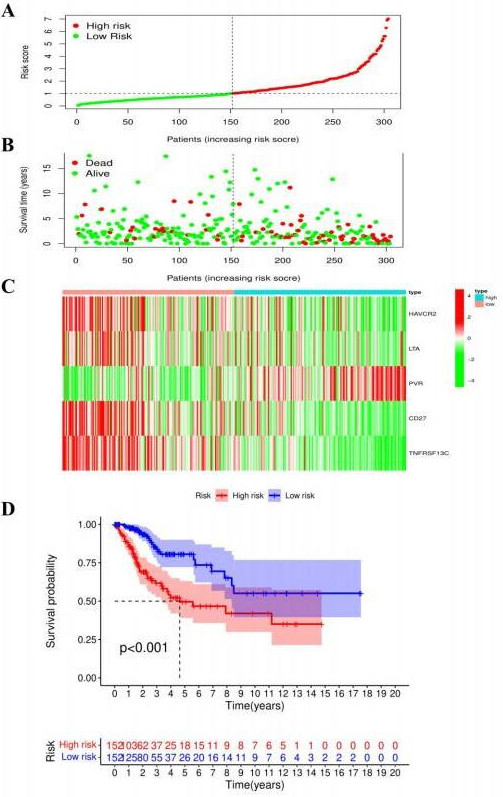
Risk score model diagnostic and prognostic analyses. Univariate Cox regression analysis of the risk score factor (A). Multivariate Cox regression analysis of the risk score factor (B). Receiver operating characteristic curve and value of AUC of clinical characteristics prognostic signature (C). AUC: area under the curve.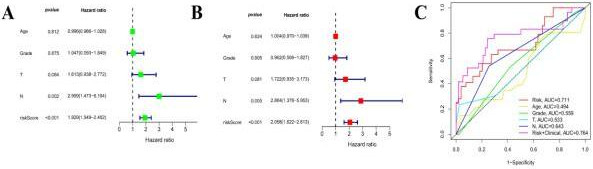
Nomogram and calibration curve for patients with CC. The nomogram predicts the OS of patients with CC (A). The calibration curve shows the accuracy of the predicted 3-year OS of the nomogram (B). CC: cervical cancer; OS: overall survival.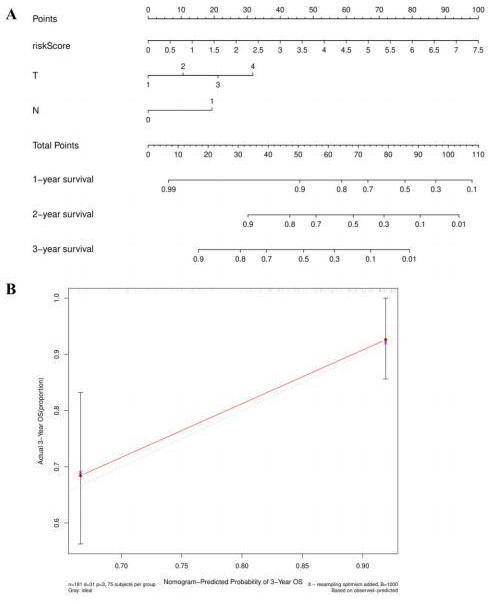
The risk score of patients with CC in TCGA database was calculated according to the risk score calculation formula. Then, according to the median risk score, the patients with CC were divided into high- and low-risk groups (Fig. 13A). The survival status of patients with CC suggested that the higher the risk score, the higher the mortality of patients (Fig. 13B). The heatmap shows the expression of HAVCR2, LTA, PVR, CD27, and TNFRSF13C in two different groups of CC; the expression of LTA and PVR was positively correlated with the risk score (Fig. 13C). Survival analysis showed that the low-risk group had a better OS (
N stage and risk score were found to be independent prognostic factors for patients with CC through univariate and multivariate Cox regression analysis (
Nomogram
A nomogram was constructed based on the combination of clinical information of CC patients in the TCGA database and risk scores to determine the prognosis of CC patients (Fig. 15A). According to the patient’s clinical characteristics and total points of risk scores, the 1-, 2-, and 3-year OS of patients with CC could be predicted. In the calibration curve, the red and grey curve shows the predicted and actual 3-year OS of patients with CC in TCGA database, respectively (Fig. 15B).
Discussion
Although the number of patients with CC has decreased with the use of CC vaccines, CC continues to threaten women’s health worldwide. Thus, there has been an increase in the number of in-depth CC-related research studies. Many studies are devoted to identifying biomarkers of CC, and research has revealed that the immune environment has a great impact on CC. In the present study, we explored immunoinhibitors and immunostimulators related to RFC5 in CC, providing a new prognostic model for CC.
Owing to the biological function of RFC5 in tumors and its high expression in CC, it is considered a new predictive gene and therapeutic target for CC [21, 31]. The present study analyzed RFC5 expression in normal and CC tissues; consistent with previous results, RFC5 was highly expressed in CC. Next, RFC5 expression in CC was also verified in the GEO database. Through the survival and ROC curve analyses, it was concluded that RFC5 has diagnostic and prognostic value in CC. Immune cells and genes associated with RFC5 were further explored. Increased populations of T helper, Tcm, and Th2 cells were positively correlated with RFC5 expression; these cell populations were markedly increased in the RFC5 high-expression group. Th cells can directly activate cytolytic T lymphocytes, and tumor antigen-specific B cells and can kill tumor cells [32]. Th cell populations were correlated with OS in patients with advanced CC after chemoradiotherapy and were considered predictors of survival in these patients [33].
After antigen activation, Tcm cells produce T cells with long-term memory and can home to lymph nodes to receive antigen re-stimulation. Tcm cells have a strong anti-tumor ability [34], which can be reinfused into the body to directly kill tumor cells [35]. Furthermore, they have self-renewal and replication abilities and can survive for long periods in the body, mediating long-term anti-tumor effects [36]. Th2 cells and the type 2 immune response of Th2 cells have both anti-tumor and tumor-promoting effects, and the effect that dominates depends on the type and stage of the tumor [37].
The expression of lymphotoxin alpha (LTA) and poliovirus receptor (PVR) was positively correlated with the risk score and negatively correlated with the prognosis of patients with CC. The expression of HAVCR2, CD27, and TNFRSF13C reduced the risk score and was revealed as a protective factor for the prognosis of patients with CC. PVR is a multifunctional cell surface protein [38], which is related to immunosuppression by immune cells [39]. In the present study, PVR was a high-risk factor for the prognosis of CC. PVR downregulation can inhibit the killing activity of NK cells against tumor cells [40]. PVR was overexpressed in melanoma [41], colorectal cancer [42], and lung adenocarcinoma [43]. In non-small cell lung and bladder cancers, PVR expression was an indicative factor in the poor prognosis of patients [44, 45]. PVR plays an important role in regulating the activity of NK cells and T cells to eliminate tumors, providing a possibility for targeted therapy of tumors [39]. LTA is a pro-inflammatory factor produced by T lymphocytes, mainly involved in inflammation and immune responses [46]. LTA polymorphisms are significantly associated with the occurrence of cancer and are associated with genetic susceptibility to CC [47, 48]. PVR and LTA may serve as new targets in the development of CC treatment.
According to the risk score formula, the risk score of patients with CC in TCGA database was calculated, and as shown in the survival time chart, the higher the risk score, the greater the probability of poor survival. As shown in the risk heatmap, the high expression of PVR and LTA of patients with CC implies poor OS, whereas the expression of HAVCR2, CD27, and TNFRSF13C was negatively associated with the risk of patients with CC, suggesting that PVR and LTA were high-risk factors and HAVCR2, CD27, and TNFRSF13C were low-risk factors. Prognostic, univariate, and multivariate independent prognostic analyses indicated that risk scores could be used as independent prognostic factors for patients with CC. The AUC value of ROC curves of risk scores combined with clinical features was 0.764, which was significantly higher than those obtained using only clinical features. Therefore, the model and clinical characteristics can be considered simultaneously when predicting the survival of patients, yielding more accurate results. Combining clinical characteristics with the risk score model, a nomogram was created to predict the survival of patients with CC. Comparing the 3-year survival curve of patients with CC in TCGA database to the 3-year survival curve predicted by the nomogram, the two curves were very similar, suggesting the good predictive performance of the nomogram. Nonetheless, this prognosis is only based on data from TCGA database; additional experimental and clinical data are required for further verification.
Conclusions
This study analyzed the expression of RFC5 in patients with CC and validated it as a biomarker for CC. By exploring the immune genes associated with RFC5 and those associated with the prognosis of CC, a risk score model was developed. The risk score model was combined with patient clinical characteristics to create a nomogram that predicted the survival of patients with CC. In future clinical work, the application of the nomogram to the evaluation of the prognosis of patients with CC will facilitate the selection of treatment options. To apply the nomogram to clinical practice, a large number of clinical trials are required. Nevertheless, this study has some limitations, such as the lack of experiments to verify the expression of RFC5 in CC tissues and clinical follow-up, which is needed to understand the correlation between the nomogram and the survival time of patients with CC. In future research, experimental verification and follow-up of patients with CC are essential.
Footnotes
Acknowledgments
The authors would like to thank TCGA, GEO, TIMER, HPA, TISIDB, WebGestalt, and SPRING databases for providing free research resources. This work was supported by the Department of Science and Technology of Yunnan Province (approval No: 202001BA07001133), Liangshan Prefecture Science and Technology Program Key R&D Project (approval No: 21ZDYF0173), Medical Discipline leader of Yunnan Provincial Health and Family Planning Commission (approval No: D-2017057), Department of Education of Yunnan Province, Key Laboratory of Research in Colleges and Universities of Yunnan Province; and Yunnan Provincial Department of Education, Obstetrics and Gynecology, graduate tutor team (2020.01–2023.12).
Conflict of interest
The authors declare no potential conflict of interest related to the research, authorship, and/or publication of this manuscript.
Author contributions
Conception: Yuanyuan Zhang and Guangming Wang.
Interpretation or analysis of data: Huaqiu Chen and Huanyu Xie.
Preparation of the manuscript: Huaqiu Chen, Huanyu Xie.
Revision for important intellectual content: Yuanyuan Zhang and Guangming Wang.
Supervision: Yuanyuan Zhang and Guangming Wang.
