Abstract
BACKGROUND:
Epithelial ovarian cancer (EOC) is the leading cause of death from gynecologic malignancies. The poor prognosis of EOC is mainly due to its asymptomatic early stage, lack of effective screening methods, and a late diagnosis in the advanced stages of the disease.
OBJECTIVE:
This study investigated metabolomic abnormalities in epithelial ovarian cancers.
METHODS:
Our study developed a novel strategy to rapidly identify the metabolic biomarkers in the plasma of the EOC patients using Internal Extraction Electrospray Ionization Mass Spectrometry (IEESI-MS) and Liquid Chromatography-mass Spectrometry (HPLC-MS), which could distinguish the differential metabolites in between plasma samples collected from 98 patients with epithelial ovarian cancer, including 78 cases with original (P), and 20 cases with self-configuration (ZP), as well as 60 healthy subjects, including 30 cases in the original sample (H), 30 cases in self-configuration (ZH), and 6 cases in a blind sample (B).
RESULTS:
Our study detected 880 metabolites based on criteria variable importance in projection (VIP)
CONCLUSIONS:
Identifying the abnormal metabolites of EOC patients through metabolomics analyses could provide a new strategy to identify valuable potential biomarkers for the screening and early diagnosis of EOC.
Introduction
Ovarian cancer remains the most common cause of death among women with gynecological malignancies worldwide [1]. According to the American Cancer Society assessment, there were about 21,000 new cases and 14,000 deaths of ovarian cancer in 2021 [2]. Epithelial ovarian cancer (EOC) accounts for more than 95% of ovarian malignancies. Histologically, EOCs are divided into five subtypes: high-grade serous, low-grade serous, clear cell, endometrioid, and mucinous. High-grade serous ovarian cancer is the most common histologic subtype, accounting for more than 70% of EOC [3, 4]. EOC is characterized by the following aspects. Approximately 75% of women with EOC are diagnosed in the advanced stages (IIIc and IV). The five-year survival rate of EOC patients with stage IIIC-IV is less than 30%, with death occurring within the first two years of diagnosis [5]. The standard treatment for EOC includes tumor reduction surgery and platinum-based chemotherapy [6]. Despite a successful initial response to initial treatment, approximately 80% of patients experienced disease relapse, with many receiving multiline therapy after relapse. Each subsequent first-line treatment is characterized by short disease-free intervals, which eventually leads to resistance and death [7]. The low survival rate of ovarian cancer patients is mainly due to the high percentage of late diagnoses in the advanced stages of the disease. Therefore, early diagnosis could be the key to improving the prognosis of ovarian cancer patients because they will gain benefits from screening to have a better clinical outcome. However, currently, there are no effective early screening methods. The most commonly used screening methods for EOC in the clinic are CA125 (cancer antigen 125), HE4 (human epididymis protein 4), and the TVS (transvaginal ultrasonography single or combined inspection). Although positive results could be found using those screening methods in some small studies [8], however, the UK Collaborative Trial of Ovarian Cancer Screening (UKCTOCS) still considered that using those biomarkers for screening EOC patients has not significantly reduced the number of deaths from ovarian and fallopian tube cancers, it is not recommended for the general population [9]. Thus, it is necessary to develop new screening methods for EOC, which could accurately determine the occurrence and progression of the disease and predict the prognosis of patients.
On the other hand, due to the maintenance of the survival of cancer cells, some metabolic changes in cancer patients could be detected prior to the occurrence of symptoms and signs, such as abnormalities in biological fluids, including blood, plasma, urine, and ascites [10]. Metabolomics is an emerging discipline that has been gradually developed and applied in recent years. By analyzing metabolites in patients’ blood, urine, and other body fluids, valuable biomarkers used for clinical diagnosis, prognosis, and monitoring can be identified, which are also applied for screening due to their ability to detect the new molecular therapeutic targets [11, 12]. Since they are products and substrates of metabolism, metabolites also reflect changes in cell functions related to tumorigenesis and cancer development, such as cell proliferation and apoptosis [13]. The metabolomics combined with the multivariate analysis has been applied to diagnose different types of cancer [14, 15, 16]. For example, a simple solid-phase extraction method based on octadecylsilyl (ODS) coupled with internal extractive electrospray ionization mass spectrometry (ODS-SPE-iEESI-MS) was developed for the analysis of blood plasma samples, with features of high sensitivity, high specificity, low sample consumption, and high throughput [17]. Moreover, the iEESI-MS and HPLC-MS techniques can identify the plasma metabolites of the EOC patients, which can directly or indirectly regulate the metabolic function of EOC. This study evaluated the value of the novel biomarkers identified from the plasma metabolites of the EOC patients by iEESI-MS and HPLC-MS in the application of screening, early diagnosis, and prognostic prediction of EOC patients.
Materials and methods
Sample information
The enrolled EOC patients
The participants in this study were divided into two groups: the EOC patients and healthy subjects. All EOC patients were newly diagnosed with EOC, confirmed by surgery and pathology in our hospital’s oncology department. They were treated at the First Hospital of Jilin University during the period from July 2014 to July 2017. A total of 78 patients with EOC were enrolled in this study. Patients enrolled in the study were not allowed to receive chemotherapy or radiation before taking blood samples. Prior to enrolling any patients, all the participating institutions must obtain the approval of the local institutional review board. All the patients had provided written informed consent before collecting their blood samples. Patients with a history of malignancy other than skin basal cell carcinoma were excluded from this study. Patients underwent baseline assessment before entering the study, including physical examination, blood count, CA-125 levels, and preoperative computed tomography. The clinical stages of those patients were determined according to the International Society of Obstetricians and Gynecologists (FIGO) staging standard in 2005. Stage I tumors are confined to the ovaries or fallopian tubes. Stage II tumors involve one or both ovaries, fallopian tubes with pelvic extension (below the pelvic margin), or primary peritoneal carcinoma. Stage III tumors involve one or both ovaries or fallopian tubes or primary peritoneal carcinoma with cytological or histological evidence of spread to the peritoneum outside the pelvis and/or metastasis to retroperitoneal lymph nodes. Stage IV tumors have distant metastases, excluding peritoneal metastasis [18]. The ovarian cancer patients who met the enrollment criteria and agreed to participate in stage I–IV EOC could be enrolled in this study. All of them were admitted to our hospital between July 2014 and July 2017, among which there were 22 cases of stage I, 7 cases of stage II, 46 cases of stage III, and 3 cases of stage IV. Their cell classification included high-grade 39 cases and low-grade 10 cases. Their cell types were listed as serous adenocarcinoma (55 cases), clear cell carcinoma (5 cases), mucinous cystadenocarcinoma (6 cases), endometrial adenocarcinoma (4 cases), and other epithelial carcinomas (8 cases) (Table 1). In addition, 30 healthy women underwent physical examination in our hospital during the same period as controls in our study. All the healthy control group participants provided written informed consent before collecting their blood samples. The investigation of the plasma metabolites of the EOC patients was approved by the Medical Committee in the Hospital Institutional Review Board of the First Hospital of Jilin University, Jilin, P.R. China (2014-337).
Clinical information related to patients’ plasma samples
Clinical information related to patients’ plasma samples
The collected plasma samples were stored in a
Materials and chemicals
The chemicals used in this study were HPLC-grade methanol, acetonitrile (ACN), and formic acid (FA), all purchased from Merck KGaA (Darmstadt, Germany). Other materials included ultrapure water obtained from the Milli-Q purification system (Millipore; Bedford, MA, USA), nylon 66 pin type filter (pore size 0.22
Preparation of octadecylsilyl using the organic imported nylon 66 needle type filter
Octadecylsilyl (ODS), also called C18, is an organic group. ODS silica gel is the most commonly used packing for reversed-phase chromatography. As shown in Fig. 1A, the mixture of octadecylsilyl and methanol was made in a glass beaker (homogenate concentration of 2 mg/mL), followed by transferring the mixture (0.5 ml) into a syringe (Hongda company, Nanchang, China) (Fig. 1B) for capturing the octadecylsilyl (groups) by an organic imported nylon 66 needle type filter, where ODS (groups) were held. However, methanol waste was allowed to pass through and discharge into a glass beaker (Fig. 1C). Then, ODS captured by the filter was washed with 1 mL ACN/H2O (v/v, 95/5) (Fig. 1D (1)) and 1 mL ACN/H2O (v/v, 5/95) (Fig. 1D (2)) in turn.
The schematic diagram of the concept of the iEESI-MS method. The Orbitrap Fusion™Tribrid™mass spectrometer (Thermo Scientific, San Jose, CA, USA) was used for all the experiments coupled with a homemade iEESI source. The MS system was performed using the positive-ion (ESI
SPE based on ODS coupled with iEESI-MS
The solid-phase extraction (SPE) based on the octadecylsilyl coupled with internal extractive electrospray ionization mass spectrometry (ODS-SPE-iEESI-MS) is a novel detection method for the analysis of plasma metabolites (Fig. 1). The Orbitrap Fusion™ Tribrid™mass spectrometer (Thermo Scientific, San Jose, CA, USA) was used for all the experiments coupled with a homemade iEESI source. Briefly, the plasma samples were thawed at room temperature (25
Analysis using HPLC-MS technology
Samples were loaded by auto-sampler via the LC system (Thermo Fisher Scientific, Waltham, MA). In this system, the mobile phase A was H
Data preprocessing
Statistical and machine learning methods are essential for analyzing mass spectrometry (MS)-generated complex metabolic data. Construction of the statistical model with calculation algorithms for multivariate analysis and feature selection of metabolomic data requires the initial “training” step and subsequent verification steps. In the “training” step, the orthogonal partial least squares discriminant analysis (OPLS-DA) was used to improve the PLS-DA algorithm for the multivariate analyses of the metabolomic data. The statistical model based on OPLS-DA can discriminate (divide) two or more groups (classes) using multivariate data. Discriminating variables were selected according to variable VIP
Results and discussion
Evaluation of the capability of the iEESI-MS and the HPLC-MS based on the analysis of metabolomic data using OPLS-DA
The analysis of metabolomic data of all groups was performed using OPLS-DA, including the EOC patients (P group), normal subjects (N group), and blind subjects (B group). The OPLS-DA score plots showed a clear separation trend between the P and N, with satisfactory modeling and predictive abilities, according to the analysis results of the metabolomic data collected from iEESI-MS (R2Ycum
The data analysis using the OPLS-DA method to evaluate the capability of the iEESI-MS and the HPLC-MS. A–B: OPLS-DA score plots (A) and S-plots (B) based on the data collected from iEESI-MS (A and B) for distinguishing P and N groups. The variables with VIP 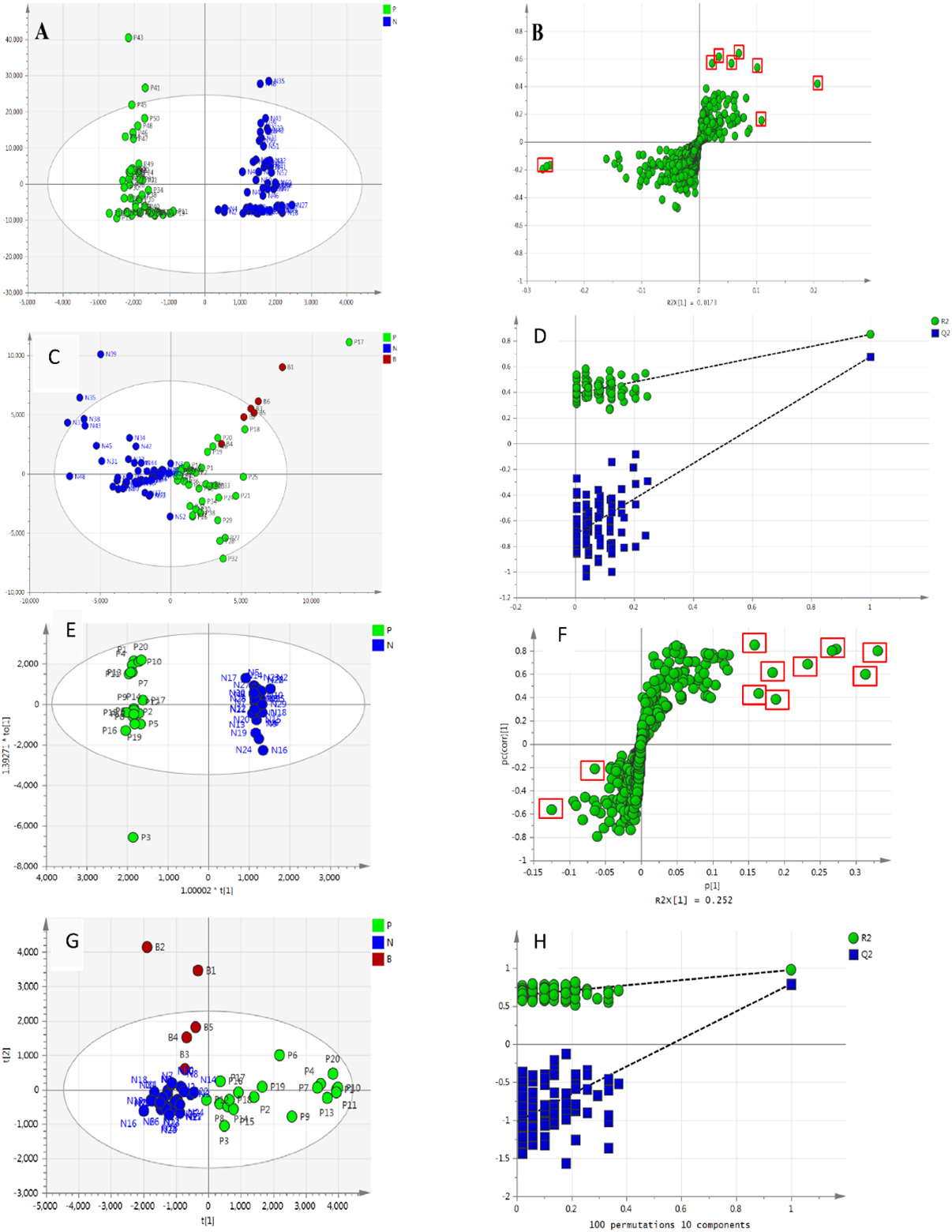
iEESI-MS mass spectrograms of plasma samples collected from the EOC patients (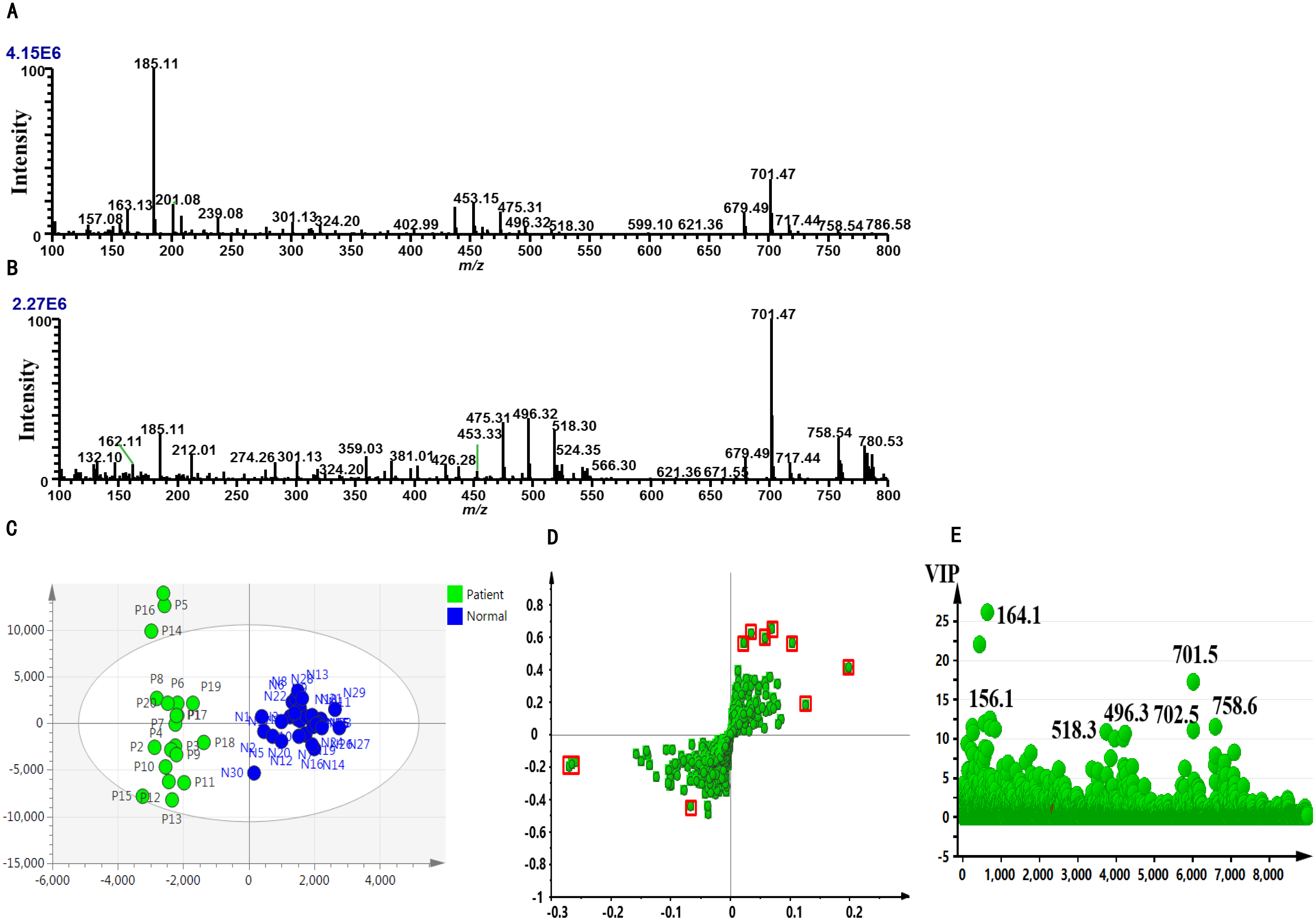
The metabolomic data of plasma samples of the EOC patients (60 cases) and normal subjects (60 cases) were collected by IEESI-MS. The data analysis showed that there were significant differences in the m/Z (100–800) mass spectrograms between the EOC patients and normal subjects (Fig. 3A: EOC patients and Fig. 3B: normal subjects). The analysis results of iEESI-MS mass spectrometry data by OPLS-DA using SIMCA software are presented as the score plots in Fig. 3 CE. As shown in the two-dimensional and S-Plot scatter diagram of OPLS-DA (Fig. 3C and D), respectively, there was a significant difference in the ratio of plasma metabolites between the EOC patients and healthy women. S-plot dispersion plots also showed the differential metabolizable product-charge ratios in plasma between the EOC patients and healthy women, with greater differences at points further dispersed. As shown in Fig. 3E, VIP scatter plots were used to distinguish group P from group N the possible markers (VIP
PLS-DA recognition score map of the IEESI-MS mass spectrometry data (including self-configured plasma samples)
The EOC patients and healthy subjects could be distinguished in the PLS-DA recognition score map of the IEESI-MS mass spectrometry data of their plasm samples. In addition, all of the blind samples were also distinguished from those of the healthy subjects, suggesting that the analysis results were reliable. All of the analysis results of 98 EOC patients [78 in the original sample (P) and 20 in the self-configured sample (ZP)], normal subjects [30 in the original sample (H) and 30 in the self-configured sample (ZH)], and six in the blind sample (B) were presented in Fig. 4A–C, including (A) PLS-DA diagram of the two-dimensional analysis, (B) PLS-DA diagram of the three-dimensional analysis, and (C) S-plot dispersion (points further apart, showing the difference in metabolites between the EOC patients and healthy subjects). As shown in Fig. 4A–C, the analysis of iEESI-MS mass spectrometry data by the PLS-DA method can also effectively and accurately distinguish the EOC patients and healthy subjects, even under the interference of self-configured plasma samples and blind samples. Further analysis was performed by excluding the self-dispensing and blind samples. Only 78 samples of the EOC patients (P) and 30 samples of the normal subjects (N) were used for analysis. The OPLS-DA recognition score map of IEESI-MS mass spectrometry data was presented in Fig. 4D–F, including (D) the PLS-DA diagram of two-dimensional analysis, (E) the PLS-DA diagram of three-dimensional analysis, and (F) S-plot dispersion. Similar results were obtained to prove our conclusion.
Potential biomarkers for distinguishing P and N groups by analysis of data from IEESI-MS
Potential biomarkers for distinguishing P and N groups by analysis of data from IEESI-MS
PLS-DA recognition score map of IEESI-MS mass spectrometry data. A–C are the results of the EOC patients (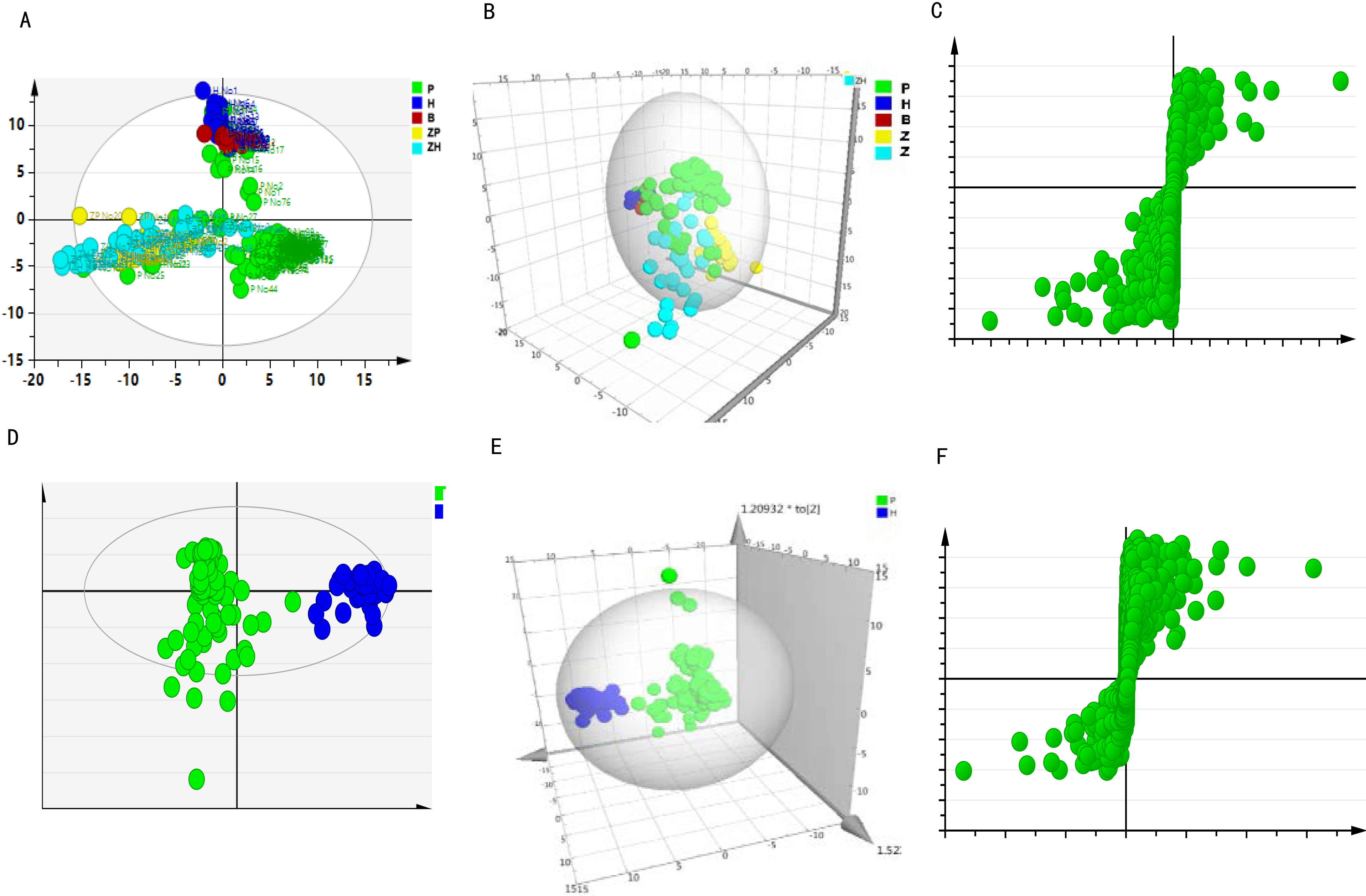
The VIP value reflects the influence of each variable on classification and the proportion of an individual metabolite in all metabolites. Thus, the VIP value of each metabolite is varied. It is generally considered that the substances (VIP
Potential biomarkers for distinguishing P and N groups by HPLC-MS
Potential biomarkers for distinguishing P and N groups by HPLC-MS
There were significant differences in HPLC-MS chromatograms of plasma samples or the plasma mass spectrogram between the EOC patients (
Identification of the potential markers for discriminating the groups of the EOC patients and normal subjects
Our study demonstrated that the plasma metabolomics analysis by iEESI-MS and HPLC-MS is a valuable tool for discriminating the groups of the EOC patients and normal subjects (
Potential metabolic markers and hypothesis of mechanisms. A: identification of potential markers for discriminating the EOC patients and normal subjects by the PLS-DA recognition score map using the mass spectrometry data of IEESI-MS and HPLC-MS. B: the possible mechanisms of these metabolic molecules associated with EOC development.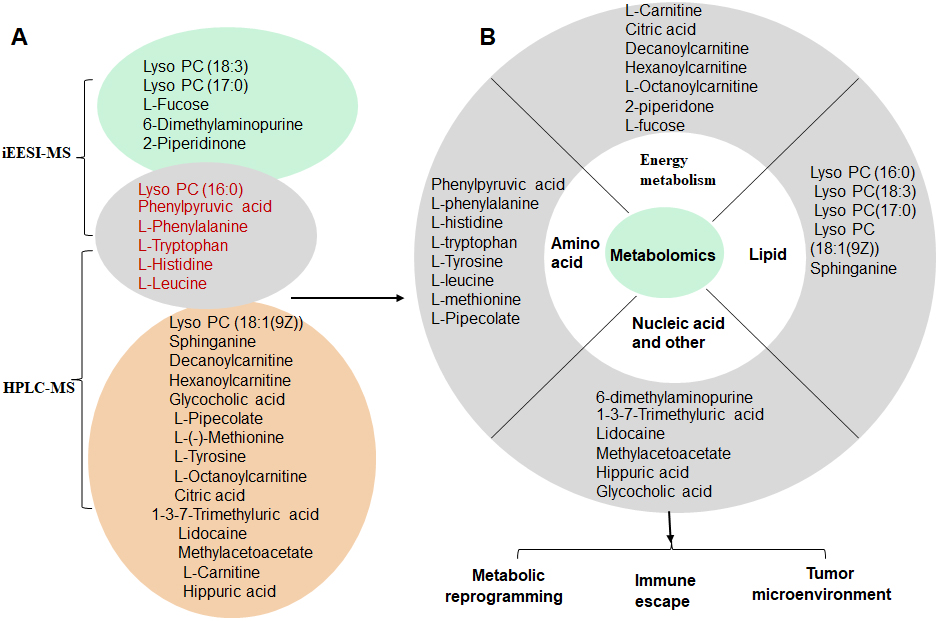
In our study, the differential metabolites-related metabolic pathways were explored by the enrichment analysis according to the highest correlation of the differential metabolites with the metabolic pathways. Using the KEGG analysis, which is a comprehensive database that integrates genome, chemistry, and system function information, metabolic pathways associated with the differential metabolites were explored (Fig. 6). The major metabolism-related pathways and molecules included (1) Metabolites involved in Biosynthesis of amino acids, such as L-Leucine, Citric acid, L-Histidine, L-Tyrosine, Phenylpyruvic acid, L-Phenylalanine, and L-Tryptophan; (2) Metabolites involved in Central carbon metabolism in cancer, such as Citric acid, L-Leucine, L-Phenylalanine, L-Histidine, L-Tryptophan, and L-Tyrosine; (3) The molecules involved in 2-Oxocarboxylic acid metabolism, such as L-Leucine, Citric acid, L-Phenylalanine, L-Tryptophan, L-Tyrosine, and Phenylpyruvic acid; (4) The molecules involved in Aminoacyl-tRNA biosynthesis, such as L-Tryptophan, L-Tyrosine, L-Leucine, L-Phenylalanine, and L-Histidine; (5) Metabolic molecules involved in Protein digestion and absorption, such as L-Leucine, L-Phenylalanine, L-Tryptophan, L-Histidine, and L-Tyrosine. Suggesting the involvement of the metabolic network of EOC. The information on the EOC-associated metabolic pathways might provide directions for future studies. The enrichment analysis results of the differential metabolites are presented in Table 4. KEGG enrichment analysis showed that the differentially expressed genes were mainly involved in the cancer-related signaling pathways, such as energy metabolism, cell migration, and transmembrane transporter.
The results of the enrichment analysis of the differential metabolites
The results of the enrichment analysis of the differential metabolites
Enrichment analysis of the differential metabolites.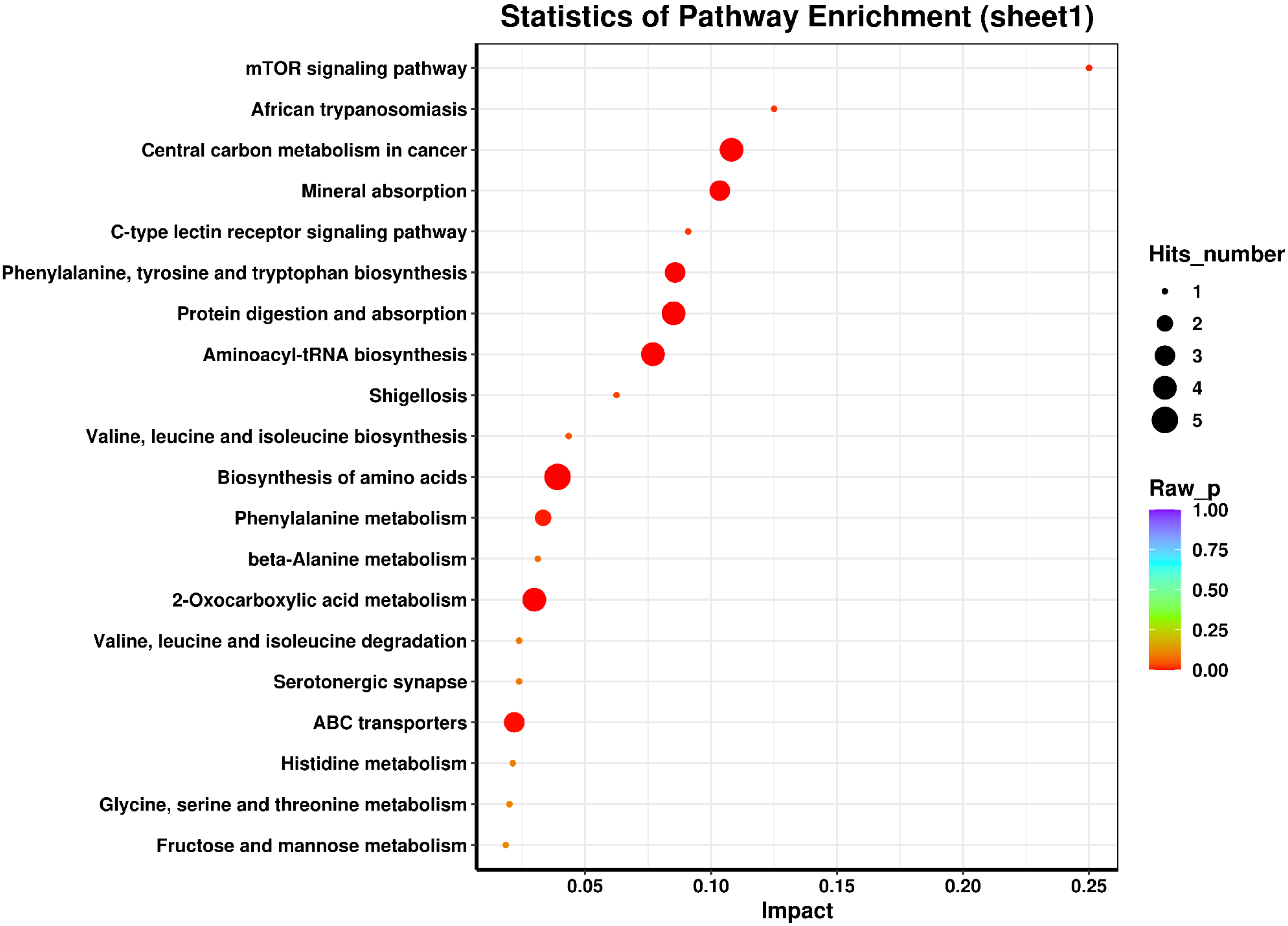
Lipid metabolism
Lipids are a class of heterogeneous metabolites involved in many vital cellular functions, such as membrane construction, energy storage, and signal transduction [19]. Membrane fluidity can directly affect the motility and behavior of tumor cells at different stages of metastatic spread [20]. Lipid metabolism may be involved in cancer-related cell proliferation, inflammation, drug resistance, tumor immune escape, and apoptosis [21, 22]. In our study, aberrant expression of some lipids was found, including Lyso PC (16:0), Lyso PC (18:3), Lyso PC (17:0), Lyso PC (18:1(9Z)), sphinganine, and intermediate products of lipid metabolism: decanoylcarnitine, hexanoylcarnitine, and glycocholic acid. EOC cells tend to migrate to the omentum, which is mainly composed of adipocytes. Co-culture of adipocytes and EOC cells revealed that adipocytes could directly transfer lipid to ovarian cancer cells, promoting tumor growth in vitro and in vivo [23]. The nutritional lipid environment, and lipid subclasses produced by phospholipid cleavage are closely associated with cancer development [24]. Abnormalities of both lysophospholipid, a kind of product of catabolic metabolism of phospholipid (the basic substance of cell membrane), and lipoprotein A (LPA), a type of Low Density Lipoprotein (LDL) cholesterol, have been detected in blood and ascites of the EOC patients. Both are valuable biomarkers and potential therapeutic targets [25, 26, 27]. In this study, abnormal expression of Lyso PC (16:0), a potential biomarker, is found in the plasma of the EOC patients. Aberrant expression of Lyso PC (16:0) is also found in other solid tumors, such as laryngeal [28] and colorectal cancers [29]. Moreover, PC (16:0/16:1) can promote the proliferation of colorectal cancer cells by activating Akt and ERK pathways. PC (16:0/16:1) was added to the culture media to maintain colorectal cancer and non-cancerous cell lines. The growth rate of the colorectal cancer cell line increased about twofold than that of the non-cancerous cell line. Further studies also found that PC (16:0/16:1) could promote the generation of lysophosphatidic acid, thereby increasing the expression of lysophosphatidic acid receptor and activating Akt and ERK pathways, leading to enhancing cell survival and proliferation [29]. Many metabolic enzymes are abnormally expressed in ovarian cancer cells, affecting lipid synthesis or degradation, resulted in disorder of lipid metabolism which is associated with cancer development. At present, rate-limiting metabolic enzyme inhibitors combined with first-line chemotherapy drugs have become a new strategy in treating ovarian cancer [30, 31].
Amino acid metabolism
Metabolism of Amino acids is important for biological processes because of their involvement in the biosynthesis of nucleotides, antioxidants (glutathione), glucosamine, and polyamines. The metabolites of amino acids are also involved in energy-producing processes, such as the tricarboxylic acid cycle [32]. A total of 20 left-handed amino acids are the cornerstone of the human body. Abnormalities of several amino acids were found in this study, including phenyl pyruvic acid, L-phenylalanine, L-histidine, L-tryptophan, L-Tyrosine, L-leucine, L-methionine, and L-Pipecolate. Methionine, phenylalanine, tryptophan, and leucine are nutritionally essential amino acids; the others are semi-essential amino acids. Tryptophan is an essential amino acid that must be obtained from the diet to support physiological processes, including cell growth and maintenance [33]. The L-tryptophan [34] promotes the inherent malignancy of tumor cells and restricts anti-tumor immunity. For this reason, L-tryptophan might be a potential therapeutic target for cancer immunotherapy. In addition, various roles and functions of the tryptophan catabolic pathway in tumor-related immunosuppression have been explored through mechanism-based studies [35]. For example, the indoleamine 2,3-dioxygenase 1 (IDO1)-induced tryptophan depletion in the tumor microenvironment (TME) can inhibit mammalian rapamycin complex (mTORC) and activate the general control nonderepressible 2 (GCN2), a stress response kinase in tumor-infiltrating T cells, leading to incompetence and apoptosis [36]. In preclinical models, the enhanced kynurenine pathway (KP) has been associated with the failure of anti-tumor immunity and tumor progression, which more likely depends on AhR activation [37]. Therefore, the tryptophan catabolic pathway is considered an essential step for tumor cells to evade the innate and adaptive immune system [38]. Besides tryptophan, arginine and histidine also play a significant role in enhancing immunity [39]. Specifically, as previously reported [40], histidine content was significantly decreased in the cancer patient, compared to that in the normal subjects, including colonic carcinomas, prostate cancer [41], and breast cancer [42]. In addition to the differential diagnosis of malignant diseases, histidine can also be used as a predictor. It has been reported that hypoxanthine and histidine in early treated serum are predictive biomarkers for the response to programmed cell death 1 ligand 1 (PD-1) blockade in patients with advanced non-small-cell lung cancer (NSCLC) [12].
Amino acids are the building blocks of proteins and are metabolized as energy substances or function as signaling molecules. Phenylalanine and phenylpyruvate can inhibit the glycolysis of hepatocytes [43]. Destructive effects of phenylalanine and its metabolites on brain glucose metabolism were also reported. Phenylpyruvate may be responsible for reducing glucose consumption by decreasing hexokinase activity, resulting in decreased ATP production in brain cells [44]. Amino acid metabolism was proven to regulate immune response and cancer development by maintaining tumor proliferation and homeostasis. Targeting amino acid metabolism has emerged as a potential therapeutic strategy for cancer patients.
Cell division and apoptosis
6-dimethylaminopurine (6-DMAP) is a protein phosphorylation inhibitor, which could increase intracellular calcium levels in oocytes and inhibit the extrusion of the second polar body. It has been reported that 6-DMAP can induce apoptosis in S-phase stalled HeLa cells, activate the M-phase promoters, and induce premature mitosis in similarly treated hamster cell lines. However, there was little effect of 6-DMAP on asynchronously grown HeLa cells or lovastatin early G1 stagnate cells [45]. Moreover, treatment with 0.6 mM of 6-DMAP for 6 h can induce cell-cycle arrest at the G2-phase [46]. 6-DMAP also induced apoptosis through a p53-independent pathway in a human lymphoma U937 cell line. 6-DMAP-mediated apoptosis is associated with activating a mitochondrial caspase-dependent pathway and [Ca2
Substances related to metabolism
Citric acid
Citrate is essential in metabolic energy production and is involved in synthesizing cholesterol, fatty acids, and isoprene compounds. Citrate also links carbohydrates to fatty acid metabolism [48]. Reduced citrate concentrations have been observed in multiple tumor tissues and in the blood of cancer patients, which might be used to indicate cancer aggressiveness and metabolic prognosis [49]. The determination of citric acid in semen may have a crucial impact on improving the diagnosis of prostate cancer in the early stages of the disease [50]. The levels of citric acid and isocitric acid, both of which are the metabolites of tricarboxylic acid (TCA) cycle, were found elevated in A2780, a human EOC cell line, after treatment with paclitaxel but without change in A-PM3 cells, which is a cell line derived from A2780 exposed to paclitaxel [51]. Citrate can regulate the reprogramming of lipid metabolism in tumor cells, control the fate of tumor cells, promote excess lipid biosynthesis, and cause the senescence of tumor cells. Thus, citrate has been used in tumor therapy [52].
Decanoylcarnitine, hexanoylcarnitine, and L-octanoylcarnitine
L-octanoylcarnitine is a combination of 8-carbon fatty acid with carnitine, which is a crucial product of fatty acid catabolism, suggesting that all fatty acids must enter the inner membrane of mitochondria in the form of lipoylcarnitine for
L-fucose
Fucose is a 6-deoxyhexose with a structure similar to glucose. Fucose has a unique L-configuration, whereas all other naturally occurring sugars in mammals are in the D conformation, making fucose a relatively common component of glycan modification on proteins and lipids. Elevated serum level of fucose was detected in cancer patients, especially in advanced and metastatic tumors, such as stomach and ovarian cancer [56]. Increasing fucose glycosylation of tumor cells contributes to abnormal characteristics such as decreased adhesion and uncontrolled tumor growth. Therefore, detecting serum levels of fucose may be a promising method for early diagnosis and prognosis of various types of cancer [57].
2-piperidone
2-piperidone is a derivative of piperidine, which affects the activity and expression of cytochrome P450 (CYP) 2E1, thereby indirectly affecting oxidative stress and metabolic activation of low molecular weight toxins. A decrease in the plasma level of 2-piperidone was found in the EOC patients [58, 59]. A selective piperidone inhibitor (compound 2) was identified as a highly potent murine double minute-2 inhibitor, with remarkable pharmacokinetic properties and in vivo anti-tumor activity in the SJSA-1 osteosarcoma xenograft models [60].
The differential metabolites related to energy metabolism
Metabolic substrates or the substrates of metabolic pathways are the products that can affect metabolism and participate in various signaling pathways as regulatory factors. The enrichment analysis results of the differential metabolites are presented in Table 4 and Fig. 6 (KEGG enrichment analysis). The enrichment pathways for the abnormal metabolite citric acid included the following aspects: citrate cycle (TCA cycle), glyoxylate and dicarboxylate metabolism, glyoxylate and dicarboxylate metabolism, 2-oxocarboxylic acid metabolism, taste transduction, carbon metabolism, glucagon signaling pathway, biosynthesis of amino acids, and central carbon metabolism in cancer. Moreover, 2-piperidone indirectly affected oxidative stress and metabolic activation by regulating the activity and expression of cytochrome P450 (CYP) 2E1. A decreased plasma level of 2-piperidone in EOC patients has been reported [58, 59]. Our study also obtained similar results. Although we could not identify the pathways or biological functions associated with 2-piperidone, L-fucose, which was abnormally expressed in this study, plays an important role in cell biology. L-fucose and fucosylation are essential for maintaining the immune system and organ development processes, which play an important role in tumor biology and immunology [61]. Therefore, honing and exploiting fucosylation might provide potential therapeutic opportunities in EOC. This study found that the abnormal generation of decanoylcarnitine, l-octanoylcarnitine, and hexanoylcarnitine in patients with EOC may be related to the abnormal fat metabolism of ovarian cancer cells [58, 59]. Fatty acid
The balance between nutrient absorption and energy production, macromolecular biosynthesis, and oxidation-reduction reaction is needed in rapidly proliferating cancer cells. In advanced and metastatic cancer, metabolic reprogramming supports metabolic plasticity and high energy production. Therefore, the therapy targeting metabolic pathways has become the focus of cancer treatment [13].
Comparison between our study and other studies
Compared to previous studies which used HPLC-MS or/and iEESI-MS methods for diagnosis of EOC, we made a summary including the following aspects: (1) Abnormal generation of methionine, histidine, tryptophan, leucine, L-phenylalanine, and 2-Pipecolate in ovarian cancer [58, 63, 64, 65], suggesting the metabolic characteristics of the EOC patients, and implying high amino acid uptake and consumption in tumor tissue; (2) Abnormal generation of lipid metabolites, such as lyso PC (16:0), lyso PC (17:0) [65], and sphinganine/C16 sphinganine [66] in ovarian cancer; (3) HPLC-MS [66] analysis showed that the metabolic characteristics between the normal control group, platinum-sensitive group, and platinum-resistant group were significantly different. These findings may provide theoretical clues for predicting prognosis and chemotherapy response, reversing drug resistance, and discovering new targets for therapeutic intervention. Although different studies found different results, we attributed those differences to the histological and staging characteristics of the ovarian cancer tissue samples and the metabolome analysis techniques.
The clinical application of our findings in future
The biomarkers found in this study, such as lyso PC (16:0), l-phenylalanine, l-leucine, phenylpyruvic acid, l-tryptophan, and l-histidine may be used for clinical application in various ways, such as screening of the ECO patients by distinguishing benign tumors and ECO, as well as excluding the diseases which are easily mixed with EOC, like pelvic inflammatory disease and tuberculosis. Further research, particularly on the relationship between the levels of those biomarkers and the long-term prognosis of ECO, is required to predict the pathological type, clinical stage, and curative effect of therapeutic approaches on ECO.
Limitations and future continuation of this study
First, only a few potential EOC biomarkers were identified in this study, performed in a single center. Therefore, it is required to conduct further multi-center studies which have larger sample sizes for validation and quantitative testing.
Second, changes in plasma metabolomics in EOC patients reflect the disease progression. Therefore, to explore the potential biomarkers by screening, the differences in metabolomics between early (stage I/II) and late (stage III/IV) EOC should be investigated in future studies. It is worth noting that chemotherapy resistance is the main cause of disease progression. Therefore, comparing the changes in plasma metabolomics before and after treatment with platinum-resistant drugs between the platinum-sensitive and platinum-resistant groups is necessary. The comparison results may provide useful clues for predicting chemotherapy response, reversing drug resistance, and even discovering new targets for therapeutic intervention.
Conclusion
Our study provides a comprehensive understanding of the metabolic changes in plasma between the EOC patients and normal control and the significance of the differential metabolites as screening markers for early diagnosis of EOC. This study confirmed that the iEESI-MS is a reliable screening method for EOC patients because it is simple to carry out analysis, with fast and easy-to-use sample pretreatment. This method is also helpful for discovering new potential biomarkers. Metabolomic screening biomarkers might provide reliable indicators for screening and diagnosing EOC and cancer prevention.
Authors’ contributions
Songling Zhang: Put forward the conception; Jiajia Li, Zhenpeng Wang: Interpretation or analysis of data; Zhentong Wei, Wenjie Liu: Preparation of the manuscript; Dongzhen Liu, Linsheng Tan: Supervision; Songling Zhang, Zhentong Wei, Jiajia Li, and Yunhe Yu: Revision for important intellectual content.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-220250.
sj-docx-1-cbm-10.3233_CBM-220250.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-220250.docx
Footnotes
Acknowledgments
The authors appreciate the International Cooperative Joint Laboratory of Nanoarchitecture Chemistry. The work was supported by the Jilin Municipal Department of Finance – Application of metabolomics based on mass spectrometry in early diagnosis and recurrence monitoring of ovarian cancer (No. 20190303143SF), The First Hospital of Jilin University – Application of metabolomics based on mass spectrometry in the diagnosis of ovarian cancer (No. JDYYJCHX003), and the Jilin Provincial Finance Department – Ovarian cancer cells up-regulated NRF2 signaling to adapt to autophagy inhibition (No. JLSWSRCZX2020-088); Special health project of the Department of Finance of Jilin Province (no. JLSCZD2019-078).
Conflict of interest
The authors declare that the research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
