Abstract
Background
Gamma delta (γδ) T cells play dual roles in human tumors, with both antitumor and tumor-promoting functions. However, the role of γδT cells in HPV-infected cervical cancer is still undetermined. Therefore, we aimed to identify γδT cell-related prognostic signatures in the cervical tumor microenvironment.
Methods
Single-cell RNA-sequencing (scRNA-seq) data, bulk RNA-seq data, and corresponding clinical information of cervical cancer patients were obtained from the TCGA and GEO databases. The Seurat R package was used for single-cell analysis, and machine learning algorithms were used to screen and construct a γδT cell-related prognostic signature. Real-time quantitative PCR (RT-qPCR) was performed to detect the expression of prognostic signature genes.
Results
Single-cell analysis indicated distinct populations of γδT cells between HPV-positive (HPV+) and HPV-negative (HPV-) cervical cancers. A trajectory analysis indicated γδT cells clustered into differential clusters with the pseudotime. High-dimensional Weighted Gene Co-expression Network Analysis (hdWGCNA) identified the key γδT cell-related gene modules. Bulk RNA-seq analysis also demonstrated the heterogeneity of immune cells, and the γδT-score was positively associated with inflammatory response and negatively associated with MYC stemness. Eight γδT cell-related hub genes (GTRGs), including ITGAE, IKZF3, LSP1, NEDD9, CLEC2D, RBPJ, TRBC2, and OXNAD1, were selected and validated as a prognostic signature for cervical cancer.
Conclusion
We identified γδT cell-related prognostic signatures that can be considered independent factors for survival prediction in cervical cancer.
Introduction
Human papillomavirus (HPV) is a small-double-stranded circular DNA virus with a tropism for squamous epithelium and has identified more than 70 HPV types [12, 13]. HPV infection leads to HPV-associated cancers in both men and women, such as cervical cancer, head and neck cancers (HNCs) and anogenital cancers (vaginal, penial, and anal cancers) [16].
Cervical cancer is one of the most frequently diagnosed cancers in women and ranks as the fourth leading cause of cancer-related death worldwide. In 2020, nearly half of the women diagnosed with cervical cancer succumbed to the disease. 1 The incidence and mortality rates of cervical cancer have declined in most areas of the world over the past few decades, but these rates have increased in China.2-4 Cervical cancer ranks as the sixth most common cancer and seventh leading cause of cancer-related death in China, with 591, 688 cases and 435, 860 deaths reported in 2020. 5 A retrospective analysis has revealed that the mortality rate of cervical cancer is rapidly rising in younger women in urban China, whereas it is declining in all age groups in rural China. 6 Several reports have found that the incidence of cervical cancer in most regions of China exceeds the national average.7-9 Infection of HPV, human immunodeficiency virus (HIV) and Chlamydia trachomatis, smoking, the higher number of childbirths, and long-term use of oral contraceptives are considered risk factors for cervical cancer. 10 Most cervical cancer cases are caused by HPV16 and HPV18 infection, with HPV16 accounting for more than 50% of these cases [14, 15]. Although the testing and prevention of cervical cancer have significantly improved over the past decades, anogenital cancers remains more difficult to diagnose definitively. 11 Exploring the promising therapies for cervical cancer is imperative for further studies.
γδT cells are a unique subset of T cells characterized by the expression antigen receptors composed of gamma (γ) chain and delta (δ) chain. 12 γδT cells also reveal the dual roles in human tumors, possessing both antitumor and tumor-promoting functions. The role are associated with their potent cytotoxicity interferon-γ (INF-γ) production, and interleukin-17 (IL-17) production.12,13 γδT cells have recently garnered attention as an ideal tool for cancer immunotherapy due to their antitumor function and role in immune surveillance. 14 Additionally, γδT cell infiltration into tumors severs as a positive prognostic marker in many solid tumors and is associated with improved clinical outcomes.15-18 It has been found that γδT cells play key roles in persistent HPV infection and early-stage cervical carcinogenesis. 19 A study has reported that HPV16 oncoproteins induce recognition of the local epithelial-associated γδT cells subpopulations, which promote tumor aggression in uterine cervical squamous cell carcinoma (UCSCC). 20 γδT cells can be potentially used as a therapeutic strategy against HPV infection cervical cancer patients.
In the present study, we explored the correlation between γδT cells and HPV infection, identifying γδT cells-related signatures in cervical cancer associated with HPV infection using single-cell RNA-sequencing (scRNA-seq) and bulk RNA-seq data. Additionally, we established a prognostic signature related to γδT cells and validated it in both HPV-infected cervical cancer and head and HNSCC. Our findings provide the potential biomarkers for predicting the clinical outcomes of both HPV-infected and non-infected patients.
Methods
Data Collecting and Processing
The mRNA expression data and corresponding clinical information as well as single-cell RNA-seq data of cervical cancer (CC) and head and neck squamous cell carcinoma (HNSC) were obtained from the Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov/) and the Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/). The details of these data are found by accessing the links.
scRNA-Seq Data Processing
The scRNA-seq data set of two HPV+ and two HPV- CC samples were downloaded from the GEO database. “Seurat” (version 4.5.0) R package was used for scRNA-seq data processing. Cells with more than 1000 and less than 50,000 unique molecular identifiers (UMIs) in a single cell, less than 80% of ribosomal genes, and less than 10% of mitochondrial genes were included in the subsequent analysis. After quality control of the cells, “SCTransform” function was utilized for data normalization through linear regression analysis, considering the normalized expression level of each gene, total UMI count per cell, and cell cycle. “FindVariableFeatures” was used to identify the top 3000 high variable genes (HVGs). Next, principal component analysis (PCA) was conducted using the “RunPCA” function based on 3000 HVGs. The Louvain clustering method was applied for clustering, followed by visualization using Uniform Manifold Approximation and Projection (UMAP) by “RunUMAP” function. Differentially expressed gene markers in each cluster were identified though manual annotation, referencing common marker genes reported in the literature for tumor tissues.
Trajectory Analysis of γδT Cell Lineages
The “Monocle 2” package was used for single-cell trajectory analysis to discover the changing trend of cell states by performing the following steps, (i) Feature selection involved ranking with the top 1000 HVGs in cell lineage. (ii) Dimension reduction was conducted using the “reduceDimension” function with the parameters method = “DDRTree”. (iii) Trajectory construction was executed with “plot_cell_trajectory” function. (iv) Branch expression analysis modeling (BEAM) was used to identify genes that distinguish cells into branches, and the results were visualized using the“plot_genes_branched_heatmap” function. (v) The trend of gene expression with the pseudo-time was analyzed and visualized using the “plot_genes_in_pseudotime” function. The enrichment of gene ontology (GO) terms for the genes in each cluster was explored using the “ClusterProfilter” package.
Comparison of the Immune Characteristics Between HPV+ and HPV- Cervical Cancers
To unveil the tumor immune microenvironment of cervical cancer, the “IOBR” R package was used for immune cell infiltration analysis in this study. The immune-related gene set consisting of 22 types of immune cells was obtained from previous literature sources. 21 The different proportions of immunes between HPV+ and HPV- groups were determined using Wilcoxon’s test. We also downloaded the reference gene set about immune-related pathways from the MSigDB (https://www.gsea-msigdb.org/gsea/index.jsp). SsGSEA was performed using the “GSVA” R package to investigate the enriched pathways between HPV+ and HPV- groups. Moreover, ssGSEA is also employed for calculating the γδT score of each sample based on the expression of marker genes specific to γδT cells, including XCL1 FCER1G, XCL2, TYROBP, KLRC1, GNLY, B3GNT7, TRDC, NCAM1, and LAT2. The correlation between the γδT score and inflammatory response, as well as γδT score and MYC stemness, were analyzed.
Identification of The γδt Cell Relevant Hub Modules Based on A High-Dimensional Weighted Gene Co-Expression Network Analysis (hdWGCNA)
hdWGCNA is a WGCNA method tailored for high-dimensional transcriptomics data. It is utilized for identifying robust modules of interconnected genes and providing context for these modules through various biological knowledge sources. 22 In the present study, an “hdWGCNA” R package with Seurat was utilized to construct a hdWGCNA and identify the key modules relevant to γδT cells. Firstly, the Seurat object was set up for WGCNA before hdWGCNA processing. Secondly, the K-nearest neighbor (KNN) algorithm was used to identify similar progenitor cells and calculate their gene expression, facilitating the construction of a metacell expression matrix. Thirdly, the expression matrix was established, and the soft threshold was selected. Subsequently, an hdWGCNA was constructed using the “ConstructNetwork” function. Fourth, module eigengenes (MEs) were computed and harmonized using “ModuleEigengenes” and “Harmony” functions. Afterward, eigengene-based connectivity (kME) was determined using the “moduleConnectivity” function, and the genes in each module ranked were visualized by kME using the “plotkME” function. AUCell algorithm was used to calculate the gene score for the hub genes by kME for each module. The correlation between each module, or modules and cell states based on their hub gene scores was visualized using the “corrplot” R package. Finally, the “clusterProfiler” package in R was used to investigate the GO analysis for each module.
Development of A γδt Cell-Related Risk Score And Validation Of The γδt Cell-Related Risk Score In HPV + cancers
Based on the hub genes identified from the hdWGCNA, the prognostic value of the γδT cell-related hub genes (GTRGs) was estimated using patients from the TCGA cohort for subsequent analysis. Univariate Cox analysis was performed to identify the prognostic GTRGs associated with overall survival according to the cutoff value of less than 0.05. Afterward, the least absolute shrinkage and selection operator (LASSO) regression analysis was performed using the “glmnet” package in R to establish a prognostic gene signature by reducing redundant genes and obviating model overfitting.
23
Then, the risk score of each sample was calculated based on the following formula, risk score =
Specimens
A total of 12 cervical cancer patients at the First People’s Hospital of Yunnan Province (Kunming, China), were enrolled in this study between November 2023 and February 2024. All the patients included in the study had been diagnosed with cervical cancer, and none had received any preoperative treatment. The patients underwent surgical resection, and the tumor tissues and adjacent normal tissues were collected on the day of surgery. All samples were frozen in liquid nitrogen quickly and stored at −80°C. The protocol for collecting clinical samples was approved by the Ethics Committee of the First People’s Hospital of Yunnan Province (No. KHLL2023-KY196) on December 13, 2023, and the patients provided informed consent before samples were collected. The reporting of this study conforms to REMARK guidelines. 24
RNA Isolation and qRT-PCR Detection
Trizol reagent (TAKARA, Dalian, China) was used to isolate the total RNA from cells following the standard protocol, and complementary DNAs (cDNAs) were synthesized using PrimeScript™ FAST RT reagent Kit with gDNA Eraser (TAKARA, Dalian, China). To compare expression levels between groups, quantitative real-time PCR (qRT-PCR) was performed using an TB Green® Premix Ex Taq™ (TAKARA, Dalian, China) and an Stratagene Mx3000P Real time PCR system (Agilent, CA, USA). The qRT-PCR results were analyzed with the 2−ΔΔCt method. GAPDH was used for normalization the expression levels of genes. The primers used are listed as follows: GAPDH, 5’-TGTTCGTCATGGGTGTGAAC-3’and 5’-ATGGCATGGACTGTGGTCAT-3’, ITGAE, 5’-TGACGAGGACTCAGTTACTAAAAG-3’and 5’-GATGCTGGGTCTCCAAGTCC-3’,IKZF3, 5’-CCATACTGGTGAACGCCCAT-3’ and 5’-CAACCAGTACCAGTGTCCCC-3’, LSP1, 5’-ATGGCGGAGGCTTCGAGTG-3’ and LSP1, 5’-ACTCGGTCCTGTCGATGAGT-3’, NEDD9, 5’-CAGCAGGACTAGCTGTCACC-3’ and 5’-TGATGAGGGAGGGATGTCGT-3’, CLEC2D, 5’-CCTGGAGGTGGGCAGAAAAA-3’ and 5’-ACTTCCTCCACTGCCAGAGA-3’, RBPJ, 5’-CGCCTGTTGTGACAGGGAAA-3’ and 5’-TCCAGGAAGCGCCATCATTT-3’, TRBC2, 5’-CTTCCGCTGTCAAGTCCAGT-3’ and 5’-ATGGGATGCACACCACTCAG-3’, OXNAD1, 5’-ACCACCTGAAGGGGTGTCAA-3’ and 5’-CTCAGGCTGCTAAAACGGGA-3’.
Statistical Analysis
Statistical analyses were performed for all experiments with the GraphPad Prism Software (Version 8.0, San Diego, CA). Experimental results are presented as the means ± standard deviation (S.D.) from at least 3 independent experiments. The statistical differences were calculated by the Student’s t-test. P < 0.05 was considered as being statistically significant.
Results
Identification of the Cell Populations of the HPV+/HPV- Cervical Cancer
The workflow of this study is shown in Figure 1. In the present study, scRNA-seq data from two HPV+ and two HPV- cervical cancer samples in the GSE171894 dataset were used for subsequent analysis. “Seurat” package was used for quality control and downstream analysis. After the quality control, a total of 14,213 cells were used for analysis (Figure 2(A)). As shown in Figure 2(D), all cells were clustered into 20 clusters, and cells were manually annotated with the known marker genes of tumor tissues from the published literature sources.25-28 Such as, Epithelial cells (EPCAM, KRT7, KRT8, KRT17, SPRR3), T cells (CD3E, CD3D, TRBC1, TRBC2, TRAC), myeloid cells (LYZ, CD86, CD68, FCGR3A), B cells (CD79 A, CD79 B, JCHAIN, IGKC, IGHG3), and fibroblasts (DCN, C1R, COL1A1, FGF7) (Figure 2(B)–(D)). Moreover, we also clustered the subpopulations of the T cells, annotating eight subtypes of T cells (Figure 2(D)), including Naïve T cells (IL7R, CCR7, TCF7), cytotoxic T cells (cyto1_T and cyto2_T, GZMK and CXCL13), regulatory T cells (Treg, FOXP3), γδT cells (TRDC and XCL1-2), memory T cell (Tem, IL7R, and TNF), Th17 cells (IL17 A and KLRB1), natural killer T cells (NKT, NKG7 and GNLY), and ISG1_T cells (IFIT1-3 and ISG15). We speculated that these cells might associate with the heterogeneity of patients with or without HPV-infection in cervical cancer. Workflow of this study. Identification of the cell populations of the HPV+/HPV- cervical cancer (A). Violin plots showing the counts levels (nCount_RNA), genes levels (nFeature_RNA), the percentages of mitochondria genes (percent.mt), and the percentages of hemoglobin genes (percent. rb) for each sample. (B) UMAP plots showing the distribution of different cell types (fibroblasts, T cells, epithelial cells, myeloid cells, and B cells) of cervical cancer. (C) Bubble plots showing the cell type marker genes. (D) Heatmap expression levels of specific markers in each cell cluster, and UMAP cluster showing the specific marker genes across the cell clusters in cervical cancer.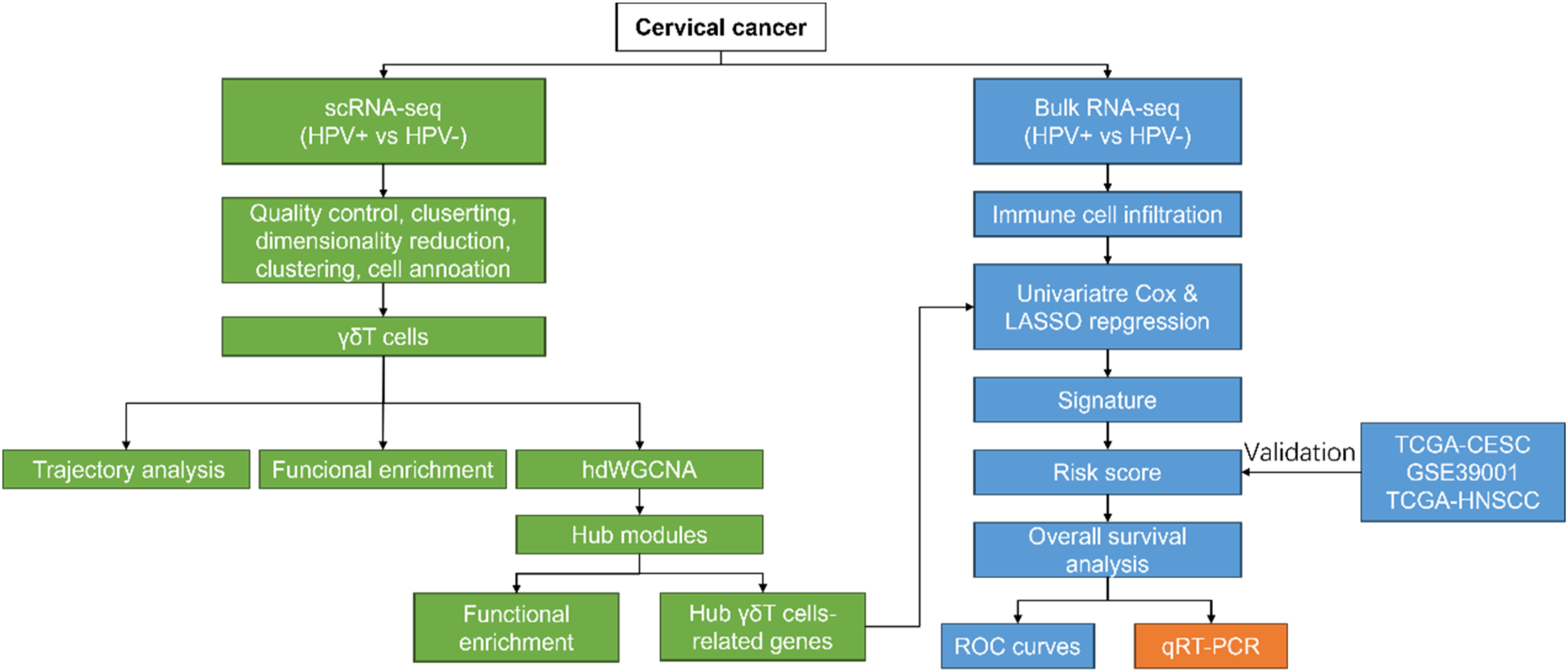

Heterogeneity of Cells in Cervical Cancer
Cell proportions between HPV+ and HPV- cervical cancer samples were explored, showing that abundant subgroups of epithelial cells (Epi_1, Epi_4, Epi_5) were significantly enriched in the HPV + group compared with the HPV- group (Figure 3(A) and (B)). Besides, we also found decreased immune cell infiltration in the HPV + group compared with the HPV- group, such as subgroups of T cells, including cytotoxic_1 (cyto_1 T), cyto_2 T, γδT cells (gammaT), Th17, NKT. Subgroups of myeloid cells, including plasmacytoid dendritic cells (pDC), tumor associated macrophage (TAM), and monocyte. Subgroups of B cells (B_plasma cells) (Figure 3(B)). These results indicated that the presence of heterogeneous cells between HPV+ and HPV- cervical cancer samples, HPV-positive status might be associated with immune suppression. Heterogeneity of cells in cervical cancer A. uniform manifold approximation and projection plots showing the distribution of different cell types of HPV+/− cervical cancer. (B) Histogram showing the proportion of different cell types of HPV+/− cervical cancer.
Heterogeneity of γδT Cells and Its Developmental Trajectory
Furthermore, we focus on the populations of γδT cells between HPV+ and HPV- cervical cancer samples. As shown in Figure 4(A), we found a significantly decreased proportion of γδT cells in the HPV + group compared with the HPV- group. Besides, the pseudotime analysis of γδT cells was performed using “monocle2”. Of note, γδT cells were ordered into five branches/states (Figure 4(B)). Moreover, five branches/states also were clustered into different clusters (Figure 4(C)). We identified the main expression genes with the pseudotime, two unique expression patterns were produced, and two marker genes of γδT cells (XCL1 and XCL2) were significantly expressed following the pseudotime (Figure 4(D) and (E)). To understand the heterogeneity of γδT cells in cervical cancer, GO analysis was performed using the clusterProlfiler package to illustrate the diversity. As shown in (Figure 4(F)), we found γδT cells associated with T cell activation, migration, adaptive immune response, etc. These results indicated that the heterogeneity of γδT cells, and the γδT cells acted as an important role in regulating T cell activities in the tumor microenvironment. Heterogeneity of γδT cells and its developmental trajectory A. Histogram showing the proportion of γδT cells of HPV+/− cervical cancer. (B) Monocle prediction of γδT cell developmental trajectory with pseudotime, and UMAP plots showing the distribution of γδT cell-subclusters with pseudotime. (C) UMAP plots showing the specific marker genes (GZMA, CCL5, and CSF1) across the γδT cell subclusters with pseudotime. (D) Pseudotime heatmap plots the changes of nine genes (CTLA4, CD3D, CD8A, HBB, XCL1, CRTAM, EGR1, AREG, and XCL2) following the pseudotime changes. (E) The pseudo-time trajectory of each representative functional gene in each γδT cell-subcluster. (F) Histogram showing the GO analysis of γδT cell-subclusters with pseudotime.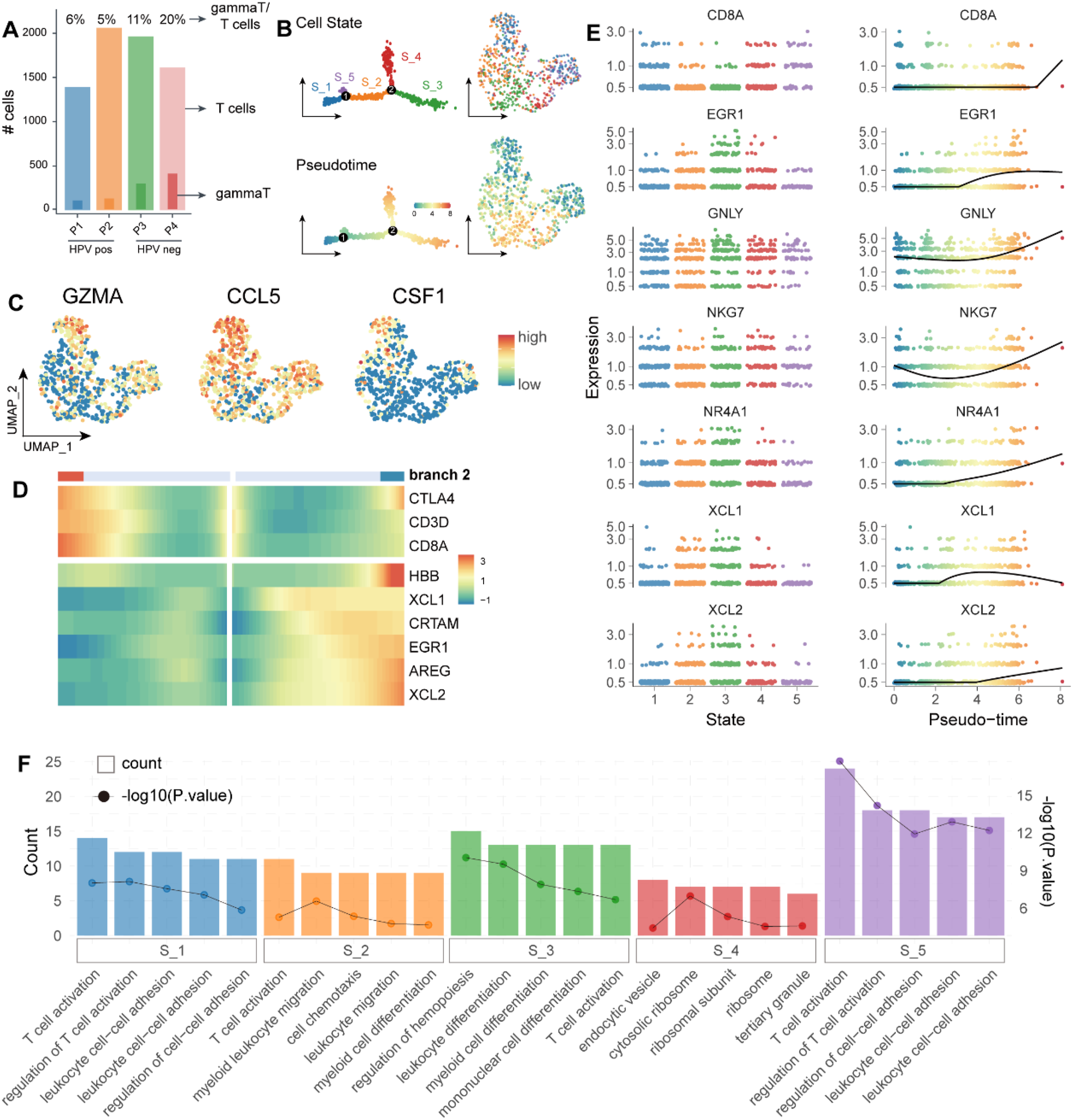
Characteristics of the Infiltrating Immune Cells in HPV+/HPV- Cervical Cancer
Meanwhile, we also investigated the immune characteristics in TME based on the bulk RNA-seq data. The results revealed the differences in infiltrating immune cells between HPV+ and HPV- cervical cancer samples. The results revealed that proportions of the CD8 T cells, CD4 T cells, resting memory CD4 T cells, resting NK cells, and macrophage M2 were decreased, but activated memory CD4 T cells, helper follicular T cells, activated NK cells, monocytes, macrophage M0, activated DCs, resting mast cells, eosinophils, and neutrophils were increased in HPV + group compared with HPV- group (Figure 5(A)). Furthermore, our GSEA results showed that the HPV + group was mainly associated with metabolic pathways (oxidative phosphorylation, cholesterol homeostasis, and glycolysis), cell proliferation-related pathways (G2M Checkpoint, E2F Targets, MYC Targets V1 and V2, and Mitotic Spindle), PI3K/AKT/MTOR signaling, TGF-β signaling, and MTORc1 signaling (Figure 5(B)). We also found HPV- group was significantly associated with several immune-related pathways, such as IL-6/JAK/STAT3 signaling, TNF-а signaling via NF-кB, and Wnt-β catenin signaling (Figure 5(B)). The above results indicated that HPV infection was associated with distinct immune cell infiltrating and cell proliferation. We further detected the relationship between γδT score and inflammatory response, as well as between γδT score and MYC stemness, resulting that γδT score was positively associated with inflammatory response, but γδT score was negatively associated with MYC stemness (Figure 5(C)). Our finding suggested that γδT cells might be involved in the inflammation and cell proliferation of HPV + cervical cancer. Characteristics of the infiltrating immune cells in HPV+/HPV- cervical cancer A. Boxplots showed the differences in infiltrated immune cells between HPV+ and HPV- cervical cancer. (B) GSEA results showed the biological function in HPV+ and HPV- cervical cancer in the TCGA training cohort. (C) Scatter plots showing the correlation between γδT-score and inflammatory response as well as MYC stemness.
Identification of Four Hub Modules Relevant to γδT Cells by hdWGCNA
HdWGCNA was used to explore the potential function of γδT cells in cervical cancer. “hdWGCNA” R package was performed to construct a hdWGCNA (Figure 6(A)). In detail, the soft threshold was set as 4 with the scale-free topology mode fit more than 0.8 (Figure 6(B)). Correlation analysis between all identified models by hdWGCNA revealed that strong correlation between M1 and M4, M2 and M1, as well as M2 and M6 (Figure 6(C)). Therefore, four gene modules (M1, M2, M4, and M6) were obtained and the modules that were associated with γδT cells and cytotoxic T cells were identified (Figure 6(D)). GO analysis of each module showed that M1 and M4 modules were mostly associated with T cell activation and lymphocyte differentiation (Figure 6(E) and (G)). M2 module was enriched in cell-substrate adhesion and junction, focal adhesion, cell-matrix adhesion, and cell leading edge (Figure 6(F)). And M6 module was connected to nuclear speck and cAMP-dependent protein kinase (Figure 6(H)). These findings suggested that γδT cells contribute to immune cell infiltrating in cervical cancer. Identification of four hub modules relevant to γδT cells by hdWGCNA. (A) The hdWGCNA dendrogram of γδT cells. (B) Selection of soft power for running hdWGCNA. Max, median, and mean connectivity were shown, respectively. (C) Correlation analysis between all identified modules by hdWGCNA. (D) Distribution of module scores in γδT cells. There are a total of six gene modules for γδT cells. (E)–(H). Protein-protein interaction network of module genes and functional analyses for four γδT cell-related modules, respectively.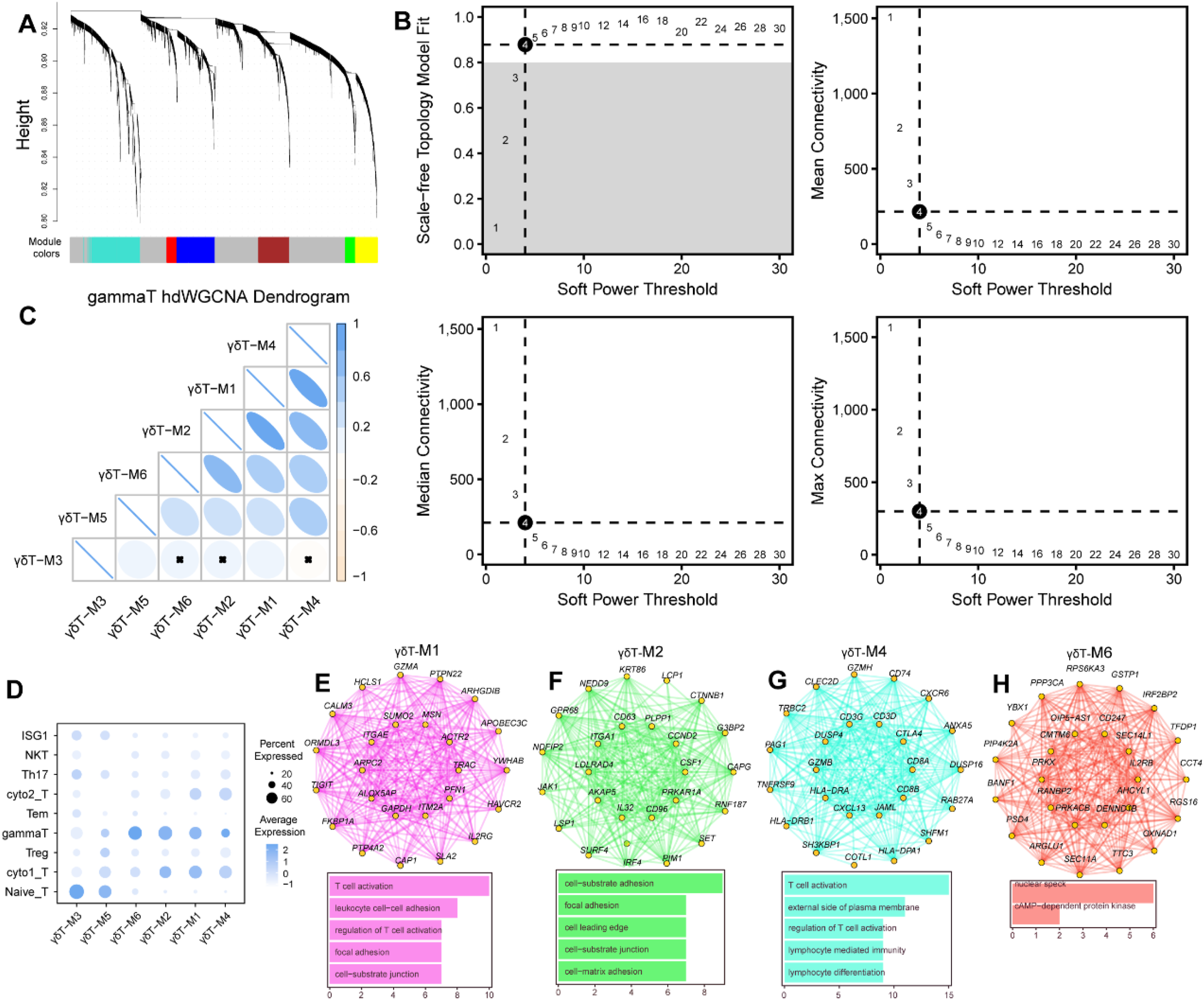
Development and Validation of the γδT Cell-Related Risk Score in Cervical Cancer
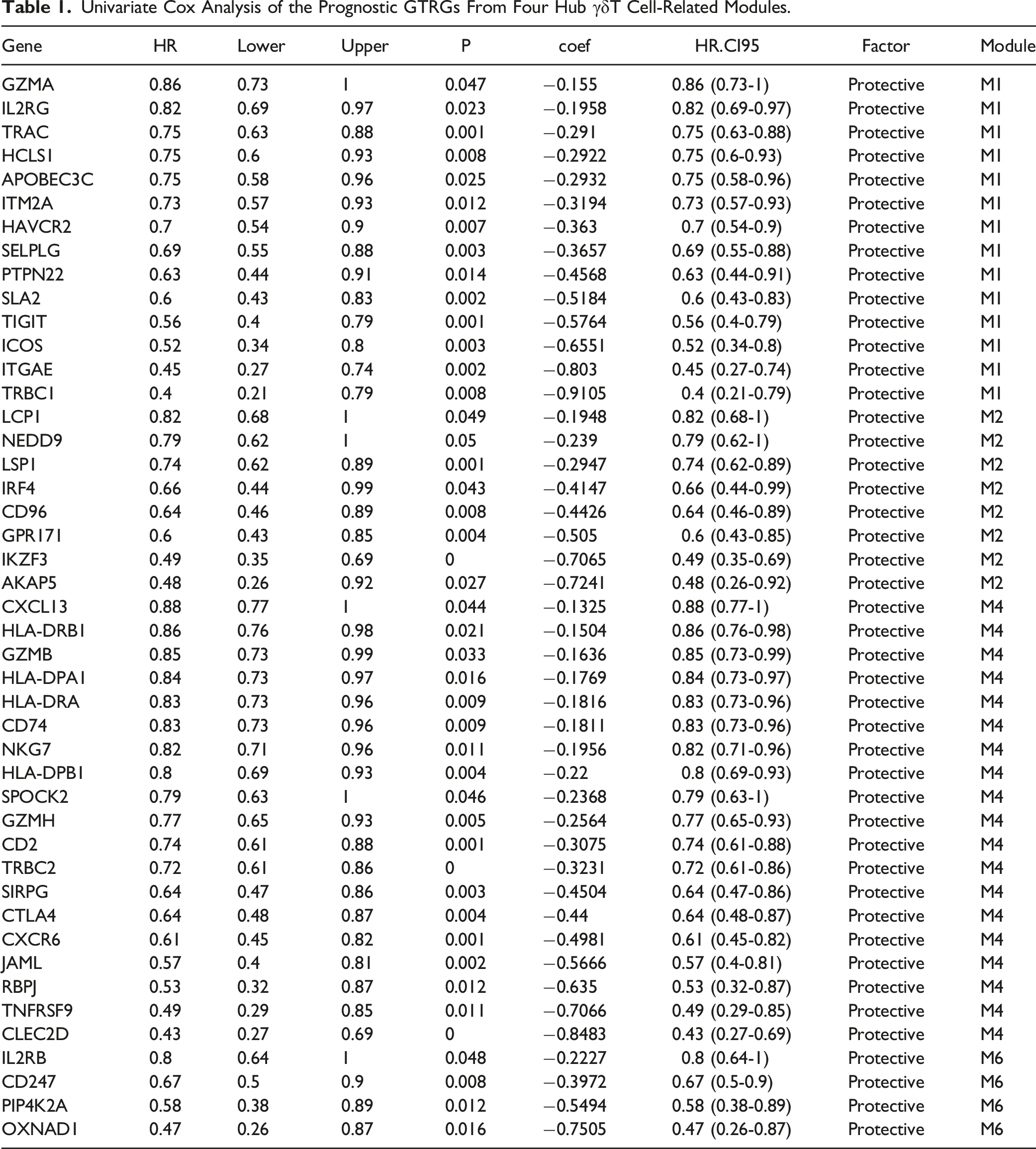
Based on the hdWGCNA results, a total of 45 prognostic GTRGs were obtained from the four γδT cell-related modules (M1, M2, M4, and M6) using univariate Cox analysis (Figure 7(A) and Table 1). 45 GTRGs then were incorporated into the LASSO regression model, resulting in eight GTRGs, ITGAE, IKZF3, LSP1, NEDD9, CLEC2D, RBPJ, TRBC2, and OXNAD1, were selected based on the coefficient of each gene (Figure 7(B)). Then, the risk score of each sample in the training set (TCGA-CESC cohort) was calculated based on the expression and coefficient of each gene, patients then were divided into high- and low-risk groups according to the median risk score (Table S1). Moreover, we observed the correlation between the expression patterns of GTRGs, ITGAE, IKZF3, LSP1, NEDD9, CLEC2D, RBPJ, TRBC2, and OXNAD1 and risk score. As a result, the expression levels of GTRGs, ITGAE, IKZF3, LSP1, NEDD9, CLEC2D, RBPJ, TRBC2, and OXNAD1 were upregulated in the high-risk group compared with the low-risk group (Figure 7(C)). Kaplan-Meier curve for OS indicated that the high-risk group showed a worse survival rate than the low-risk group (Figure 7(D)). Consistently, the AUC values of time-dependent ROC curves for 3- and 5-years were 0.694 and 0.701, respectively in the training set (Figure 7(E)), indicating the robustness of the prognostic model. To detect the predictive ability of the prognostic signature, the GSE39001 cohort of cervical cancer patients was selected as an external validation set to verify the predictive ability of the prognostic model, the AUC values of time-dependent ROC curves for 3- and 5-years were 0.746 and 0.801, respectively (Figure 7(F)), which also verified the predictive ability of the prognostic model. Moreover, we also investigated whether the prognostic model applied to other HPV-infected tumors. The AUC values of time-dependent ROC curves for 3 years were 0.786 (Figure 7(G)), indicating that the prognostic model able to predict HPV infection in HNSCC. These finding suggested that the potential valuable of the γδT cell-related risk model for predicting the prognosis of HPV infected cancers. Development and validation of the γδT cell-related risk score in cervical cancer A. Univariate Cox analysis of the prognostic genes for cervical cancer based on TCGA-CESC cohort. (B) Left: Ten-fold cross-validation of parameter selection in the LASSO model. Right: LASSO regression coefficient of the five genes. (C) Heatmap showed the expression of prognostic genes with a risk score. (D) Kaplan-Meier OS curves for OS between high- and low-risk groups in cervical cancer patients in TCGA-CESC cohort. (E)–(G) Time-dependent ROC curves for OS of cervical cancer patients in TCGA-CESC cohort, GSE39001, and TCGA-HNSCC cohorts. Univariate Cox Analysis of the Prognostic GTRGs From Four Hub γδT Cell-Related Modules.
Validation of the Expression of γδT Cell-Related Prognostic Signature Genes
In the present study, qRT-PCR was used to detect the expressions of γδT cell-related prognostic signature genes in cervical cancer. As shown in (Figure 8(A)–(H)), we found that the increased ITGAE, IKZF3, TRBC2, and OXNAD1, but downregulated expression of LSP1, NEDD9, CLEC2D, and RBPJ in cervical tumor tissues compared with adjacent normal tissues. These finding suggested that the significantly differential expressed γδT cell-related signature genes in cervical cancer. Validation of the expression of γδT cell-related prognostic signature genes (A)–(H). The histograms of the expression of ITGAE, IKZEF3, LSP1, NEDD9, CLEC2D, RBPJ, TRBC2, and OXNAD1 between cervical tumor tissues and adjacent normal tissues. *P < 0.05, **P < 0.01, ***P < 0.001.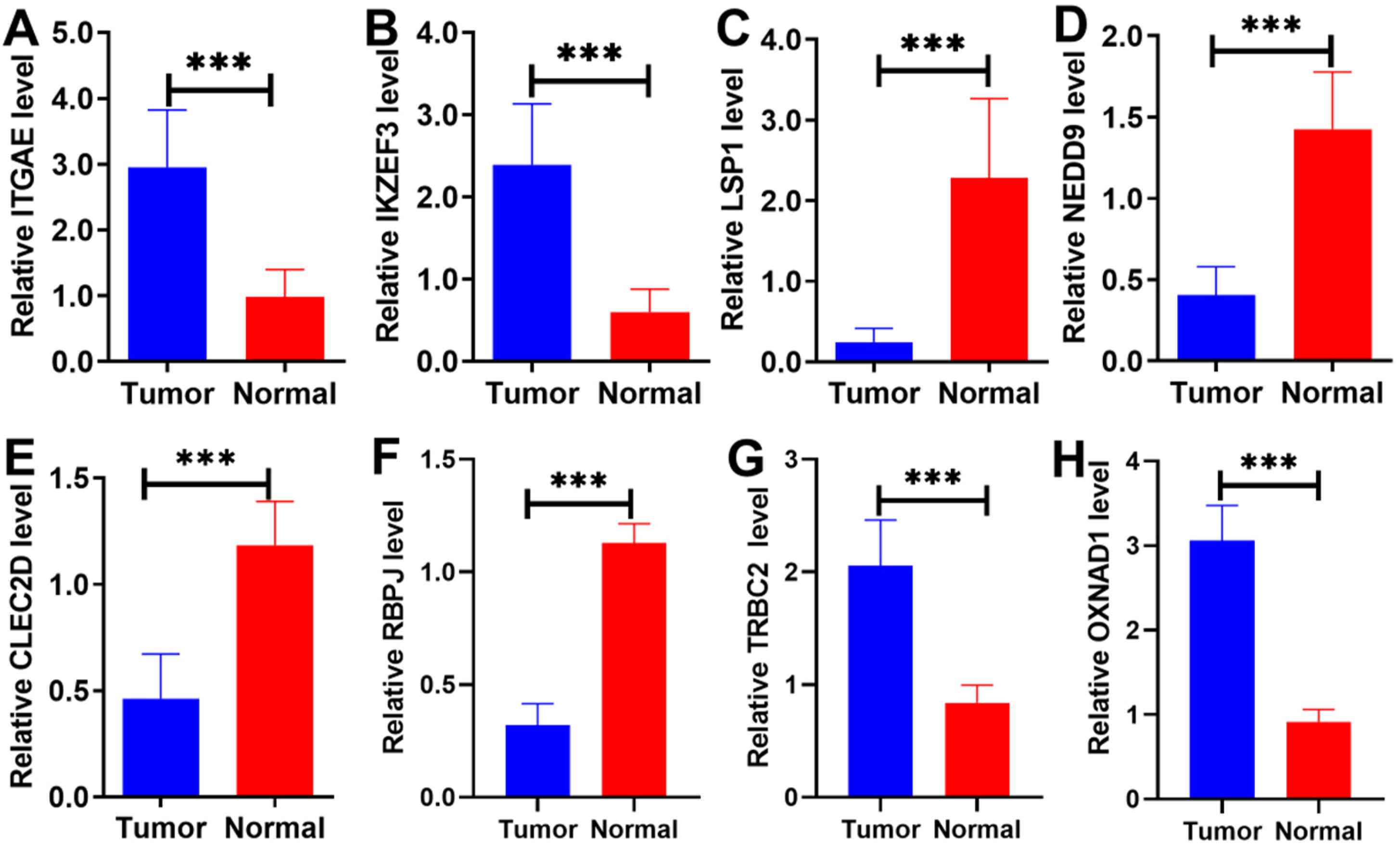
Discussion
Increasing evidence reveals that γδT cells act as a major component of tumor-infiltrating lymphocytes (TILs), playing a crucial role in tumor immunity. However, they often exert immunosuppressive functions in multiple solid tumors.29,30 Edelblum KL, et al have found that γδT cells exert both pro- and anti-tumor effects in tumors by regulating a wide rang of functional plasticity. 31 γδT cells have been reported to contribute to human colorectal cancer progression by polymorphonuclear myeloid-derived suppressor cells (PMN-MDSCs)-mediated immunosuppression. 32 In pancreatic ductal adenocarcinoma (PDA), we found that γδT cells promote tumorigenesis by inhibiting αβT cells. 33 Besides, hypoxia accelerates γδT cells differentiated to IL-17 producing-γδT cells in oral cancer. 34 Dogan S, et al have revealed that the alterations in population and function of γδ T cells contribute to persistent HPV infection and carcinogenesis in the early stage of cervical cancer. 19 Although it has been reported that γδ T cells are components of the mucosal immune defense of the female genital tract against HPV, the role of γδ T cells in HPV-infected cervical cancer remains unclear.
An increasing number of studies have discovered that γδT cells play a dual role in tumors, acting both pro- and anti-tumors, and influencing the outcome of the immune response through exerting cytotoxic, cytolytic, and immune-regulatory functions.13,35 Here, single-cell analysis, hdWGCNA, immune cell infiltration, biological functional analysis, and construction of prognostic signature were performed to explore the role of γδT cells in HPV-infected cervical cancer and to identify the γδT cells-related prognostic signature. Results indicated that the increased epithelial cells (Epi_1, Epi_4, Epi_5) and decreased immune cells (cyto_1 T, cyto_2 T, γδT, Th17, NKT, TAM, monocyte, B_plasma cells) in HPV-positive infection cervical cancer. Notably, we found a decrease in the population of γδ T cells in HPV-positive infection cervical cancer. Biological functional analysis indicated that γδT cells were involved in the T cell activation in TME. The above results also demonstrated that γδT cells contribute to the connection between innate and adaptive anti-tumor immune response. 35 The pseudotime analysis of γδT cells indicated that γδT cells were ordered into five branches/states, and two marker genes of γδT cells (XCL1 and XCL2) were significantly expressed with the pseudotime. The NK/NKT cell marker genes (NKG7 and GNLY) were found by dynamic changes in gene expression levels with the pseudotime. This finding supports that γδT cells could differentiate into effectors under the viral infection state. 36
We also found the distinction of infiltrating immune cells between HPV+ and HPV- cervical cancer samples. Functional analysis revealed that HPV-infected cervical cancer is involved in metabolic pathways, cell proliferation-related pathways, PI3K/AKT/MTOR signaling, TGF-β signaling, and MTORc1 signaling. However, the non-HPV-infected cervical cancer is associated with several immune-related pathways. Although there were no significant changes in a fraction of γδT cells between HPV+ and HPV- cervical cancer patients based on bulk RNA-seq data may be due to patient heterogeneity, significant differences were found in the scRNA-seq data. We found that γδT score was positively associated with inflammatory response, but γδT score was negatively associated with MYC stemness. Previous studies have demonstrated that γδT cells can be used in potential therapeutics against HPV in infected patients, 19 and increasing the cytotoxic ability of γδT cells may improve the immunotherapeutic results.17,37
Therefore, we identified four γδT cell-related gene modules based on hdWGCNA and then selected eight γδT-related gene signatures (ITGAE, IKZF3, LSP1, NEDD9, CLEC2D, RBPJ, TRBC2, and OXNAD1) for HPV-infected cancer. Integrin Alpha E, also named itgae, αE, and CD103), which widely expressed in migrating and resident T cells and DCs in many organs and tumors.38,39 CD103 interacts with E-cadherin to promote anti-tumor cytotoxic T lymphocyte (CTL) activity.40-42 Studies have indicated that CD103 participates in the regulation of immune activation after viral and bacterial infections.43,44 Fleming C, et al have found that microbiota-activated CD103+ DCs drive γδT cell proliferation and activation. 45 Moreover, several studies reported that a high intratumor abundance of CD103+ immune cells of HPV-infected oropharyngeal cancers (HPV + OPSCC) is associated with an excellent prognosis for survival.46-48 Recent studies also reveal that CD103 expression is associated with prognostic benefit and therapy response in cervical cancer.49,50 Here, we also demonstrated that CD103 acts as a protector in cervical cancer that involves regulating γδT cells.
IKZF3 (encoding for the protein Aiolos) is a member of the Ikaros Zinc finger family of transcription factors and expressed in various immune cell types.51-55 IKZF3 has been identified as a prognostic biomarker in HPV-infected cervical cancer and HNSC.56,57 Our results also supported previous findings. Leukocyte-specific protein 1 (LSP1) polymorphisms are widely reported in breast cancer.58-61 It is first reported in cervical cancer. NEDD9 is a focal adhesion scaffolding protein and is associated with tumor invasion and metastasis ,62-65 while it is rarely reported in cervical cancer. 66 In the present study, NEDD9 has been identified as a potentially prognostic gene for cervical cancer.
C-type lectin-like domain family 2 (CLEC2D, LLT1) is a ligand for NK cell inhibitory receptor NKRP1A (CD161) and is significantly expressed in the tumor cells, as well as inhibits NK cytolytic function.67,68 Sanchez-Canteli M, et al, have discovered that CLEC2D acts as an independent prognostic gene in HPV-negative oropharyngeal squamous cell carcinoma. 69 In this study, we also found that CLEC2D acts as an important prognostic biomarker in HPV-negative cervical cancer. Pan B, et al have found that tecombination signal binding protein for immunoglobulin kappa J region (RBPJ) inhibits the CD8+ T cell-mediated killing function and acts as a novel immunotherapeutic target in hepatocellular carcinoma. 70 Here, RBPJ has been selected as a prognostic biomarker for cervical cancer. T cell receptor β-chain constant (TRBC) 2 is a potentially therapeutic target for cancer by discriminating malignant from healthy (normal) T cells.71-73 The expression of OXNAD1 is downregulated in the peripheral blood mononuclear cell (PBMC) aging and it has been identified as a specific biomarker of immune system aging. 74 In the present study, we first selected OXNAD1 as a promising biomarker for the prognosis of cervical cancer.
Based on γδT cells-related gene signature, we calculated the risk score for each patients in cervical cancer and HNSCC and found that the poor survival status in patients with high-risk compared with low-risk scores. We also collected the clinical specimens to demonstrate the differential expression of those genes in cervical cancer. γδT cells are widely used in the filed of immunotherapy in recent years with their extraordinary antitumor potential and prospects for clinical application. 75 Our finding described the landscape of the γδT cells in HPV+ and HPV- cervical, and provided potential γδT cells-related biomarkers for diagnosis and treatment of cervical cancer, even for other HPV-infected cancers. However, there are some limitations exist in the present study. The main conclusion of this study, that T cells exhibit heterogeneity in HPV-infected tumors and are associated with prognostic markers, has not been thoroughly validated through cell and animal experiments. Therefore, in future studies, we will further apply for animal ethics approval and conduct in vivo and in vitro experiments to comprehensively investigate and validate the above conclusions.
Conclusion
In summary, we elucidated the heterogeneity of γδT cells between HPV+ and HPV- infection cervical cancer. Moreover, eight γδT-related gene signatures (ITGAE, IKZF3, LSP1, NEDD9, CLEC2D, RBPJ, TRBC2, and OXNAD1) were identified for cervical cancer prediction. Our finding provided the reliable evidence for targeting γδT cells and their related signatures might a viable therapeutic strategy for treating patients with cervical cancer.
Supplemental Material
Supplemental Material - Integrating Single-Cell RNA-Seq and Bulk RNA-Seq to Construct a Novel γδT Cell-Related Prognostic Signature for Human Papillomavirus-Infected Cervical Cancer
Supplemental Material for Integrating Single-Cell RNA-Seq and Bulk RNA-Seq to Construct a Novel γδT Cell-Related Prognostic Signature for Human Papillomavirus-Infected Cervical Cancer by Xiaochuan Wang, Yichao Jin, Liangheng Xu, Sizhen Tao, Yifei Wu, and Chunping Ao in Cancer Control.
Footnotes
Author Contributions
Chunping Ao, Yichao Jin, and Yifei Wu conceived and designed the experiments. Chunping Ao, Liangheng Xu, and Sizhen Tao performed the program. Chunping Ao and Xiaochuan Wang analyzed the data. Yifei Wu provided the analysis tools. Chunping Ao wrote the original draft. Chunping Ao and Xiaochuan Wang wrote, reviewed, and edited the article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the Yunnan Provincial Key Laboratory of Clinical Virology Open Project (202205AG070053), Yunnan Provincial Department of Science and Technology-Kunming Medical University Joint Special Project on Applied Basic Research (202201AY070001-251), and Yunnan Provincial Basic Research Program (202401AT070051).
Ethical Statement
Data Availability Statement
The datasets presented in this study can be found in TCGA (https://portal.gdc.cancer.gov/), and GEO (![]() ). The names of the accession numbers are shown in the methods in the article.
). The names of the accession numbers are shown in the methods in the article.
Supplemental Material
Supplemental material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
