Abstract
BACKGROUND:
It has been discovered that lncRNA ARAP1-AS1 is upregulated and operates as a tumor promoter in many cancers. However, its pattern of expression and potential mechanism in lung adenocarcinoma (LUAD) is still unknown.
METHODS:
The levels of lncRNA ARAP1-AS1, miR-8068, and CEACAM5 expressions in LUAD cell lines and tissues were assessed by conducting western blot and RT-qPCR analyses. MiR-8068’s potential targeting relationships with lncRNA ARAP1-AS1 and CEACAM5 were ascertained by performing bioinformatics analysis. The interaction of lncRNA ARAP1-AS1 with miR-8068 was validated by means of by RIP and luciferase reporter experiments. CCK-8, cell adhesion, and Transwell migration experiments were conducted to study how lncRNA ARAP1-AS1 affects LUAD cell migration, adhesion, and proliferation. To confirm the function of lncRNA ARAP1-AS1 in vivo, a tumor formation experiment was executed.
RESULTS:
An elevated expression of lncRNA ARAP1-AS1 was observed among the LUAD cells and tissues. The overexpression of lncRNA ARAP1-AS boosted cell proliferation, adhesion, and migration in LUAD and also favored in vivo tumor growth. MiR-8068 was found to be lncRNA ARAP1-AS1’s target gene. MiR-8068 overexpression partially antagonized lncRNA ARAP1-AS1’s promotive effect on proliferation, viability, and adhesion. Meanwhile CEACAM5 could alleviate the miR-8068-induced inhibition of tumor growth. The negative correlation of miR-8068 with lncRNA ARAP1-AS1 or CEACAM5 was also revealed.
CONCLUSION:
To upregulate CEACAM5 expression lncRNA ARAP1-AS1 targeted miR-8068, thus promoting the progression of LUAD. This indicates that the lncRNA ARAP1-AS1/miR-8068/CEACAM5 axis has potential as a therapeutic target in LUAD treatment.
Introduction
Lung cancer is the second most commonly diagnosed cancer and leading cause of cancer death worldwide [1]. Lung adenocarcinoma (LUAD) represents the primary histological type of lung cancer with unsatisfactory prognosis [2, 3]. In spite of the developments in novel cancer therapies, LUAD still has a 5-year survival rate that is below 20% [4]. Previous research have revealed numerous long non-coding RNAs (lncRNAs) participating in LUAD progression [5, 6, 7]. However, their types and specific mechanisms have yet to be explored completely. Hence, searching for robust biomarkers may help tailor targeted therapeutic treatments for patients with LUAD.
LncRNAs endogenous RNAs that do not code for proteins and contain over 200 nucleotides are involved in numerous malignant cancers. They perform a pivotal role in various cellular functions by regulating gene expression, translation, and chromatin organization [8]. In the recent years, several lncRNAs have been found to be dysregulated in LUAD cells, wherein they served as tumor promoters or suppressors. For example, lncRNA ZFPM2-AS1 induces LUAD by negatively regulating miR-18b-5p and upregulating VMA21 expressions [9]. The pyroptosis-related lncRNA FAM83A-AS1 exerts a carcinogenic effect on LUAD through the miR-150-5p/MMP14 axis [10]. Meanwhile lncRNA LINC00968 suppresses cell invasion, migration, and proliferation in LUAD by targeting the miR-22-5p/CDC14A axis [7]. It has been documented that the lncRNA ARAP1 antisense RNA 1 (ARAP1-AS1) promotes tumorigenesis and metastasis in many cancer [11, 12, 13]. One study documented a high expression of ARAP1-AS1 in lung cancer [14]. Nonetheless, the detailed mechanism of lncRNA ARAP1-AS1 remains unknown.
LncRNAs that influence cancer occurrence and development via gene regulation have complex mechanisms [15]. This includes binding to targeting miRNAs that promote or suppress tumorigenesis. For instance, studies have shown a variety of miRNA-regulated pathways that participate in cancer progression. The highly expressed miR-3529/miR-150/miR-31/miR-30d were found to inhibit the malignancy of LUAD [10, 16, 17, 18]. As for miR-8068, its upregulation is detectable in autoimmune hepatitis, a liver inflammation status related to liver cancer. MiR-8068 is also associated with the pathogenicity of SARS-CoV-2 and host immune responses. However, its role in lung cancer has yet to be reported.
CEACAM5 is located at chromosome 19q13.2 and consists of 10 exons. CEACAM5 is responsible for encoding a cell surface glycoprotein, which represents the founding member of the carcinoembryonic antigen (CEA) protein family. The encoded protein has been utilized as a clinical biomarker for colorectal, gastrointestinal, and lung cancers, as it may promote tumor development via its involvement in cell adhesion [19, 20, 21]. In addition, the encoded protein may modulate cell polarity, differentiation, and apoptosis [22]. Nevertheless, how CEACAM5 regulates LUAD remains obscure.
In this research, we intended to study the effects of lncRNA ARAP1-AS1 on LUAD. Further, bioinformatics analyses clarified the interaction of miR-8068 with CEACAM5 and lncRNA ARAP1-AS1. This novel interaction axis was validated in vitro. Our findings may contribute in uncovering the molecular mechanisms involved in LUAD and help identify novel biomarkers for its targeted therapies.
Materials and methods
Collection of tissue specimens
Thirty-eight sets of lung cancer and normal adjacent tissues were gathered from patients suffering from LUAD. The Ethics Committee of The First Affiliated Hospital of Chengdu Medical College granted approval to the protocols used in this study. All participants who agreed to the usage of their specimens in the study provided a written informed consent. The execution of this study complied with the World Medical Association’s Declaration of Helsinki.
Cell culture
The human normal lung cell BEAS-2B and four human LUAD cell lines (A549, PC9, Calu-3, DV-90) were bought from the Chinese Academy of Sciences, Shanghai Institute of Biochemistry and Cell Biology (Shanghai, China). DMEM (Invitrogen, USA) with an added fetal bovine serum (FBS; 10%; Hyclone, USA) was used in culturing all cells. The cultures were incubated in a 37
RNA extraction and RT-qPCR
The cells and tissues were treated with the TRIZOL reagent (ThermoFisher, USA) to extract their total RNAs. Afterward, cDNA were synthesized by subjecting the total RNAs to reverse transcription using an RT primer and AMV reverse transcriptase (Takara, Japan). MiRNA, lncRNA, and mRNA were quantified with Taqman probes (ThermoFisher, USA). RT-qPCR was accomplished in an ABI 7300 Sequence Detection Systems (Applied Biosystems) with a Taqman PCR Kit (ThermoFisher, USA). The endogenous controls adopted for the expressions of miR-8068, CEACAM5, and lncRNA ARAP1-AS1 were U6 and GAPDH. Table 1 lists the primer sequences.
Real-time PCR primer sequences
Real-time PCR primer sequences
Synthgene, China synthesized the miR-8068 mimic (miR-8068), mimic NC (miR-NC), pcDNA-CEACAM5 overexpression vector (OE-CEACAM5) its empty vector (OE-NC), siRNA targeting ARAP1-AS1(si-ARAP1-AS1), and its negative control (si-NC). First, 1
For the ectopic expression of lncRNA ARAP1-AS1, lentiviral particles containing pLenti-CMV-lncRNA ARAP1-AS1 (OE-lnc) and empty pLenti-CMV (OE-NC) were produced by OBio, Shanghai, China. The lentiviral expression vectors were transfected into 293T cells, followed by a 7-week puromycin maintenance. The lentiviruses produced were then utilized to infect A549 and PC9 cells. Forty-eight hours later, RT-qPCR was performed to analyze the cells.
Localization within the cytoplasm and nucleus
Following the manufacturer’s instructions, NE- PERTM Nuclear and Cytoplasmic Extraction Reagents Kit (Thermo Scientific) was employed to locate lncRNA ARAP1-AS1 within the A549 and PC9 cell lines. The cytoplasmic and nuclear components of the cells were separated and collected. RT-qPCR was conducted to estimate lncRNA ARAP1-AS1 expressions within the cell cytoplasm and nucleus. GAPDH was adopted as the cytoplasmic positioning control while U6 was utilized as the nuclear positioning control.
Cell counting Kit-8 (CCK-8) assay
The proliferation and viability of the cells were evaluated by means of the CCK-8 experiment. On 96-well plates, a 100
Cell adhesion assay
A 96-well culture plate was pre-coated with 30
Transwell migration assay
Transwell inserts (Corning, Inc. USA) and 24-well plates were used for this assay. A549 and PC9 cells (
Dual-luciferase reporter assay
The relationship of miR-8068 with the target genes was tested by means of the luciferase reporter experiment. MiR-8068’s wild-type (WT) and mutated (MUT) binding domains in each target gene were amplified and subsequently inserted into psiCHECK-2 vectors (Synthgene Biotech, Nanjing, China). Four luciferase reporter plasmids were then constructed: ARAP1-AS1-WT ARAP1-AS1-Mut, CEACAM5-3’-UTR-Mut and CEACAM5-3’-UTR-WT The A549 and PC9 cell lines were inoculated on 24-well culture plates and then allowed to reach 60% confluence. A Lipofectamine 2000 (Thermofisher, USA) was used in transfecting the A549 and PC9 cell lines with a combination of a luciferase reporter plasmid (0.5
RNA-binding protein immunoprecipitation (RIP) assay
The RIP assay was carried out with an EZ-Magna RIP Kit (EMD Millipore) to identify RNA-protein interactions. In brief, PC9 and A549 cells were lysed in a RIP lysis buffer supplemented with cocktail (Roche Diagnostics). The cell supernatants were then extracted and then maintained at 4
Tumor formation experiment in nude mice
BALB/c nude mice with thymus-deficiency (
Western blot
A RIPA lysis buffer supplemented with PMSF was utilized to lyse the cells at 4
Statistical analysis
The statistical analyses of the data were performed in SPSS 18.0 (SPSS, Inc.). Each experiment was conducted thrice, and the collected data were indicated in the following format: mean
Results
ARAP1-AS1 overexpression boosts the growth of LUAD cells in vivo and in vitro
We first assessed lncRNA ARAP1-AS1 expression among the LUAD cell lines and in the 38 sets of normal and LUAD tissues. Relative to the normal lung cell BEAS-2B, all the tumor cell lines (A549, PC9, DV-90, and Calu-3) exhibited an upregulation of ARAP1-AS1 (Fig. 1A). The highest ARAP1-AS1 levels were observed among the A549 and PC9 cells, hence they were selected for the succeeding experiments. The LUAD and adjacent normal tissues were evaluated for ARAP1-AS1 expression via RT-qPCR. As shown in Fig. 1B, ARAP1-AS1 had a significantly higher expression in LUAD tissues than in normal ones. The proportions of ARAP1-AS1 expression within the cytoplasm and nuclei of PC9 and A549 cell lines were detected. The results showed that ARAP1-AS expression was mainly enriched in cytoplasmic part of A549 and PC9 (Fig. 1C). To investigate ARAP1-AS1’s oncogenic ability and role in LUAD progression, its overexpression vector, knockdown vector, and negative control were transfected into the PC9 and A549 cell lines. The transfection efficiency is validated in Fig. 1D, showing that ARAP1-AS1 overexpression vector led to the increase of ARAP1-AS1 expression, whereas ARAP1-AS1 knockdown vector led to the decrease of ARAP1-AS1 expression in PC9 and A549 cells The CCK-8 assay revealed that ARAP1-AS1 overexpression significantly promoted the viability of the PC9 and A549 cell lines, but ARAP1-AS1 knockdown inhibited the viability (Fig. 1E). Moreover, the outcomes of the Transwell migration experiment suggested that the overexpression of ARAP1-AS1 stimulated cell migration more than the NC group, whereas ARAP1-AS1 impaired cell migration (Fig. 1F). It was demonstrated in the cell adhesion assay that the overexpression of ARAP1-AS1 in the PC9 and A549 cell lines strengthened tumor cell adhesion, but ARAP1-AS1 knockdown showed the opposite effect (Fig. 1G). Collectively, lncRNA ARAP1-AS1 exerted oncogenic effects on LUAD growth in vitro. We also conducted an in vivo experiment in mice to examine how ARAP1-AS1 affected LUAD tumor growth. The outcomes revealed that mice with ARAP1-AS1 overexpression had LUAD tumors that are larger in appearance and measurements than those from the NC mice (Fig. 2A and B). Moreover, the tumors from the OE-lnc group were heavier than the tumors from the OE-NC group (Fig. 2C). Our findings evidence that ARAP1-AS1 operates as a tumor oncogene during the progression of LUAD.
LncRNA ARAP1-AS1 overexpression promotes cell viability, proliferation, and adhesion. (A) LncRNA ARAP1-AS1 expression was quantified in normal lung cells (BEAS-2B) and all LUAD cell lines (A549, PC9, Calu-3, and DV-90) via RT-qPCR. 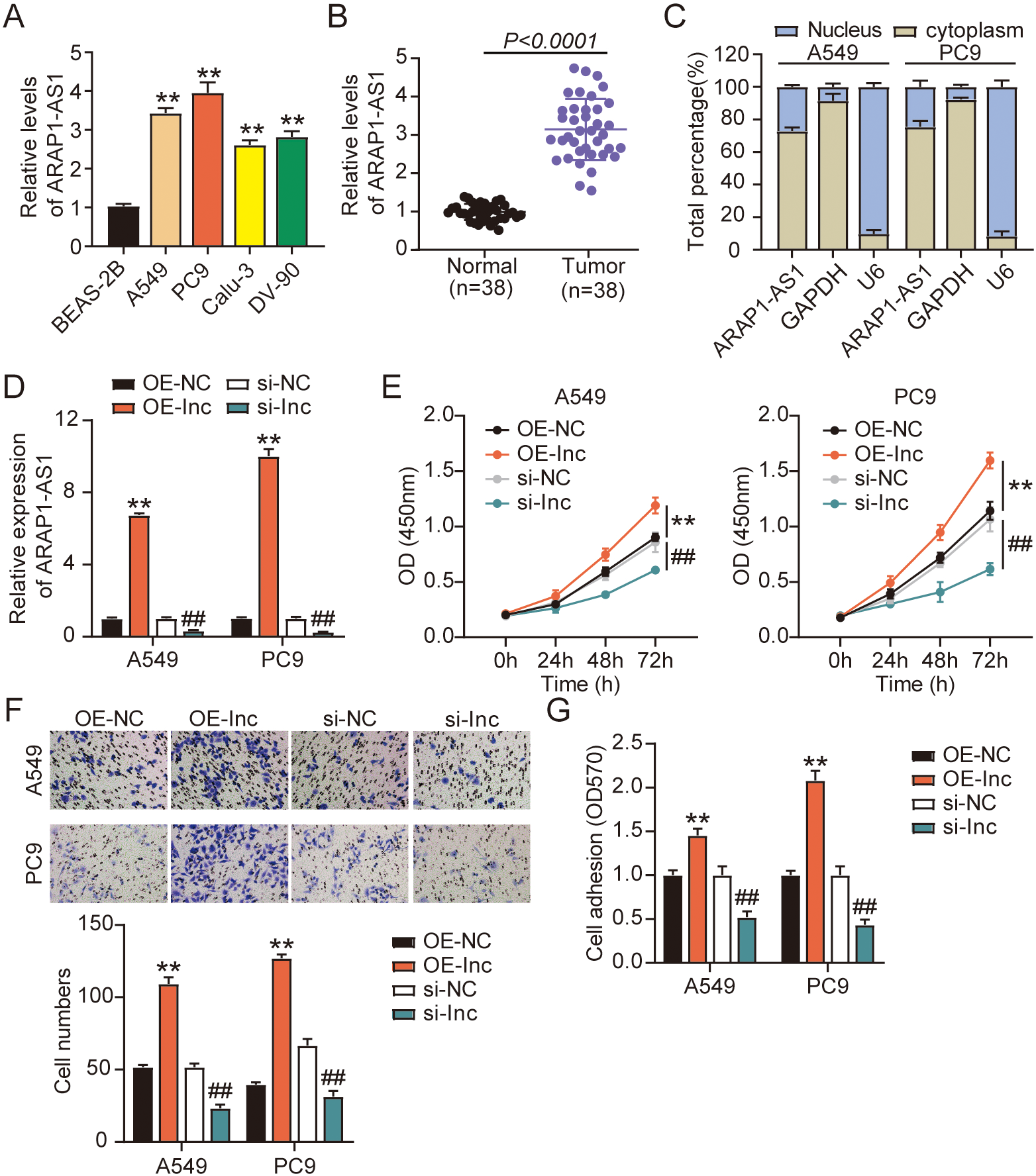
LncRNA ARAP1-AS1 boosts tumor growth in vivo. Nude mice were subcutaneously injected with infected A549 cells. (A) Photograph of the xenograft tumors, 6-weeks after the subcutaneous implantation of A549 cells. (B) Growth curves of xenograft tumors. (C) Weight of xenograft tumors on the 6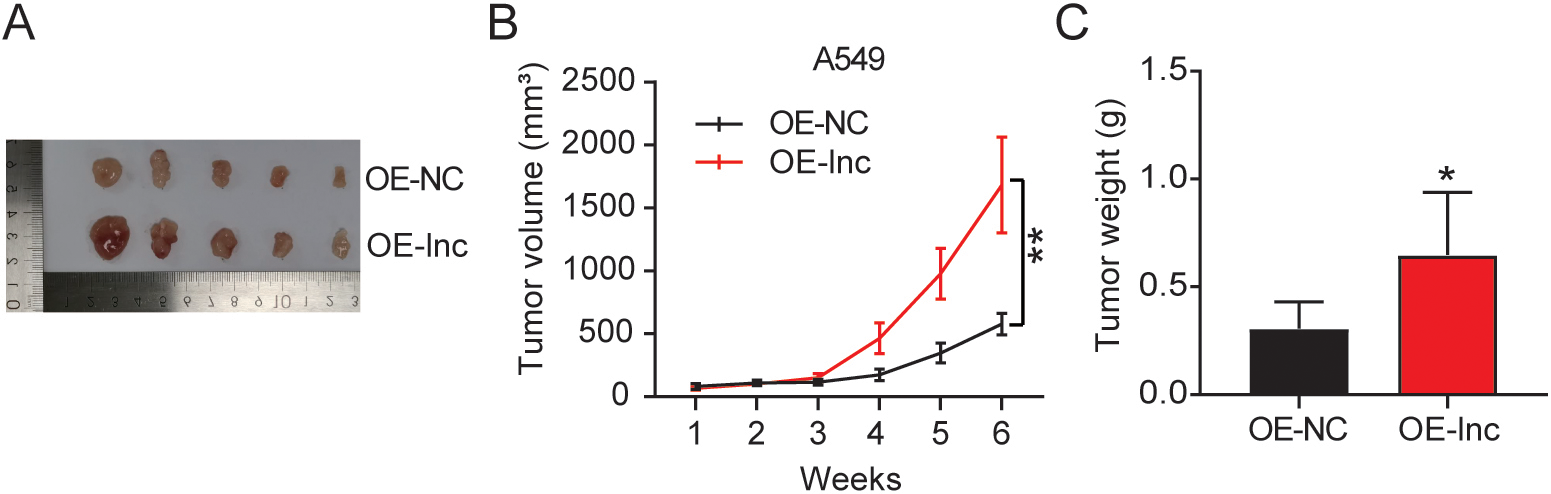
LncRNA ARAP1-AS1 directly interacts with miR-8068. (A) The promising ARAP1-AS1 binding site in the region of miR-8068 was identified by starBase. (B) The relative luciferase signals in PC9 and A549 cells that had a combined transfection of ARAP1-AS1-WT/MUT luciferase reporter vectors and a miR-8068 mimic or mimic NC. 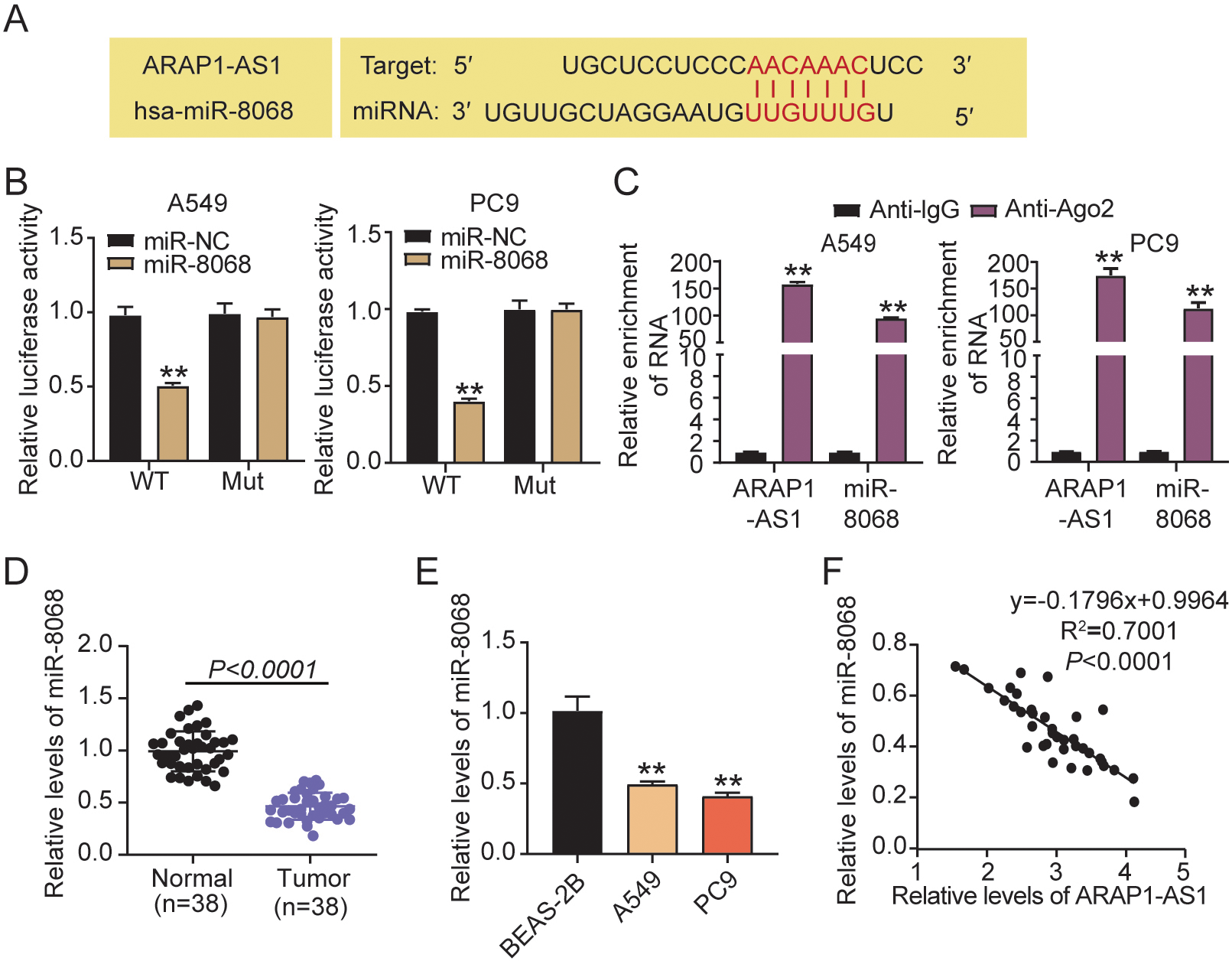
The suppression of cell viability, migration, and adhesion by miR-8068 upregulation is counteracted by ARAP1-AS1 overexpression. PC9 and A549 cells were transfected with ARAP1-AS1 overexpressing vectors (OE-lnc), OE-NC, miR-8068 mimic (mimic), mimic-NC, and OE-lnc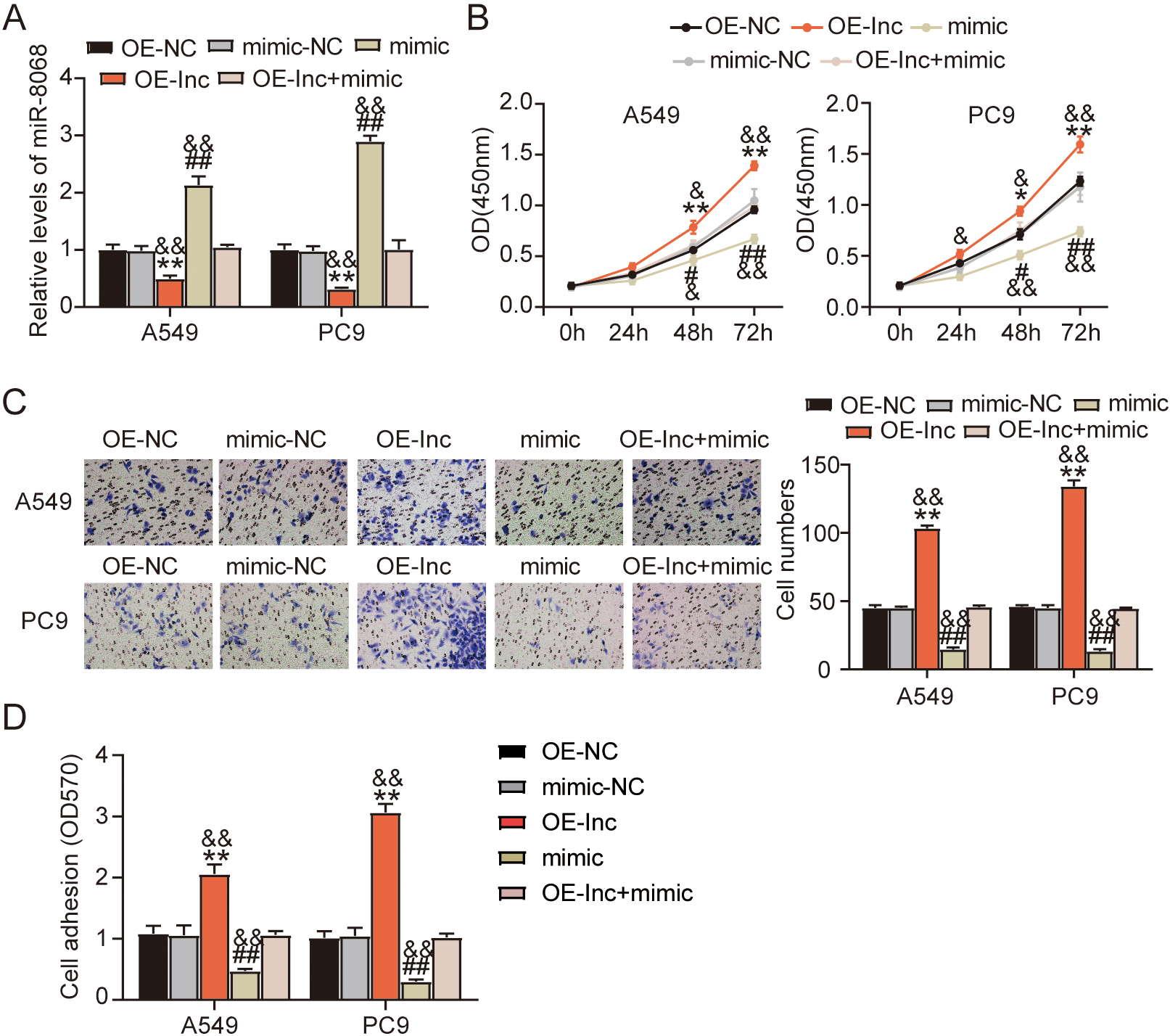
CEACAM5 is a miR-8068 target gene. (A) Three genes (CD24, EPCAM, CEACAM5) were related to cell proliferation and cell adhesion based on STRING analysis. (B) RT-qPCR analysis of the expression levels of CD24, EPCAM, and CEACAM5 in tumoral and normal tissues. (C) TargetScan predicted the promising miR-8068 binding sites in CEACAM5 3’UTR. (D) Relative values of the luciferase signals were assessed in PC9 and A549 cells that had a combined transfection of a CEACAM5 3’UTR-WT/MUT luciferase reporter vector and a miR-8068 mimic or mimic NC. 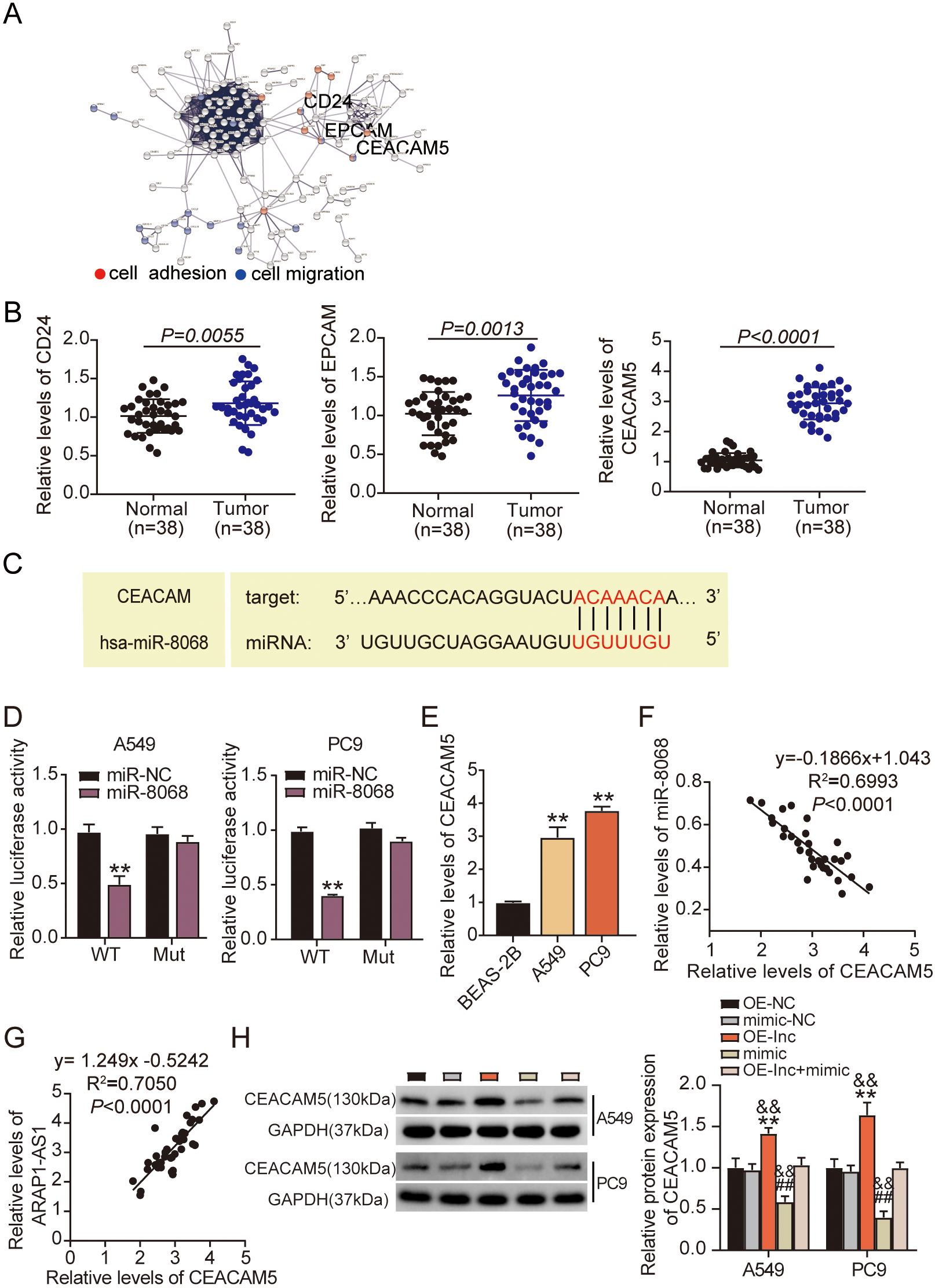
MiR-8068 tones down CEACAM5 expression and exerts a suppressive effect in LUAD progression. CEACAM5 overexpression vectors (OE-CEACAM5), OE-NC, miR-8068 mimic (mimic), mimic-NC, and OE-CEACAM5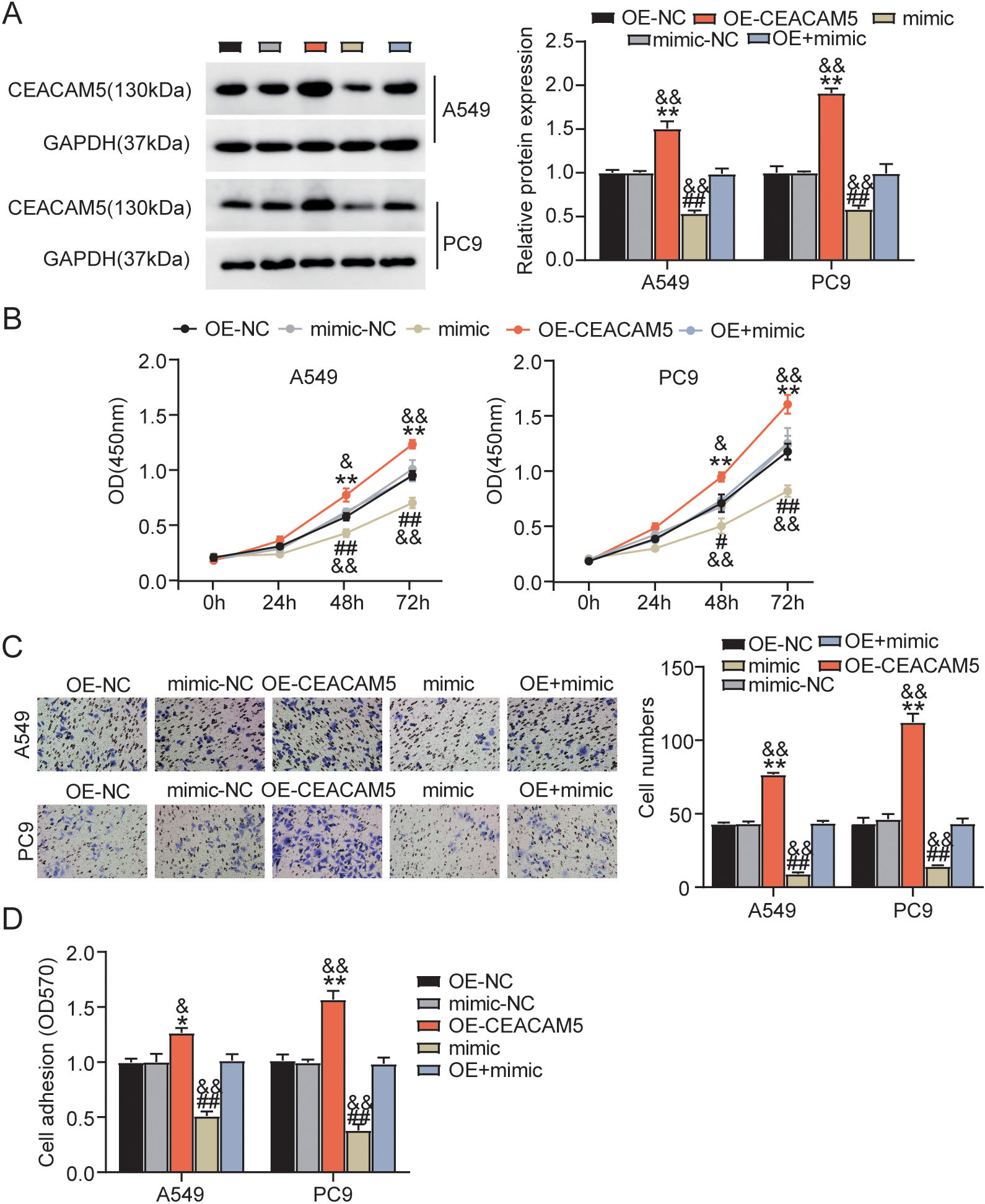
The mechanism of ARAP1-AS1/miR-8068/CEACAM5 axis in LUAD.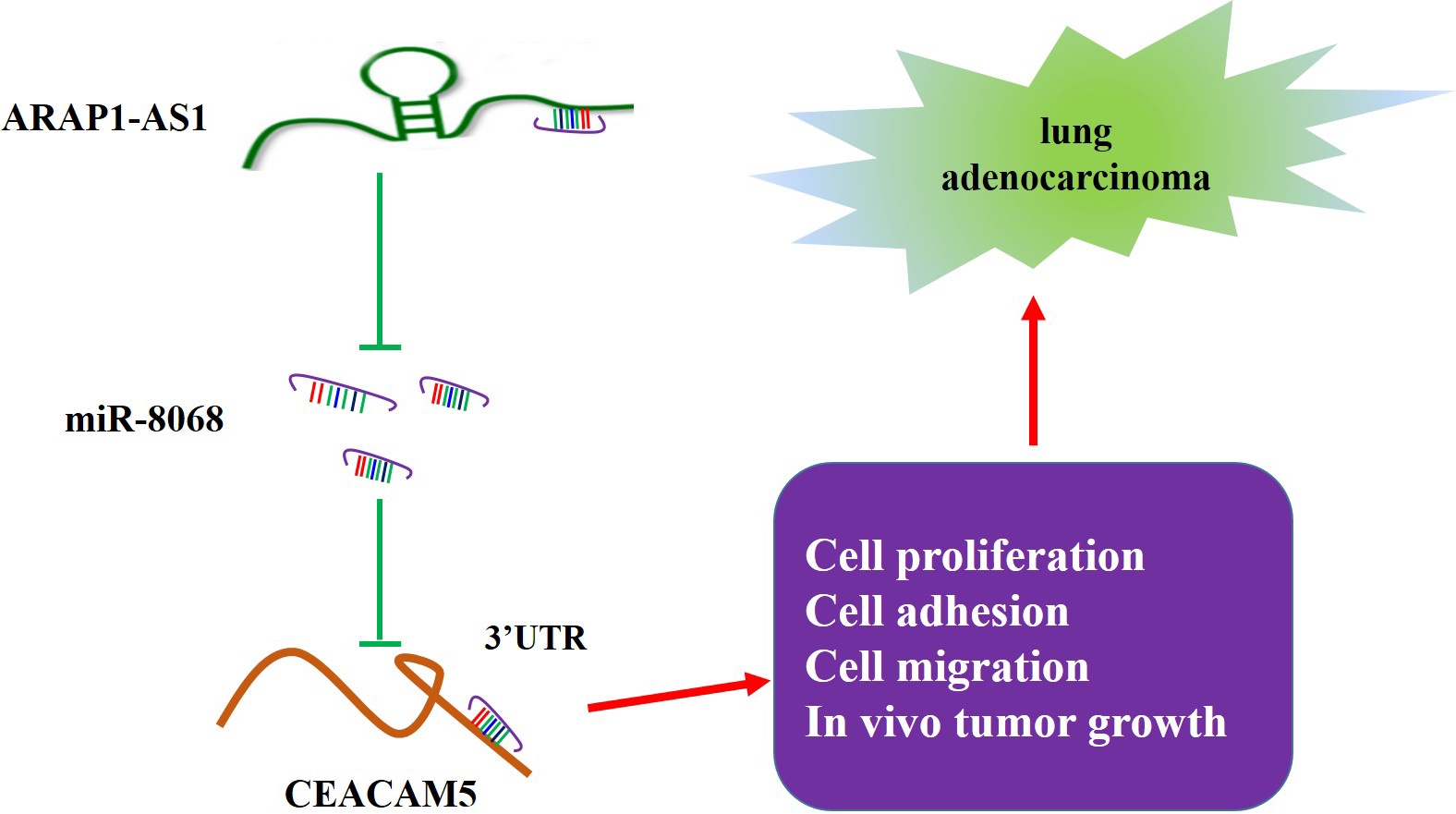
We conducted bioinformatics analyses in starBase (
MiR-8068 is involved inARAP1-AS1-mediated oncogenicity of LUAD in vitro
To investigate miR-8068’s relevance in the ARAP1-AS1-mediated LUAD progression, RT-qPCR was carried out to quantify miR-8068 expression in the PC9 and A549 cell lines transfected with OE-lnc and miR-8068 mimic. Overexpressing ARAP1-AS1 hampered the miR-8068-mimic-induced stimulation of miR-8068 expression in LUAD cells Fig. 4A. In the functional analyses, CCK-8 results revealed that exogenous miR-8068 significantly decreased the cell viability (Fig. 4B). However, this decrease in viability was rescued by the simultaneous transfection of miR-8068 mimic and OE-lnc. Furthermore, a similar trend was observed when the combined transfection of the two restored to normal levels the suppressed LUAD cell migration effected by the miR-8068 mimic (Fig. 4C). The cell adhesion analysis demonstrated that the miR-8068 mimic also lessened the cell adhesion, and yet this inhibitory effect was offset by ARAP1-AS1 overexpression (Fig. 4D). To conclude, the miR-8068 mimic inhibits the LUAD progression induced by the overexpression of ARAP1-AS1.
MiR-8068 targets CEACAM5
We investigated the molecular mechanism underlying the ARAP1-AS1-driven LUAD progression, by utilizing the GEPIA website (
MiR-8068 tones down CEACAM5 expression and exerts its suppressive effect on LUAD progression
In view of the abovementioned findings, we further verified the mechanism of the miR-8068/CEACAM5 axis in LUAD progression by transfecting CEACAM5/miR-8068 NC mimics, CEACAM5 overexpression vector, miR-8068 mimic, and a combination of CEACAM5 and miR-8068 mimic into A549 and PC9 cells. Western blot analysis uncovered that miR-8068 overexpression remarkably suppressed the expression levels of CEACAM5 protein (Fig. 6A). This outcome, however, was reversed by overexpressing CEACAM5. The results of the CCK-8 and Transwell migration experiments indicate that a high CEACAM5 expression could promote cell viability and migration (Fig. 6B and C). Nonetheless, these promotive effects may be counteracted by the introduction of exogenous miR-8068. Moreover, the adhesion analysis revealed trends that are similar with that of the proliferation assessment (Fig. 6D). Altogether, these findings evidence the crucial role of lncRNA ARAP1-AS1 in LUAD tumorigenesis, executed via the regulation of the miR-8068/CEACAM5 axis.
Discussion
LUAD is the most prevalent lung cancer and is linked to high morbidity and poor prognosis worldwide [3]. Clinically, the targeted treatment based on molecular alteration, which drives cancer development and maintenance, has made progress in LUAD [23]. Studies have reported multiple lncRNAs involved in the progression and prognosis of LUAD [16, 24]. Hence, dissecting the roles of these lncRNAs in LUAD is important to the development of personalized therapies. Our present study demonstrated that lncRNA ARAP1-AS1 acted as an oncogenic driver of LUAD by partly regulating the miR-8068/CEACAM5 axis (Fig. 7). Our findings might provide a novel target for LUAD intervention.
Evidence have shown the strong association of altered LncRNAs with LUAD. For example, LncRNA LINC01287 stimulates the proliferation and inhibits the apoptosis of LUAD cells by upregulating RNASEH2A after competitively binding with miR-3529-5p [16]. The novel lncRNA, ARAP1-AS1, was previously found be to a tumor promoter in many cancers. For example, ARAP1-AS1 activates the Wnt/
As lncRNAs harbor miRNA response elements (MREs), lncRNAs may serve as ceRNAs to inhibit miRNA expression [26]. Our findings suggest that miR-8068 was ARAP1-AS1’s target miRNA. We identified and validated the negative regulatory association between miR-8068 and ARAP1-AS1 in LUAD cells. The re-upregulation of miR-8068, accomplished by transfecting miR-8068 mimics into the A549 and PC9 cell lines, considerably repressed the viability, adhesion, and migration of cells. The oncogenic effect of ARAP1-AS1 in LUAD progression antagonized the suppressive effect of miR-8068. This indicates that ARAP1-AS1 may serve as a progress promoter in LUAD by interacting miR-8068. MiR-8068’s differential expression has been detected in serval neurodegenerative disorders as well as in osteoarthritis. In cancers, bioinformatics analyses indicate that miR-8068 might be involved in the kidneys’ clear cell renal carcinoma. Nevertheless, no study has reported its role in LUAD malignancy. To our knowledge, this study has been the first to reveal miR-8068’s inhibitory role in cancer progression, shedding new light on the specific molecular regulatory mechanism of cancer.
Through bioinformatics analyses, we screened out CEACAM5 as a potential miR-8068 target gene. The CEACAM gene family mutations have been reported to contribute in numerous cancers, such as breast, colon, gastric, and lung cancers [27, 28, 29]. Among them, CEACAM5 is a commonly recognized biomarker of cancer occurrence and recurrence. A recent study found that CEACAM5 upregulation could stimulate the migration and proliferation of tumor cells to induce non-small-cell lung cancer [29]. Comparable results were presented in our research. We studied the expression levels of CEACAM5 mRNA in LUAD tissues and further demonstrated that its overexpression augmented the oncogenic potential of LUAD cells. However, the overexpression of miR-8068 mimics neutralized the effect of CEACAM5 on LUAD cell malignancy. Our findings evidence that miR-8068 executes a vital role in LUAD progression through CEACAM5 regulation.
More detailed mechanisms and signal pathways of lncRNA ARAP1-AS1 in lung cancer requires further studies. Also, the functions of the miR-8068/CEACAM5 axis needs further verification through clinical investigations.
Conclusion
We presented the elevated expression of lncRNA ARAP1-AS1 in LUAD cells and tissues, which was also accompanied by an upregulated CEACAM5 and downregulated miR-8068 expressions. The overexpression of miR-8068 could partly antagonize the stimulating influence of ARAP1-AS1 on the adhesion, migration, and viability of LUAD cells. Furthermore, CEACAM5 overexpression can inhibit the miR-8068-induced suppression of proliferation among the LUAD cell lines. Our findings evidence that the lncRNA ARAP1-AS1/miR-8068/CEACAM5 axis is a promising therapeutic target in LUAD treatment.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Ethics approval
The Ethics Committee of The First Affiliated Hospital of Chengdu Medical College (Chengdu, China) approved this research. This study was executed in accordance with the World Medical Association’s Declaration of Helsinki. All patients submitted signed informed consents.
The Ethics Committee of The First Affiliated Hospital of Chengdu Medical College granted approval to this study. The execution of the study observed the national and institutional guidelines for animal care and use. All animal experiments had been conducted in strict adherence to the ARRIVE guidelines.
Consent for publication
Consent for publication was obtained from all participants.
Competing interests
The authors declare that they have no conflicts of interests.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by The First Affiliated Hospital of Chengdu Medical College High-level Talents Research Start-up Fund Project (CYFY-GQ26).
Authors’ contributions
Conception: XFZ.
Interpretation or analysis of data: ZQW and XFZ.
Preparation of the manuscript: HW.
Revision for important intellectual content: XBW and ZQW.
Supervision: all authors.
All authors have read and approved this manuscript.
Footnotes
Acknowledgments
None.
